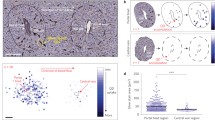
Abstract
Introduction
Lipid nanoparticles (LNPs) tend to accumulate in the liver due to physiological factors. Whereas the biological mechanisms that promote LNP delivery to hepatocytes have been reported, the mechanisms that promote delivery to other cell types within the liver microenvironment are poorly understood. Single cell profiling studies have recently identified subsets of Kupffer cells and hepatic endothelial cells with distinct gene expression patterns and biological phenotypes; we hypothesized these subtypes would differentially interact with nanoparticles.
Methods
To test the hypothesis, we quantified nucleic acid (i) biodistribution and (ii) functional mRNA delivery within the liver microenvironment using two clinically relevant LNPs in vivo.
Results
We found that these LNPs distribute nucleic acids distribute to Kupffer cells and liver endothelial cells as efficiently as they distribute to hepatocytes, yet result in more functional mRNA delivery to endothelial cells. Additionally, we found these LNPs differentially accumulate in Kupffer and endothelial cell subsets.
Conclusions
These data suggest subsets of liver microenvironmental cells can differentially interact with nanoparticles in vivo, thereby altering LNP delivery. More generally, the data suggest that nucleic acid biodistribution is not sufficient to predict functional nucleic acid delivery in vivo.




Similar content being viewed by others
References
Adams, D., et al. Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis. N. Engl. J. Med. 379:11–21, 2018.
Bahl, K., et al. Preclinical and clinical demonstration of immunogenicity by mRNA vaccines against H10N8 and H7N9 influenza viruses. Mol. Ther. 25:1316–1327, 2017.
Chen, D., et al. Rapid discovery of potent siRNA-containing lipid nanoparticles enabled by controlled microfluidic formulation. J. Am. Chem. Soc. 134:6948–6951, 2012.
Dahlman, J. E., et al. In vivo endothelial siRNA delivery using polymeric nanoparticles with low molecular weight. Nat. Nano 9:648–655, 2014.
Dong, Y., et al. Lipopeptide nanoparticles for potent and selective siRNA delivery in rodents and nonhuman primates. Proc. Natl. Acad. Sci. USA 111:3955–3960, 2014.
Finn, J. D., et al. A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing. Cell Rep. 22:2227–2235, 2018.
Jiang, C., et al. A non-viral CRISPR/Cas9 delivery system for therapeutically targeting HBV DNA and pcsk9 in vivo. Cell Res. 27:440–443, 2017.
Kauffman, K. J., et al. Optimization of lipid nanoparticle formulations for mRNA delivery in vivo with fractional factorial and definitive screening designs. Nano Lett. 15:7300–7306, 2015.
Kauffman, K. J., et al. Rapid, single-cell analysis and discovery of vectored mRNA transfection in vivo with a loxP-flanked tdTomato reporter mouse. Mol. Therapy Nucleic Acids 10:55–63, 2018.
Lorenzer, C., M. Dirin, A. M. Winkler, V. Baumann, and J. Winkler. Going beyond the liver: progress and challenges of targeted delivery of siRNA therapeutics. J. Control Release 203:1–15, 2015.
Love, K. T., et al. Lipid-like materials for low-dose, in vivo gene silencing. Proc. Natl. Acad. Sci. USA 107:1864–1869, 2010.
MacParland, S. A., et al. Phenotype determines nanoparticle uptake by human macrophages from liver and blood. ACS Nano 11:2428–2443, 2017.
MacParland, S. A., et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9:4383, 2018.
Miller, J. B., et al. Non-viral CRISPR/Cas gene editing in vitro and in vivo enabled by synthetic nanoparticle co-delivery of Cas9 mRNA and sgRNA. Angew. Chem. Int. Ed. Engl. 56:1059–1063, 2017.
Patel, S., et al. Boosting intracellular delivery of lipid nanoparticle-encapsulated mRNA. Nano Lett. 17:5711–5718, 2017.
Paunovska, K., et al. Analyzing 2000 in vivo drug delivery data points reveals cholesterol structure impacts nanoparticle delivery. ACS Nano 12:8341–8349, 2018.
Paunovska, K., et al. Nanoparticles containing oxidized cholesterol deliver mRNA to the liver microenvironment at clinically relevant doses. Adv. Mater. 31:e1807748, 2019.
Richner, J. M., et al. Modified mRNA vaccines protect against Zika virus infection. Cell 168:1114–1125, 2017.
Sabnis, S., et al. A novel amino lipid series for mRNA delivery: improved endosomal escape and sustained pharmacology and safety in non-human primates. Mol. Ther. 26:1509–1519, 2018.
Sago, C. D., et al. Modifying a commonly expressed endocytic receptor retargets nanoparticles in vivo. Nano Lett. 18:7590–7600, 2018.
Sago, C. D., et al. High-throughput in vivo screen of functional mRNA delivery identifies nanoparticles for endothelial cell gene editing. Proc. Natl. Acad. Sci. 115:e9942–e9952, 2018.
Sago, C. D., et al. Barcoding chemical modifications into nucleic acids improves drug stability in vivo. J. Mater. Chem. B 6:7197–9203, 2018.
Sato, Y., et al. Understanding structure-activity relationships of pH-sensitive cationic lipids facilitates the rational identification of promising lipid nanoparticles for delivering siRNAs in vivo. J. Control Release 295:140–152, 2019.
Semple, S. C., et al. Rational design of cationic lipids for siRNA delivery. Nat. Biotechnol. 28:172–176, 2010.
Shalek, A. K., et al. Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature 510:363–369, 2014.
Tavares, A. J., et al. Effect of removing Kupffer cells on nanoparticle tumor delivery. Proc. Natl. Acad. Sci. USA 114:e10871–e10880, 2017.
Tsoi, K. M., et al. Mechanism of hard-nanomaterial clearance by the liver. Nat. Mater. 15:1212–1221, 2016.
Villani, A. C., et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 2017. https://doi.org/10.1126/science.aah4573.
Yin, H., et al. Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing. Nat. Biotechnol. 35:1179–1187, 2017.
Zeisel, A., et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347:1138–1142, 2015.
Acknowledgments
The authors thank Sommer Durham and the Georgia Tech Cellular Analysis and Cytometry Core. Additionally, the authors thank Dalia Arafat and the Genome Analysis Core. J.E.D. thanks Jordan Cattie and Taylor E. Shaw.
Conflict of interest
Cory D. Sago is co-founder of Guide Therapeutics and an employee at Guide Therapeutics. James E. Dahlman is a co-founder of Guide Therapeutics and consultant for Guide Therapeutics. Brandon R. Krupczak is an employee at Guide Therapeutics. Melissa P. Lokugamage declares no conflict of interest. Zubao Gan declares no conflict of interest.
Ethical Approval
All animal studies were carried out in accordance with the institutional and national guidelines, following an animal protocol approved by the Georgia Institute of Technology IACUC committee. No human studies were performed as part of this research.
Funding
C.D.S. and J.E.D. were funded by Georgia Tech startup funds (awarded to J.E.D.). C.D.S. was funded by the NIH-sponsored Research Training Program in Immunoengineering (T32EB021962). M.P.L was funded by the NIH-sponsored Research Training Program in Computational Biology and Predictive Health Genomics (T32GM105490). This content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
C.D.S. and J.E.D. designed experiments, performed experiments, and analyzed data. B.Z.K., M.P.L., Z.G. performed experiments. C.D.S. and J.E.D. wrote the paper, which was reviewed by all other authors.
Corresponding author
Additional information
Associate Editor Michael R. King oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
James E. Dahlman is an Assistant Professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory. His lab works at the interface of drug delivery, nanotechnology, genomics, and gene editing. James has designed nanoparticles that deliver RNA to blood vessels in the heart and lung; these nanoparticles have been validated by > 20 labs and have been licensed for clinical development. James also uses molecular biology to design the genetic drugs he delivers. He designed ‘dead’ guide RNAs to turn on genes using active Cas9. Similarly, using his background in nanoparticle chemistry, in vivo RNA delivery, and genomics, his lab has designed a series of increasingly sensitive DNA barcoding systems that can measure how > 200 nanoparticles target cells 30 different cell types at once, directly in vivo. James’ nanoparticle barcoding work led to his placement on the MIT Tech Review TR35 list. James has won scientific awards at every stage of his career, including the NSF, NDSEG, NIH OxCam, Whitaker, and LSRF Fellowships, and the Weintraub Graduate Thesis Award. He has been named a young / leading investigator by Bayer, the Parkinson’s Disease Foundation, and the Journal of Materials Chemistry B. At the age of 32, his research has been published in Science, Cell, Nature Nanotechnology, Nature Biotechnology, Nature Cell Bio, Science Translational Medicine, PNAS, Advanced Materials, JACS and other leading journals. He has given > 75 invited talks on drug delivery, gene editing, and nanoparticle DNA barcoding across the world, and is a co-founder of GuideRx.

This article is part of the 2019 CMBE Young Innovators special issue.
Rights and permissions
About this article
Cite this article
Sago, C.D., Krupczak, B.R., Lokugamage, M.P. et al. Cell Subtypes Within the Liver Microenvironment Differentially Interact with Lipid Nanoparticles. Cel. Mol. Bioeng. 12, 389–397 (2019). https://doi.org/10.1007/s12195-019-00573-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12195-019-00573-4