Abstract
A method combining multiplex real-time polymerase chain reaction (PCR) with high-resolution melting (HRM) analysis for rapid and specific simultaneous detection of Salmonella, Listeria monocytogenes, and Staphylococcus aureus was developed. The method included a melting-curve analysis of products and was evaluated by specificity, sensitivity and reproducibility analyses. Sensitivity and reproducibility analyses was both conducted by genomic DNA extracted from serial dilutions for each target pathogen. Assays with artificially inoculated and naturally contaminated samples after enrichment were also conducted. In the specificity test, there was no nonspecific amplification of the 44 nontarget pathogens, whereas the actual T m values were 79.38 ± 0.14, 82.54 ± 0.15, and 77.36 ± 0.14 °C for Salmonella, L. monocytogenes, and S. aureus, respectively. The sensitivity of the method was 3.5 × 102 CFU ml−1 for Salmonella and L. monocytogenes and 3.5 × 103 CFU ml−1 for S. aureus. The coefficients of variation of T m values ranged 0.51–1.03 % for Salmonella, 1.63–2.11 % for L. monocytogenes, and 0.75–2.17 % for S. aureus in intraassay, and ranged 0.81–2.43 % for Salmonella, 1.97–2.35 % for L. monocytogenes, and 0.93–3.93 % for S. aureus in interassay. The detection limit in artificially inoculated samples (n = 50) was 5 CFU (25 g)−1 food for the three tested pathogens. In the naturally contaminated samples (n = 120),Salmonella DNA was detected by HRM, sequencing, and conventional culture-based methods at a positive rate of 25.00, 25.00, and 24.17 %, respectively; the corresponding rates for L. monocytogenes were 14.17, 14.17, and 14.17 %, respectively, while those for S. aureus were 16.7, 16.7, and 16.7 %, respectively.
Similar content being viewed by others
Introduction
Foodborne diseases are widespread public health concerns affecting both developed and developing countries. Among the foodborne pathogens, Salmonella enterica, Listeria monocytogenes, and Staphylococcus aureus are recognized as the major causes of food contaminations (Kim and Bhunia 2008), as well as the pathogens that predominantly cause the foodborne human diseases in the world. Thus, rapid and specific methods to detect these pathogens are needed.
Currently, the most used method for identification of foodborne pathogens from food products is still cultivation based conventional diagnostics methods, which generally require bacterial culture and subsequent biochemical assays. The procedures of conventional diagnostics methods are labor-intensive, time-consuming, and expensive due to the necessity of separate cultivation for each target species (Xu et al. 2012). As a replacement for conventional microbiological approaches, real-time polymerase chain reaction (PCR) possesses sensitivity equal to culture methods, but it is rapid and allow testing to be completed in 48 h (Navas et al. 2006). In addition, contrary to the conventional PCR, it eliminates the risk of cross-contamination from gel electrophoresis, reducing both the workload and analysis time (Omiccioli et al. 2009). Protocols of real-time PCR for simplex detection of S. enterica, L. monocytogen, and S. aureus have been employed (Almeida et al. 2013; Vanegas et al. 2009; Fusco et al. 2011). However, because many food, such as milk, dairy products, meat, poultry, fruits, and vegetables, are commonly contaminated with carriers of multiple pathogens, such as S. enterica, Escherichia coli O157:H7, S. aureus, and L monocytogenes a simultaneous, rapid, and accurate multiplex detection of these foodborne pathogens becomes important, which will make the task of food surveillance much more manageable and will be of significance for tracing the outbreaks of bacterial pathogens within the food supply (13). In addition, the multiplex PCR technique is attractive and economically favorable, because it can reduce the total space requirement for handling a large number of samples, as well as the bench space, supplies, reagents, and labor needed, hence reducing the overall cost of testing per pathogen (Ruiz-Rueda et al. 2011; Singh et al. 2012; Elizaquivel and Aznar 2008). Multiplex real-time PCR methods for detecting S. enterica, L. monocytogenes, and S. aureus have been reported (Chen et al. 2013; Chiang et al. 2012; Garrido et al. 2013). It is clear that these pathogens continue to be a health hazard and a serious problem for the food industry and need to be detected with fast and reliable methods.
In recent years, real-time PCR based on a variety of chemistries and detection instruments have become feasible with high-throughput technologies (Elizaquível and Aznar 2008). For example, detection systems that combined real-time PCR with high-resolution melt analysis have been introduced to the field of clinical diagnostics (Chen et al. 2013). High-resolution melting (HRM) analysis is a newly emerging technology that is able to monitor the separation of double-stranded (ds) DNA with increasing temperatures (Tong et al. 2010). HRM is based on computer analysis of DNA melting transitions, whereby it is possible to record more than 25 readings per 1 °C, via monitoring of the changes in fluorescence resulted from gradual temperature-dependent release of an intercalated fluorescent ds DNA dye (Norambuena et al. 2009). The advantages of HRM real-time PCR include the use of high-quality DNA intercalating dyes and the step by step monitoring of fluorescence over small temperature increments with high resolution and accuracy. In this closed tube technique, there is neither post-PCR handling nor gel separation steps, there is less chance for contamination, and the general analysis time is reduced (Pangasa et al. 2009; Pietzka et al. 2011). Extremely precise melting temperature discrimination power as well as the advantages of easy, rapid, high-throughput, and low-cost operation (Krypuy et al. 2007; Reed et al. 2007) make the application of HRM being applied widely for coping with heterozygous sequences and mutation, as well as for analysis of repetitive sequence (Chateigner-Boutin and Small 2007; Fortini et al. 2007). Currently, HRM technique has been rapidly introduced into diagnostic laboratories to genotype disease-associated genes as well as to differentiate mutation at single nucleotide scale, e.g., the gyrA mutations causing quinolone resistance in S. enterica serovars typhi and paratyphi (Slinger et al. 2007). It is applied not only in the rapid identification and simultaneous subtyping of bacteria, such as L. monocytogenes (Pietzka et al. 2011), and diagnosis of protozoans, such as cryptosporidiosis in humans (Pangasa et al. 2009) with one primer pair, but also in the identification and genotyping of bacteria, such as S. aureus (Lilliebridge et al. 2011) and Salmonella serovars (Chateigner-Boutin and Small 2007; Vanegas et al. 2009), and duplex detection of Macrobrachium rosenbergii nodavirus (MrNV) and extra small virus (XSV) in shrimp (Senapin et al. 2010) with several primer pairs, as well as in the classification of virus such as fowl adenovirus serotypes (Steer et al. 2009) and dengue (Naze et al. 2009).
Because each pathogen specific sequence has a precise theoretical T m value, the HRM analysis technique can be applied to multiplex detection of several pathogens. Therefore, for the first time, the present study has developed a multiplex HRM real-time PCR system for simultaneous detection of Salmonella, L. monocytogenes, and S. aureus and optimized the assay using a Rotor gene 6000 platform. The sensitivity, specificity, and reproducibility of the method were also assessed, and the application of the assay with artificially inoculated and naturally contaminated samples were evaluated.
Materials and Methods
Bacterial Strains and Culture Conditions
There were 90 strains of Salmonella, L. monocytogenes, and S. aureus used as target in this study, while another 44 nontarget pathogens were selected to assess the specificity of the assay because of their relevance to foodborne diseases (Table 1). All strains were maintained at −80 °C in nutrient broth (NB) supplemented with 20 % glycerol for long-term storage.
DNA Extraction and Primer Design
Pathogen DNA was extracted using a DNeasy tissue kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions and stored at −20 °C until use. Presumptively conserved sequences of Salmonella, L. monocytogenes, and S. aureus were used to design primers with Primer Express V2.0. All primer combinations were evaluated for the formation of primer–dimer structures and to discard putative interactions among the primers. The feasibility of all primers was subsequently validated by BLAST (http://www.ncbi.nlm.nih.gov/BLAST). Finally, primers were synthesized by Invitrogen Biotechnology (Table 2).
Multiplex HRM Real-Time PCR
Each 25 μl PCR reaction system contained 2.5 μl 10× buffer, 0.5 μl Taq polymerase (5 U μl−1), 2.5 μl dNTP (10 mmol l−1), 3 μl MgCl2 (25 m mol l−1), 0.5 μl each of the forward and reverse primers (10 μmol l−1), 2 μl DNA template of each pathogen (0.5 μmol l−1), 1.25 μl of the EvaGreen (TAKARA, Japan), 0.2 μl of BSA (20 mg ml−1), and nuclease-free water to bring the total reaction volume to 25 μl. Nuclease-free water was used as the no template control (NTC).An optimized experimental protocol for the QIAGEN Rotor gene 6000 system (Qiagen, Hilden, Germany) was as follows: 95 °C for 10 min, 40 cycles at 95 °C for 10 s, and 60 °C for 45 s. Finally, a melting curve temperature profile was undertaken from 60 to 95 °C. Four different melt rates including 0.1, 0.2, 0.4, and 0.8 °C s−1 were performed to obtain the best resolution of the melt curve. EvaGreen was used as the intercalating dye, and the data were analyzed using the manufacture’s software. Some research found that a complete translocation of SYBR green I going from 74 to 114 bp fragment occurred when using a SmartCycler® II thermal cycler (Cepheid, Sunnyvale, CA, USA) with the common melt curve analysis function (Varga and James 2006). To address whether a machine with the HRM analysis function can avoid the phenomenon of dye translocation, repetitive melting curves of 15 times of melt runs were conducted by both the LightCycler 480 and QIAGEN Rotor gene 6000 system in duplicate, and the results showed that the LightCycler 480 machine was able to carry out the common melt curve analysis, while the QIAGEN Rotor gene 6000 system were able to perform the high-resolution melt curve analysis.
Sensitivity Assay
The analytic sensitivity of the multiplexed HRM real-time PCR assays was assessed and compared by testing genomic DNA extracted from tenfold serial dilutions of S. enterica serovar enteritidis, L. monocytogenes, and S. aureus ranging from 3.5 × 104 to 3.5 × 101 CFU ml−1 shown in Table 1, respectively. Cell concentrations were measured by standard surface spread plating count. At least three replicates at each genomic DNA concentration were assessed, and a detection level was considered “positive” only if all three replicates returned positive results.
Determination of Intraassay and Interassay Reproducibility
Seven dilutions of genomic DNA extracted from 107 to 101 CFU ml−1 for each of the three target pathogens described in sensitivity assay were assayed in triplicate. Genomic DNA was amplified four times in the same run to evaluate intraexperimental reproducibility and in four separate runs that were performed on different days to evaluate interassay reproducibility. The mean, standard deviation (S.D.), and coefficient of variation (CV, standard deviation/Ct mean.) for T m value obtained for each dilution were calculated.
Application of the Multiplex HRM Real-Time PCR System to Artificially Inoculated Samples
There were 50 food samples including milk, chicken, beef, pork, and eggs collected from the Dongguan Entry–Exit Inspection and Quarantine Bureau, Donguan, China, and proven to have no target pathogens with GB 4789-2008 (GB is the National Standard of the People’s Republic of China), kept at –20 °C except for milk and eggs at 4 °C before analysis. Each 25 g food samples were spiked with S. enterica serovar enteritidis, L. monocytogenes, and S. aureus cells with 5, 10, and 20 CFU (25 g)−1 food homogenate as final concentration of each species, measured by standard surface spread plating count and stomached at 20 °C for 1 h. Then the samples were mixed with 225 ml of SSL (selective enrichment broth, which allowed the simultaneous growth of the three target pathogens) described in our previous study (Yu et al. 2010) and homogenized for 10 s. Uninoculated meat samples served as negative controls and DNA referred in sensitivity assay was used as positive control. After 24 h of incubation at 37 °C, samples were centrifuged for 5 min at 1,000 × g and filtered in a Stomacher bag (Tekmar, Cincinnati, OH) to remove coarse food particles, DNA was extracted using a DNeasy Tissue kit (Qiagen, Valencia, CA, USA) according to manufacturer’s protocol. There were 50 samples with three artificially inoculated concentration tested by real-time PCR assays with HRM analysis in triplicate.
Naturally Contaminated Samples
To further confirm the robustness of this method, we assessed naturally contaminated pork samples. A total of 120 pork samples were collected from the Dongguan Entry–Exit Inspection and Quarantine Bureau, Donguan, China, kept at −20 °C before analysis. These samples were treated by the method described in the assay of artificially inoculated samples without the artificially inoculation process. DNA was extracted using a DNeasy tissue kit (Qiagen, Valencia, CA, USA) according to manufacturer’s protocol. Samples were enriched with SSL then subjected to the HRM real-time PCR analysis, PCR amplicons were further analyzed by cloning into pUC-19 T plasmid and subjected to sequencing by Shanghai Sangong (Shanghai, China). The same panel of samples was also synchronously tested by conventional culture-based detection method according to GB 4789-2008 China. Genomic DNA of Salmonella, L. monocytogenes, and S. aureus was used as the Positive controls and nuclease-free water was used as the negative controls.
Results
Optimization of Real-Time PCR with HRM Analysis
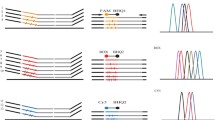
Common melt curves with 15 melt runs conducted by LightCycler 480 without HRM analysis function are shown in Fig. 1. The results of our studies showed that template concentration affected the shape of the melting curve. A higher template concentration of Salmonella led to the competitive inhibition to L. monocytogenes and S. aureus; thus, only the peak of Salmonella was recognized in the melting curves in such condition (Fig. 1a 1 and b1). Interestingly, a translocation of the EvaGreen dye, a saturable dye, from one amplicon to another was observed when the melt runs were repeated more than four times during a time-course of melt analysis in three samples with different template concentrations (Fig. 1a 1–a15, b1–b15, c1–c15). In repeated melt run treatments (total of 15) of a single sample containing coamplified targets, partial translocation of EvaGreen was observed, going from a 117 bp fragment (Salmonella) to a 86 bp fragment (L. monocytogenes), and a 147 bp fragment (S. aureus). This phenomenon is similar to the results in other research (Varga and James 2006), which found a complete translocation of SYBR green I, an unsaturated dye, going from a 74 bp fragment to a 114 bp fragment. However, the phenomenon of translocation can be avoided by Rotor gene 6000 with HRM analysis function, that is why we chose the HRM analysis instead of the common melt curve analysis for the detection system.
Translocation of EvaGreen dye in repetitive melting curve of HRM multiplex real-time PCR assay on LightCycler 480. Samples a–c represent template concentration ratios of Staphylococcus aureus : Salmonella : Listeria monocytogenes were 0.2 : 1.0 : 0.2, 0.2 : 1.0 : 0.4, and 0.5 : 0.5 : 0.1, respectively, in duplicate. Subscripts indicate the melt run number
As for the melt rate, we found that the shift in T m value from low to high is associated with a low to high shift in melt rate. In addition, lower melt rate improves the resolution of melt curve. Templates with concentration ratios of S. aureus : Salmonella : L. monocytogenes as 1.0 : 1.0 : 0.4 were subjected to the HRM assay with the same results observed (Fig. 2). To gain a better resolution, four melt runs at the melt rates of 0.1, 0.2, 0.4, and 0.8 °C s−1, respectively, were conducted on Rotor gene 6000. A lower melt rate increases the number of readings over time; therefore, the loss of fluorescence is smaller than that at a higher melt rate. In addition, a melt rate of 0.1 °C s−1 is more sensitive to changes in fluorescence than a rate at 0.8 °C s−1, as eight times more fluorescence readings are taken, especially when two amplicons melt at similar temperatures. However, as the arrow shown in Fig. 2a 1, when the melt rate was 0.1 °C s−1, the resolution was too high and formed two peaks for L. monocytogenes, which makes it difficult to distinguish several pathogens in a multiplex detection system. In the templates with concentration ratios of 1.0 : 1.0 : 0.4 for S. aureus : Salmonella : L. monocytogenes, larger and more discernible melt peaks were associated with the 0.2 °C s−1 melt rate at T m values of 77.29, 79.11, and 82.64 °C for the 147, 117, and 86 bp amplicons, respectively. However, the loss of peak resolution occurred with a ramp rate of 0.4 and 0.8 °C s−1 when the T m values of two amplicons were close. A trade-off exists where at a lower melt rate T m accuracy with a lower associated S.D. is improved but at the expense of potentially missing the melt peak of weak amplicons.
Melt curves of different melt rates in HRM real-time PCR on Rotor gene 6000. Template concentration ratios of Staphylococcus aureus : Salmonella : Listeria monocytogenes of the sample is 1.0 : 1.0 : 0.4. a 1 –a 4 represent the melt rate as 0.1, 0.2, 0.4, and 0.8 °C s−1, respectively. The arrows showed a relative higher resolution of L. monocytogenes at 0.1 °C s−1.
Specificity
The 90 Salmonella, L. monocytogenes, and S. aureus strains and 44 nontarget pathogen strains collected in this study were all detected, whereas no positive fluorescent signal was observed for nontarget pathogens strains. The results indicated the high specificity of the HRM multiplex real-time PCR assay for nontarget pathogens. The melting temperatures of the amplified products from target genes invA, hly, and Sa442 were recorded as 79.42 ± 0.14, 82.54 ± 0.15, and 77.56 ± 0.14 °C, respectively, which were 2 to 4 °C higher than the theoretical values calculated by DNAStar (DNAStar, USA) shown in Table 2. These melting temperatures were sufficiently different to distinguish three target amplicons.
Analytical Sensitivity
The analytic sensitivity of the assay was determined to be 3.5 × 102 CFU ml−1 for Salmonella and L. monocytogenes DNA and 3.5 × 103 CFU ml−1 for S. aureus DNA by HRM real-time PCR.
Reproducibility
Intraassay and interassay reproducibilities of HRM real-time PCR for Salmonella, L. monocytogenes, and S. aureus were assessed. The CVs of T m values for seven dilutions (107–101 CFU ml−1) were calculated, and the results showed that the assay was highly reproducible. While the CVs ranged from 0.51 to 1.03 % for Salmonella, from 1.63 to 2.11 % for L. monocytogenes, and from 0.75 to 2.17 % for S. aureus in intraassay, they ranged from 0.81 to 2.43 % for Salmonella, from 1.97 to 2.35 % for L. monocytogenes, and from 0.93 to 3.93 % for S. aureus in interassay. Such low CVs of both intraassay and interassay indicated a high repeatability of the developed assay and confirmed its sensitivity to 3.5 × 102 CFU ml−1 for Salmonella and L. monocytogenes and 3.5 × 103 CFU ml−1 for S. aureus.
Artificially Inoculated Samples
To analyze how different samples affected the HRM real-time PCR assay, the assay was used to detect DNA of three target pathogens extracted from milk, chicken, beef, pork, and eggs. A total of 50 artificially inoculated samples were analyzed (Table 3). Under three inoculated concentrations including 5, 10, and 20 CFU (25 g)−1, T m values of the amplicon from DNA extracted from food samples were not significantly different from those obtained directly from the bacterial culture, which may be due to the high purity of the extracted DNA, because impurity in samples may affects the T m values. Multiplex HRM real-time PCR assays after 24 h of enrichment at 37 °C showed a detection limit of 5 CFU (25 g)−1 food for all three target pathogens, which is similar to that achieved in a reported multiplex detection of Salmonella spp. and L. monocytogenes (5 CFU (25 g)−1; Garrido et al. 2013) with incubation at 37 °C overnight, but higher than another multiplex detection of S. enterica serovar typhimurium, L. monocytogenes, and E. coli O157:H7 in liquid whole egg (ten cells (25 g)−1; Germini et al. 2009) with incubation at 37 °C for 15 h.
Naturally Contaminated Samples
As shown in Table 4, Salmonella was detected in specimens of 120 porks by HRM real-time PCR, sequencing method, and conventional culture-based detection method with a positive rate of 25.00, 25.00, and 24.17 %, respectively; the positive rate for L. monocytogenes was 14.17, 14.17, and 14.17 %, respectively, and for S. aureus 16.7, 16.7, and 16.7 %, respectively. Of these samples, 30 samples were positive for Salmonella based on HRM real-time PCR assay, and 29 samples were positive based on conventional culture method. The 17 samples were positive for L. monocytogenes both according to HRM real-time PCR assay and conventional culture method, while the 20 samples were positive for S. aureus based both on HRM real-time PCR assay and the conventional method. The HRM real-time PCR-positive strain, which was negative according to the conventional culture method, was also analyzed by cloning the amplification fragments into pUC-19 T plasmid, and the sequencing result confirmed the accuracy of the multiplex HRM real-time PCR assay. Among 120 samples, three samples were found positive for Salmonella and L. monocytogenes, two samples were found positive for Salmonella and S. aureus by HRM real-time PCR and conventional method, verified by amplicon sequencing. No sample was found positive for the three target pathogens simultaneously. No false-negative or false-positive samples occurred using the developed multiplex HRM real-time PCR method, demonstrating that the newly developed HRM real-time PCR assay was more sensitive than the conventional method.
Discussion
In our study, a multiplex real-time PCR system combining with HRM analysis for the detection of Salmonella, L. monocytogenes, and S. aureus was developed using three specific pairs of primers targeting the genes invA, hly, and Sa442, respectively. Factors affected the T m values were also discussed to make sure the rationality of the criteria of T m values.
Many factors, such as size and G/C content of the amplified target fragment, machine, and parameters like melt rate, may affect the T m values. Theoretically, the T m value of each product is dependent on the length and G/C content of the sequence (Bratchikov and Mauricas 2011); a positive correlation between T m value and G/C content of the amplified DNA fragment can be observed in Table 2. Among these three DNA fragments, L. monocytogenes has the highest theoretical T m value of 78.63 °C and highest G/C content of 47.67 %. Under a similar length, DNA fragments with an increasing percentage of A/T are easier to dissociate into single strand. In addition, choosing conservative sequences of the pathogen DNA as the amplified fragment was given the utmost priority in our study, which directly affects the reliability of T m value.
Furthermore, the machine’s optics as well as software interpretation of fluorescence data also have an important impact on the T m value. In the previous research, real-time PCR assays with melt curve analysis were conducted by two styles of machine, i.e., LightCycler 480 and Rotor gene 6000. The detection result turned out that when using the LightCycler 480 without HRM module, a translocation of EvaGreen dye from one amplicon to another was observed when the melt run was repeated more than four times. This phenomenon may explain inconsistent EvaGreen fluorescence patterns associated with melt curve analysis of some amplicon complexes. The duration of the melt run may be a critical factor affecting EvaGreen binding and translocation, and its manipulation may facilitate improved resolution and simultaneous detection of multiple targets. Fortunately, such translocation phenomenon can be avoided when using Rotor gene 6000 with a HRM module, indicating the advantage of HRM on melt curve analysis. According to the results of this study, different machines with HRM module produced different T m values for the same target pathogen; however, once the machine is fixed, the T m value will not be significantly influenced when using a saturated dye (EvaGreen).
Some research have described that increasing the melt rate from 0.1 to 0.4 °C s−1 shifted the T m higher across all strains, to an average of approximately 1.08 °C, and the shift in T m was not constant across strains (Varga and James 2006). Although a similar result was gained in our experiment that T m value was positively correlated with melt rate (data not shown), once the optimal melt rate (0.2 °C s−1) was established, the T m value was not significantly affected when using the QIAGEN rotor gene 6000 with HRM module.
Five kinds of food including milk, chicken, beef, pork, and egg were chosen as the detection objects, and it was proven in this study that there was no significant effect of different sample sources on T m values, because of the proper DNA extraction process.
What is more, we found that when target template of Salmonella is in a concentration much higher than that of L. monocytogenes and S. aureus, only the peak of Salmonella is visible (Fig. 1a 1, b1, c1) by LightCycler 480 without the HRM module, suggesting possible false negative results because of competitive inhibition in amplification of multiple targets. After 15 HRM repletion, however, three obvious peaks could be observed, indicating that the lower concentration template was normally amplified. Fig. 2 was formed by QIAGEN Rotor gene 6000 system with HRM module, no translocation of EvaGreen dye was observed and templates with lower concentration was also amplified. Thus, it seems that there was no obvious competitive inhibition in the PCR amplification, and in contrast to Fig. 1, Fig. 2 showed the importance of the HRM module. Samples can effectively amplified at a similar concentrations as we can see in Figs. 2 and 1a 15, b15, c15 (after 15 HRM replicates). If the concentrations of templates vary significantly, for example, concentrations at 107 and 102 CFU ml−1 are amplified at the same time, it seems normal that high concentration will inhibit the lower ones in PCR. However, once template with higher concentration was detected, it means that the sample was contaminated, indicating that a food safety risk existed. At this moment, whether the templates with lower concentration could be amplified will not be urgent.
In spite of all the considerations taken above, there are still certain problems, e.g., how to validate the criteria of T m value, that remain to be solved. In some studies which rapidly identified and simultaneously subtyped bacteria, cutoff values were used as criteria, for example, T m cutoff value (Slinger et al. 2007), cycle threshold cutoff value (Tong et al. 2010), and relative fluorescence unit (RFU) cutoffs (Chen et al. 2013) were established to indicate the detection of Salmonella serovars and S. aureus and sample classification, respectively. The need of cutoff value was based on the very small differences in T m value, cycle threshold, and RFU due to the precise requirement of subtyping. However, it may not be necessary for our study, since the differences in T m value among Salmonella, L. monocytogenes, and S. aureus were great enough, which makes the design of primer pairs much more convenient and the differentiation between pathogens much more intuitive and reliable, avoiding the tedious calculation.
To evaluate the reliability of this assay, artificially inoculated samples as well as naturally contaminated samples were detected in our study. In consideration of the high specificity, good sensitivity (3.5 × 102 CFU ml−1 for Salmonella and L. monocytogenes and 3.5 × 103 CFU ml−1 for S. aureus), high reproducibility (0.85 ± 0.11 % for Salmonella, 1.85 ± 1.12 % for L. monocytogenes, and 1.21 ± 0.87 % for S. aureus in intraassay; 2.12 ± 1.61 % for Salmonella, 2.19 ± 2.02 % for L. monocytogenes, and 1.59 ± 1.52 % for S. aureus in interassay), high detection limit for artificially inoculated samples (5 CFU (25 g)−1 food for all three pathogens with enrichment), and a more accurate positive rate of 120 naturally contaminated pork samples than the conventional method, the HRM real-time PCR system in this study is feasible and reliable.
Finally, as a PCR-based method, an internal amplification control (IAC) should be usually used to determine or eliminate false negatives. We once included the internal amplification controls in this multiple detection system, but it led to more competitive inhibition due to the primers of the IAC. As we see in Figs. 1 and 2, though samples at lower concentration got amplification, there might be still part of competitive inhibition. However, we did a series of experiments to study how different food samples affect the T m values; as the Table 3 showed, no significant inhibition of the PCR was observed within the food panel. Of course, we may select different IAC to further improve our detection method in the future, if necessary.
Conclusions
In summary, this study has developed and utilized a low-cost, closed-tube, easily estimated, and easily designed HRM-based multiplex real-time PCR assay for rapid, specific, and sensitive detection of Salmonella, L. monocytogenes, and S. aureus. It can also be applied to the detection of artificially inoculated samples and naturally contaminated samples. The described method allows the detection of three different pathogens in a single run, from a single extract using HRM analysis in the same QIAGEN Rotor gene 6000 apparatus. This method has a potential for clinical diagnosis and epidemiological surveillance.
References
Almeida C, Cerqueira L, Azevedo N, Vieira M (2013) Detection of Salmonella enterica serovar enteritidis using real time PCR, immunocapture assay, PNA FISH and standard culture methods in different types of food samples. Int J Food Microbiol 161(1):16–22
Bratchikov M, Mauricas M (2011) Development of a multiple-run high-resolution melting assay for Salmonella spp. genotyping: HRM application for Salmonella spp. subtyping. Diagn Microbiol Infect Dis 71(3):192–200
Chateigner-Boutin A-L, Small I (2007) A rapid high-throughput method for the detection and quantification of RNA editing based on high-resolution melting of amplicons. Nucleic Acids Res 35(17):e114
Chen JH-K, Cheng VC-C, Chan JF-W, She KK-K, Yan M-K, Yau MC-Y, Kwan GS-W, Yam W-C, Yuen K-Y (2013) The use of high-resolution melting analysis for rapid spa typing on methicillin-resistant Staphylococcus aureus clinical isolates. J Microbiol Methods 92(2):99–102
Chiang Y-C, Tsen H-Y, Chen H-Y, Chang Y-H, Lin C-K, Chen C-Y, Pai W-Y (2012) Multiplex PCR and a chromogenic DNA macroarray for the detection of Listeria monocytogens, Staphylococcus aureus, Streptococcus agalactiae, Enterobacter sakazakii, Escherichia coli O157: H7, Vibrio parahaemolyticus, Salmonella spp. and Pseudomonas fluorescens in milk and meat samples. J Microbiol Methods 88(1):110–116
Elizaquível P, Aznar R (2008) A multiplex RTi-PCR reaction for simultaneous detection of Escherichia coli O157: H7, Salmonella spp. and Staphylococcus aureus on fresh, minimally processed vegetables. Food Microbiol 25(5):705–713
Elizaquivel P, Aznar R (2008) A multiplex RTi-PCR reaction for simultaneous detection of Escherichia coli O157: H7, Salmonella spp. and Staphylococcus aureus on fresh, minimally processed vegetables. Food Microbiol 25(5):705–713
Fortini D, Ciammaruconi A, De Santis R, Fasanella A, Battisti A, D’Amelio R, Lista F, Cassone A, Carattoli A (2007) Optimization of high-resolution melting analysis for low-cost and rapid screening of allelic variants of Bacillus anthracis by multiple-locus variable-number tandem repeat analysis. Clin Chem 53(7):1377–1380
Fusco V, Quero GM, Morea M, Blaiotta G, Visconti A (2011) Rapid and reliable identification of Staphylococcus aureus harbouring the enterotoxin gene cluster (egc) and quantitative detection in raw milk by real time PCR. Int J Food Microbiol 144(3):528–537
Garrido A, Chapela M-J, Román B, Fajardo P, Lago J, Vieites JM, Cabado AG (2013) A new multiplex real-time PCR developed method for Salmonella spp. and Listeria monocytogenes detection in food and environmental samples. Food Control 30(1):76–85
Germini A, Masola A, Carnevali P, Marchelli R (2009) Simultaneous detection of Escherichia coli O175: H7, Salmonella spp., and Listeria monocytogenes by multiplex PCR. Food Control 20(8):733–738
Kim H, Bhunia AK (2008) SEL, a selective enrichment broth for simultaneous growth of Salmonella enterica, Escherichia coli O157: H7, and Listeria monocytogenes. Appl Environ Microbiol 74(15):4853–4866
Krypuy M, Ahmed AA, Etemadmoghadam D, Hyland SJ, Fox SB, Brenton JD, Bowtell DD, Dobrovic A (2007) High resolution melting for mutation scanning of TP53 exons 5–8. BMC Cancer 7(1):168
Lilliebridge RA, Tong SY, Giffard PM, Holt DC (2011) The utility of high-resolution melting analysis of SNP nucleated PCR amplicons—an MLST based Staphylococcus aureus typing scheme. PLoS One 6(6):e19749
Navas J, Ortiz S, Lopez P, Jantzen MM, Lopez V, Martinez-Suarez JV (2006) Evaluation of effects of primary and secondary enrichment for the detection of Listeria monocytogenes by real-time PCR in retail ground chicken meat. Foodbourne Pathog Dis 3(4):347–354
Naze F, Le Roux K, Schuffenecker I, Zeller H, Staikowsky F, Grivard P, Michault A, Laurent P (2009) Simultaneous detection and quantitation of Chikungunya, dengue and West Nile viruses by multiplex RT-PCR assays and dengue virus typing using high resolution melting. J Virol Methods 162(1):1–7
Norambuena PA, Copeland JA, Křenková P, Štambergová A, Macek M Jr (2009) Diagnostic method validation: high resolution melting (HRM) of small amplicons genotyping for the most common variants in the MTHFR gene. Clin Biochem 42(12):1308–1316
Omiccioli E, Amagliani G, Brandi G, Magnani M (2009) A new platform for real-time PCR detection of Salmonella spp., Listeria monocytogenes and Escherichia coli O157 in milk. Food Microbiol 26(6):615–622
Pangasa A, Jex AR, Campbell BE, Bott NJ, Whipp M, Hogg G, Stevens MA, Gasser RB (2009) High resolution melting-curve (HRM) analysis for the diagnosis of cryptosporidiosis in humans. Mol Cell Probes 23(1):10–15
Pietzka AT, Stöger A, Huhulescu S, Allerberger F, Ruppitsch W (2011) Gene scanning of an internalin B gene fragment using high-resolution melting curve analysis as a tool for rapid typing of Listeria monocytogenes. J Mol Diagn 13(1):57–63
Reed GH, Kent JO, Wittwer CT (2007) High-resolution DNA melting analysis for simple and efficient molecular diagnostics. Pharmacogenomics 8(6):597–608
Ruiz-Rueda O, Soler M, Calvó L, García-Gil JL (2011) Multiplex real-time PCR for the simultaneous detection of Salmonella spp. and Listeria monocytogenes in food samples. Food Anal Methods 4(2):131–138
Senapin S, Molthathong S, Phiwsaiya K, Jaengsanong C, Chuchird N (2010) Application of high resolution melt (HRM) analysis for duplex detection of Macrobrachium rosenbergii nodavirus (MrNV) and extra small virus (XSV) in shrimp. Mol Cell Probes 24(5):291–297
Singh J, Batish VK, Grover S (2012) Simultaneous detection of Listeria monocytogenes and Salmonella spp. in dairy products using real time PCR-melt curve analysis. J Food Sci Technol 49(2):234–239
Slinger R, Bellfoy D, Desjardins M, Chan F (2007) High-resolution melting assay for the detection of gyrA mutations causing quinolone resistance in Salmonella enterica serovars typhi and paratyphi. Diagn Microbiol Infect Dis 57(4):455–458
Steer PA, Kirkpatrick NC, O’Rourke D, Noormohammadi AH (2009) Classification of fowl adenovirus serotypes by use of high-resolution melting-curve analysis of the hexon gene region. J Clin Microbiol 47(2):311–321
Tong SY, Lilliebridge RA, Holt DC, Coombs GW, Currie BJ, Giffard PM (2010) Rapid detection of H and R Panton–Valentine leukocidin isoforms in Staphylococcus aureus by high-resolution melting analysis. Diagn Microbiol Infect Dis 67(4):399–401
Vanegas MC, Vásquez E, Martinez AJ, Rueda AM (2009) Detection of Listeria monocytogenes in raw whole milk for human consumption in Colombia by real-time PCR. Food Control 20(4):430–432
Varga A, James D (2006) Real-time RT-PCR and SYBR Green I melting curve analysis for the identification of Plum pox virus strains C, EA, and W: effect of amplicon size, melt rate, and dye translocation. J Virol Methods 132(1):146–153
Xu Y, Cui L, Tian C, Li S, Cao J, Liu Z, Zhang G (2012) A multiplex polymerase chain reaction coupled with high-performance liquid chromatography assay for simultaneous detection of six foodborne pathogens. Food Control 25(2):778–783
Yu YG, Wu H, Liu YY, Li SL, Yang XQ, Xiao XL (2010) A multipathogen selective enrichment broth for simultaneous growth of Salmonella enterica serovar enteritidis, Staphylococcus aureus, and Listeria monocytogenes. Can J Microbiol 56(7):585–597
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Nos. 31101279 and 31271867), by the Doctoral Program Foundation of Institutions of Higher Education of China (No. 20110172120034), and by the Fundamental Research Funds for the Central Universities (Nos. 2013ZZ0068 and 2013ZZ0077).
Conflict of Interest
Xing-long Xiao has no conflict of interest. Li Zhang has no conflict of interest. Hui Wu has no conflict of interest. Yi-gang Yu has no conflict of interest. Yu-qian Tang has no conflict of interest. Dong-mei Liu has no conflict of interest. Xiao-feng Li has no conflict of interest. This article does not contain any studies with human or animal subjects.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xiao, Xl., Zhang, L., Wu, H. et al. Simultaneous Detection of Salmonella, Listeria monocytogenes, and Staphylococcus aureus by Multiplex Real-Time PCR Assays Using High-Resolution Melting. Food Anal. Methods 7, 1960–1972 (2014). https://doi.org/10.1007/s12161-014-9875-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-014-9875-x