Abstract
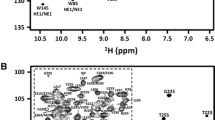
Onconase® FL-G zymogen is a 120 residue protein produced by circular permutation of the native Onconase® sequence. In this construction, the wild type N- and C-termini are linked by a 16 residue segment and new N- and C-termini are generated at wild type positions R73 and S72. This novel segment linking the native N- and C-termini is designed to obstruct Onconase’s® active site and encloses a cleavage site for the HIV-1 protease. As a first step towards the resolution of its 3D structure and the study of its structure–function relationships, we report here the nearly complete NMR 1H, 13C and 15N resonance chemical shift assignments at pH 5.2 and 35°C (BMRB deposit no 17973). The results presented here clearly show that the structure of the wild type Onconase® is conserved in the FL-G zymogen.


Similar content being viewed by others
References
Beck ZQ, Hervio L, Dawson PE, Elder JH, Madison EL (2000) Identification of efficiently cleaved substrates for HIV-1 protease using a phage display library and use in inhibitor development. Virology 274(2):391–401. doi:10.1006/viro.2000.0420
Darzynkiewicz Z, Carter SP, Mikulski SM, Ardelt WJ, Shogen K (1988) Cytostatic and cytotoxic effects of Pannon (P-30 Protein), a novel anticancer agent. Cell Tissue Kinet 21(3):169–182
Goddard TD, Kneller DG (2005) Sparky 3. University of California, San Francisco
Gorbatyuk VY, Tsai CK, Chang CF, Huang TH (2004) Effect of N-terminal and Met23 mutations on the structure and dynamics of onconase. J Biol Chem 279(7):5772–5780. doi:10.1074/jbc.M311233200
Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wüthrich K (1998) Recommendations for the presentation of NMR structures of proteins and nucleic acids. Pure Appl Chem 70(1):117–142
Plainkum P, Fuchs SM, Wiyakrutta S, Raines RT (2003) Creation of a zymogen. Nat Struct Biol 10(2):115–119. doi:10.1038/nsb884
Ribó M, Bosch M, Torrent G, Benito A, Beaumelle B, Vilanova M (2004) Quantitative analysis, using MALDI-TOF mass spectrometry, of the N-terminal hydrolysis and cyclization reactions of the activation process of onconase. Eur J Biochem 271(6):1163–1171
Schulenburg C, Low C, Weininger U, Mrestani-Klaus C, Hofmann H, Balbach J, Ulbrich-Hofmann R, Arnold U (2009) The folding pathway of onconase is directed by a conserved intermediate. Biochemistry 48(35):8449–8457. doi:10.1021/bi900596j
Wishart DS, Bigam CG, Holm A, Hodges RS, Sykes BD (1995a) 1H, 13C and 15N random coil NMR chemical shifts of the common amino acids. I. Investigations of nearest-neighbor effects. J Biomol NMR 5(1):67–81
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995b) 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J Biomol NMR 6(2):135–140
Acknowledgements
This work was supported by the projects, CTQ2008-0080, CTQ2010-21567-C02-02 and BFU2009-06935/BMC from MICINN and PUG2008A from the Universitat de Girona. MC acknowledges a fellowship from Ministerio de Educación y Ciencia.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Serrano, S., Callís, M., Vilanova, M. et al. 1H, 13C and 15N resonance assignments of the Onconase FL-G zymogen. Biomol NMR Assign 7, 13–15 (2013). https://doi.org/10.1007/s12104-012-9367-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-012-9367-0