Abstract
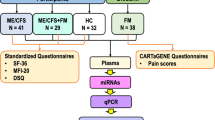
This work was aimed at investigating the circulating microRNA (miRNA) profiles in serum and saliva of patients affected by fibromyalgia syndrome (FM), correlating their expression values with clinical and clinimetric parameters and to suggest a mathematical model for the diagnosis of FM. A number of 14 FM patients and sex- and age-matched controls were enrolled in our study. The expression of a panel of 179 miRNAs was evaluated by qPCR. Statistical analyses were performed in order to obtain a mathematical linear model, which could be employed as a supporting tool in the diagnosis of FM. Bioinformatics analysis on miRNA targets were performed to obtain the relevant biological processes related to FM syndrome and to characterize in details the disease. Six miRNAs were found downregulated in FM patients compared to controls. Five of these miRNAs have been included in a linear predictive model that reached a very high sensitivity (100 %) and a high specificity (83.3 %). Moreover, miR-320b displayed a significant negative correlation (r = −0.608 and p = 0.036) with ZSDS score. Finally, several biological processes related to brain function/development and muscular functions were found potentially implicated in FM syndrome. Our study suggests that the study of circulating miRNA profiles coupled to statistical and bioinformatics analyses is a useful tool to better characterize the FM syndrome and to propose a preliminary model for its diagnosis.


Similar content being viewed by others
References
Kashikar-Zuck S, King C, Ting TV and Arnold LM (2016) Juvenile Fibromyalgia: Different from the Adult Chronic Pain Syndrome?. Curr. Rheumatol. Rep. 18, 19–016–0569-9
Buskila D, Press J, Gedalia A, Klein M, Neumann L, Boehm R, Sukenik S (1993) Assessment of nonarticular tenderness and prevalence of fibromyalgia in children. J Rheumatol 20:368–370
Gerloni V, Ghirardini M, Fantini F (1998) Assessment of nonarticular tenderness and prevalence of primary fibromyalgia syndrome in healthy Italian schoolchildren. Arthritis Rheum 41:1405
Wolfe F, Smythe HA, Yunus MB, Bennett RM, Bombardier C, Goldenberg DL, Tugwell P, Campbell SM et al (1990) The American College of Rheumatology 1990 criteria for the classification of fibromyalgia. Report of the Multicenter Criteria Committee Arthritis Rheum 33:160–172
Wolfe F, Clauw DJ, Fitzcharles MA, Goldenberg DL, Katz RS, Mease P, Russell AS, Russell IJ et al (2010) The American College of Rheumatology preliminary diagnostic criteria for fibromyalgia and measurement of symptom severity. Arthritis Care Res (Hoboken) 62:600–610
Sarzi-Puttini P, Atzeni F, Mease PJ (2011) Chronic widespread pain: from peripheral to central evolution. Best Pract Res Clin Rheumatol 25:133–139
Yunus MB (2008) Central sensitivity syndromes: a new paradigm and group nosology for fibromyalgia and overlapping conditions, and the related issue of disease versus illness. Semin Arthritis Rheum 37:339–352
Crofford LJ (2002) The hypothalamic-pituitary-adrenal axis in the pathogenesis of rheumatic diseases. Endocrinol Metab Clin N Am 31:1–13
Di Franco M, Iannuccelli C, Valesini G (2010) Neuroendocrine immunology of fibromyalgia. Ann N Y Acad Sci 1193:84–90
Rodriguez-Pinto I, Agmon-Levin N, Howard A, Shoenfeld Y (2014) Fibromyalgia and cytokines. Immunol Lett 161:200–203
Buskila D, Neumann L (1997) Fibromyalgia syndrome (FM) and nonarticular tenderness in relatives of patients with FM. J Rheumatol 24:941–944
Arnold LM, Hudson JI, Hess EV, Ware AE, Fritz DA, Auchenbach MB, Starck LO, Keck PE Jr (2004) Family study of fibromyalgia. Arthritis Rheum 50:944–952
Lee YH, Choi SJ, Ji JD, Song GG (2012) Candidate gene studies of fibromyalgia: a systematic review and meta-analysis. Rheumatol Int 32:417–426
Cortez MA, Calin GA (2009) MicroRNA identification in plasma and serum: a new tool to diagnose and monitor diseases. Expert Opin Biol Ther 9:703–711
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Tarang S, Weston MD (2014) Macros in microRNA target identification: a comparative analysis of in silico, in vitro, and in vivo approaches to microRNA target identification. RNA Biol 11:324–333
Chen Y., Gelfond J. A., McManus LM and Shireman PK (2009) Reproducibility of quantitative RT-PCR array in miRNA expression profiling and comparison with microarray analysis. BMC Genomics. 10, 407-2164-10-407
Bjersing JL, Lundborg C, Bokarewa MI, Mannerkorpi K (2013) Profile of cerebrospinal microRNAs in fibromyalgia. PLoS One 8:e78762
Bjersing JL, Bokarewa MI, Mannerkorpi K (2015) Profile of circulating microRNAs in fibromyalgia and their relation to symptom severity: an exploratory study. Rheumatol Int 35:635–642
de Boer HC, van Solingen C, Prins J, Duijs JM, Huisman MV, Rabelink TJ, van Zonneveld AJ (2013) Aspirin treatment hampers the use of plasma microRNA-126 as a biomarker for the progression of vascular disease. Eur Heart J 34:3451–3457
Burckhardt CS, Clark SR, Bennett RM (1991) The fibromyalgia impact questionnaire: development and validation. J Rheumatol 18:728–733
Iannuccelli C, Sarzi-Puttini P, Atzeni F, Cazzola M, di Franco M, Guzzo MP, Bazzichi L, Cassisi GA et al (2011) Psychometric properties of the fibromyalgia assessment status (FAS) index: a national web-based study of fibromyalgia. Clin Exp Rheumatol 29:S49–S54
Bruce B, Fries JF (2005) The health assessment questionnaire (HAQ). Clin Exp Rheumatol 23:S14–S18
Zung WW (1971) A rating instrument for anxiety disorders. Psychosomatics 12:371–379
Zung WW (1972) The depression status inventory: an adjunct to the self-rating depression scale. J Clin Psychol 28:539–543
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L et al (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Masotti A, Alisi A (2012) Integrated bioinformatics analysis of microRNA expression profiles for an in-depth understanding of pathogenic mechanisms in non-alcoholic fatty liver disease. J Gastroenterol Hepatol 27:187–188
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57
Ceribelli A, Nahid MA, Satoh M, Chan EK (2011) MicroRNAs in rheumatoid arthritis. FEBS Lett 585:3667–3674
Andersen HH, Duroux M, Gazerani P (2014) MicroRNAs as modulators and biomarkers of inflammatory and neuropathic pain conditions. Neurobiol Dis 71:159–168
Gold MS, Gebhart GF (2010) Nociceptor sensitization in pain pathogenesis. Nat Med 16:1248–1257
Bai G, Ambalavanar R, Wei D, Dessem D (2007) Downregulation of selective microRNAs in trigeminal ganglion neurons following inflammatory muscle pain. Mol Pain 3:15
Poh KW, Yeo JF, Ong WY (2011) MicroRNA changes in the mouse prefrontal cortex after inflammatory pain. Eur J Pain 15:801.e1–801.12
Russo F, Di Bella S, Bonnici V, Lagana A, Rainaldi G, Pellegrini M, Pulvirenti A, Giugno R and Ferro A (2014) A knowledge base for the discovery of function, diagnostic potential and drug effects on cellular and extracellular miRNAs. BMC Genomics. 15 Suppl 3, S4-2164-15-S3-S4
Cerda-Olmedo G, Mena-Duran AV, Monsalve V, Oltra E (2015) Identification of a microRNA signature for the diagnosis of fibromyalgia. PLoS One 10:e0121903
White RE, Giffard RG (2012) MicroRNA-320 induces neurite outgrowth by targeting ARPP-19. Neuroreport 23:590–595
Orlova IA, Alexander GM, Qureshi RA, Sacan A, Graziano A, Barrett JE, Schwartzman RJ and Ajit SK, (2011) MicroRNA modulation in complex regional pain syndrome. J. Transl. Med. 9, 195-58769-195
Shen Y, Li Y, Ye F, Wang F, Wan X, Lu W, Xie X (2011) Identification of miR-23a as a novel microRNA normalizer for relative quantification in human uterine cervical tissues. Exp Mol Med 43:358–366
Blondal T, Jensby NS, Baker A, Andreasen D, Mouritzen P, Wrang TM, Dahlsveen IK (2013) Assessing sample and miRNA profile quality in serum and plasma or other biofluids. Methods 59:S1–S6
Chhabra R, Dubey R and Saini N (2010) Cooperative and individualistic functions of the microRNAs in the miR-23a~27a~24-2 cluster and its implication in human diseases. Mol. Cancer. 9, 232-45989-232
Wada S, Kato Y, Sawada S, Aizawa K, Park JH, Russell AP, Ushida T, Akimoto T (2015) MicroRNA-23a has minimal effect on endurance exercise-induced adaptation of mouse skeletal muscle. Pflugers Arch 467:389–398
Varendi K, Kumar A, Harma MA, Andressoo JO (2014) miR-1, miR-10b, miR-155, and miR-191 are novel regulators of BDNF. Cell Mol. Life Sci 71:4443–4456
Huang EJ, Reichardt LF (2001) Neurotrophins: roles in neuronal development and function. Annu Rev Neurosci 24:677–736
Nakajo Y, Miyamoto S, Nakano Y, Xue JH, Hori T, Yanamoto H (2008) Genetic increase in brain-derived neurotrophic factor levels enhances learning and memory. Brain Res 1241:103–109
Mousavi K, Jasmin BJ (2006) BDNF is expressed in skeletal muscle satellite cells and inhibits myogenic differentiation. J Neurosci 26:5739–5749
Nielsen S, Scheele C, Yfanti C, Akerstrom T, Nielsen AR, Pedersen BK, Laye MJ (2010) Muscle specific microRNAs are regulated by endurance exercise in human skeletal muscle. J Physiol 588:4029–4037
Nielsen S, Akerstrom T, Rinnov A, Yfanti C, Scheele C, Pedersen BK, Laye MJ (2014) The miRNA plasma signature in response to acute aerobic exercise and endurance training. PLoS One 9:e87308
Chen RJ, Kelly G, Sengupta A, Heydendael W, Nicholas B, Beltrami S, Luz S, Peixoto L et al (2015) MicroRNAs as biomarkers of resilience or vulnerability to stress. Neuroscience 305:36–48
Shi W, Du J, Qi Y, Liang G, Wang T, Li S, Xie S, Zeshan B et al (2012) Aberrant expression of serum miRNAs in schizophrenia. J Psychiatr Res 46:198–204
Acknowledgments
The authors thank Dr. Letizia Da Sacco for technical discussions about the quantification of circulating miRNAs from serum.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The study was conducted according to the protocol and good clinical practice principles and the Declaration of Helsinki statements. All patients gave their informed consent and the study was approved by the local Ethical Committee.
Funding
This work was supported by the SAPIENZA University of Rome to M.D.F, by the Italian Ministry of Health to A.M. and by the Italian Ministry for Education, University and Research in the framework of the Flagship Project NanoMAX to C.B.
Conflict of Interest
The authors declare that they have no competing interests.
Additional information
Key Messages
- Serum miRNAs correlate with major fibromyalgia (FM) symptoms
- A statistical linear model made of five miRNAs could discriminate FM patients from controls
- Liquid biopsies collected from different body parts have different miRNA profiles
Andrea Masotti and Antonella Baldassarre equally contributed to the work.
Rights and permissions
About this article
Cite this article
Masotti, A., Baldassarre, A., Guzzo, M.P. et al. Circulating microRNA Profiles as Liquid Biopsies for the Characterization and Diagnosis of Fibromyalgia Syndrome. Mol Neurobiol 54, 7129–7136 (2017). https://doi.org/10.1007/s12035-016-0235-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-0235-2