Abstract
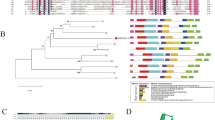
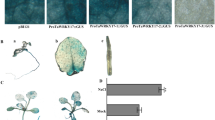
The MADS-box gene family has multiple molecular and biological functions in plants. Here, the LiSEP3 gene of the MADS-box gene family of’ ‘Sorbonne’ was obtained by homologous cloning using the petals of the flowering stage of Lilium Oriental Hybrid ‘Sorbonne.’ The ORF full-length sequence is 729 bp, encoding 242 amino acids. Bioinformatics analysis showed that the relative molecular weight of the LiSEP3 protein is 27.67 kD and the isoelectric point (pI) is 9.16. The prediction result of the gene positioning is transcription in its nucleus. Homologous alignment of amino acid sequences showed that the protein not only had typical MADS-box and K-box domains, but also contained two short and relatively conservative SEP motifs. The phylogenetic tree showed that the amino acid sequence encoded by the LiSEP3 gene had the closest relationship with SEP3 in monocotyledon plants such as Apostasia odorata. The results of real-time PCR showed that LiSEP3 gene was mainly expressed in petal. During flower development, the expression level of the LiSEP3 gene showed an overall trend of initially increasing and then decreasing. The flowering time of LiSEP3 transgenic Arabidopsis thaliana L. plants was earlier than that of wild-type Arabidopsis thaliana L. plants, compared with wild type, the number of rosette leaves is less. In the transgenic plants, the expression of flowering-associated AtSPL5 and AtGI genes was up-regulated, while the expression of AtSVP and AtFRI genes that inhibit flowering was down-regulated, which was consistent with the statistical results of the flowering time of LiSEP3 transgenic plants. Our results illustrate that the heterologous expression of SEP3 functional genes in the MADS-box family promoted the flowering period of transgenic plants of this hybrid. This research provides a theoretical basis for improving the flowering period of ornamental plants through plant genetic engineering technology and enhancing their economic and social values.






Similar content being viewed by others
References
Song, Y. H., Ito, S., & Imaizumi, T. (2013). Flowering time regulation: Photoperiod- and temperature-sensing in leaves. Trends in Plant Science, 18, 575–583.
Song, G. Q., & Chen, Q. (2018). Overexpression of the MADS-box gene K-domain increases the yield potential of blueberry[J]. Plant Science, 276, 22–31.
Yamaguchi, T., & Hirano, H. Y. (2006). Function and diversification of MADS-box genes in rice [J]. Scientific World Journal, 6, 1923–1932.
Shore, P., & Sharrocks, A. D. (1995). The MADS-box family of transcription factors. European Journal of Biochemistry, 229, 1–13.
Pelucchi, N., Fornara, F., Favalli, C., Masiero, S., Lago, C., Enrico, M. P., Colombo, L., & Kater, M. M. (2002). Comparative analysis of rice MADS-box genes expressed during flower development[J]. Sexual Plant Reproduction, 15, 113–122.
Honma, T., & Goto, K. (2001). Complexes of MADS-BOX proteins are sufficient to convert leaves into floral organs [J]. Nature, 409, 525–529.
Pelaz, S., Gustafson-Brown, C., Kohalmi, S. E., Crosby, W. L., & Yanofsky, M. F. (2001). APETALA1 and SEPALLATA3 interact to promote flower development [J]. Plant J, 26, 385–394.
Zahn, L. M., Kong, H., Leebens-Mack, J. H., Kim, S., Soltis, P. S., Landherr, L. L., Soltis, D. E., Depamphilis, C. W., & Ma, H. (2005). The evolution of the SEPLLATA subfamily of MADS-box genes: A preangiosperm origin with multiple duplications throughout angiosperm history [J]. Genetics, 169, 2209–2223.
Stellari, G. M., Jaramillo, M. A., & Kramer, E. M. (2004). Evolution of the APETALA3 and PISTILLATA lineages of MADS-box-containing genes in the basal angiosperms[J]. Molecular Biology and Evolution, 21, 506–519.
Immink, R. G., Tonaco, I. A., de Folter, S., Shchennikova, A., van Dijk, A. D., Busscher-Lange, J., Borst, J. W., & Angenent, G. C. (2009). SEPALLATA3: The “glue” for MADS box transcription factor complex formation[J]. Genome Biology, 10, R24.
Hugouvieux, V., Silva, C. S., Jourdain, A., Stigliani, A., Charras, Q., Conn, V., Conn, S. J., Carles, C. C., Parcy, F., & Zubieta, C. (2018). Tetramerization of MADS family transcription factors SEPALLATA3 and AGAMOUS is required for floral meristem determinacy in Arabidopsis[J]. Nucleic Acids Res, 46, 4966–4977.
Gregis, V., Sessa, A., Colombo, L., & Kater, M. M. (2006). AGL24, SHORT VEGETATIVE PHASE, and APETALAI redundantly control AGAMOUS during early stages of flower development in Arabidopsis[J]. The Plant Cell, 18, 1373–1382.
Michaels, S. D., Ditta, G., Gustafson-Brown, C., Pelaz, S., Yanofsky, M., & Amasino, R. M. (2003). AGL24 acts as a promoter of flowering in Arabidopsis and is positively regulated by vernalization[J]. The Plant Journal, 33, 867–874.
Hartmann, U., Höhmann, S., Nettesheim, K., Wisman, E., Saedler, H., & Huijser, P. (2000). Molecular cloning of SVP: A negative regulator of the floral transition in Arabidopsis[J]. The Plant Journal, 21, 351–360.
Angenent, G. C., Franken, J., Busscher, M., Weiss, D., & van Tunen, A. J. (1994). Co-suppression of the petunia homeotic gene fbp2 affects the identity of the generative meristem [J]. The Plant Journal, 5, 33–44.
Pelaz, S., Ditta, G. S., Baumann, E., Wisman, E., & Yanofsky, M. F. (2000). B and C floral organ identity functions require SEPALLATA MADS-box genes [J]. Nature, 405, 200–203.
Gong, B., Yi, J., Wu, J., Sui, J., Khan, M. A., Wu, Z., Zhong, X., Seng, S., He, J., & Yi, M. (2014). A novel heat stress transcription factor in lily (Lilium longiflorum), can interact with LIHSFA2 and enhance the theromotolerance of transgenic Arabidosis thaliana [J]. Plant Cell Reports, 33, 1519–1533.
Tzeng, T. Y., Hsiao, C. C., Chi, P. J., & Yang, C. H. (2003). Two lily SEPALLATA-like genes cause different effects on floral formation and floral transition in Arabidosis [J]. Plant Physiology, 133, 1091–1101.
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C (T)) method. Methods, 25, 402–408.
Song, Y. H., Smith, R. W., To, B. J., Millar, A. J., & Imaizumi, T. (2012). Fkf1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science, 336, 1045–1049.
Mandel, M. A., & Yanofsky, M. F. (1998). The Arabidopsis AGL9 MADS box gene is expressed in young flower primordia [J]. Sexual Plant Reproduction, 11, 22–28.
Ireland, H. S., Yao, J. L., Tomes, S., Sutherland, P. W., Nieuwenhuizen, N., Gunaseelan, K., Winz, R. A., David, K. M., & Schaffer, R. J. (2013). Apple SEPALLATA1/2-like genes control fruit flesh development and ripening [J]. The Plant Journal, 73, 1044–1056.
Zhou, Y., Xu, Z., Yong, X., Ahmad, S., Yang, W., Cheng, T., Wang, J., & Zhang, Q. (2017). SEP-class genes in Prunus mume and their likely role in floral organ development [J]. BMC Plant Biology, 17, 10.
Hwan Lee, J., Joon Kim, J., & Ahn, J. H. (2012). Role of SEPALLATA3 (SEP3) as a downstream gene of miR156-SPL3-FT circuitry in ambient temperature responsive flowering [J]. Plant Signaling & Behavior, 7, 1151–1154.
Rumpler, F., Theissen, G., & Melzer, R. (2018). A conserved leucine zipper-like motif accounts for strong tetramerization capabilities of SEPALLATA-like MADS-domain transcription factors [J]. Journal of Experimental Botany, 69, 1943–1954.
Goloveshkina, E. N., Shulga, O. A., Shchennikova, A. V., Kamionskaya, A. M., & Skryabin, K. G. (2012). Functional characterization of chrysanthemum SEPALLATA3 homologs CDM77 and CDM44 in transgenictobacco plants [J]. Doklady Biological Sciences, 443, 87–90.
Mendez-Vigo, B., Martinez-Zapater, J. M., & Alonso-Blanco, C. (2013). The flowering repressor SVP underlies a novel Arabidopsis thaliana QTL interacting with the genetic background[J]. PLoS Genet, 9, e1003289.
Acknowledgements
The authors thank for technical support from experimental center in college of life science in northeast agricultural university.
Funding
This research was funded by National Key Research and Development Projects (2016YFC0500306-02), ‘Young Talents’ Project of Northeast Agricultural University(18QC09), and National Natural Science Foundation of China(32001505).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cao, L., Liu, D., Jiang, F. et al. Heterologous Expression of LiSEP3 from Oriental Lilium Hybrid ‘Sorbonne’ Promotes the Flowering of Arabidopsis thaliana L. Mol Biotechnol 64, 1120–1129 (2022). https://doi.org/10.1007/s12033-022-00492-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00492-2