Abstract
A higher level of tumor-infiltrating lymphocytes (TILs) is associated with better prognosis in breast cancer patients. Adoptive transfer of lymphocytes coupled with conventional therapies has appealed to many clinicians and investigators as an effective treatment strategy for cancer patients, which necessitates efficient activation and expansion of cytotoxic T lymphocytes precisely targeting cancer cells. To comprehensively understand composition of TILs and to provide a grounding in adoptive T cell therapy, we analyzed the T cell receptor (TCR) repertoires in ex vivo–expanded TILs from nine breast cancer patients via next-generation sequencing. For the three of them, TCR repertoires of TILs gathered after the initial culture during 2 weeks were additionally analyzed and compared to those of TILs that underwent ex vivo rapid expansion procedure (REP). Diversity of TCR repertoire was variable among the patients. V/J segment usage in the clonotypes was similar among patients, with variable distribution of read counts for each V/J segment. The top 50% of most frequently observed VJ combinations was present in > 80% of the total clonotypes. Compared with TCGA data, the samples contained a similar amount of recurrent CDR3 sequences, but clonotype expansion was variable among the samples. In terms of clinicopathologic factor, presence of in vitro reactivity among triple-negative breast cancer cases seemed to be related to lower Shannon’s index, but p value was not statistically significant. In addition, the proportion of CD45RO+ cells out of CD8+ T cells were negatively correlated with Shannon’s diversity index for both TCRα and TCRβ chains (p = 0.010) via Spearman test. In this study, we identified a heterogeneous pattern of expanded T cell clones and stable usage of V/J segments in ex vivo–expanded TILs from breast cancer patients. Further large-scale studies are requisite to elucidate the clinical significance of TCR repertoires.
Similar content being viewed by others
Introduction
Undeniable evidence to suggest that immunology-based therapy is a novel promising strategy for cancer regression has emerged from the use of immune checkpoint inhibitors. This approach enhances the need for activation of cytotoxic T cells to target cancer cells. However, lack of immune cells prior to use of the immune checkpoint inhibitors is one of the intrinsic factors resulting in resistance to such therapies [1]. The strategy of expanding and activating T cells ex vivo and returning them into the patients has significantly attracted attention in the field of immunotherapy [2]. Therefore, it is important to generate sufficient amount of anti-tumoral T cells in vitro.
Since recognition of antigens is carried out by specific T cells, identifying the sequences of T cell receptors (TCRs) of tumor-infiltrating T cells can assist in vitro clonotype engineering, expansion of tumor-specific T cells, and their subsequent re-infusion into patients. The specificity of TCRs is defined by the variability in the complementarity determining region 3 (CDR3), resulting from recombination of the variable (V), diversity (D), and joining (J) segments. Tumor-infiltrating lymphocytes (TILs) might have a broad array of repertoires which are specific to tumor antigens and eventually playing a crucial role in cancer treatment [3]. Next-generation sequencing (NGS) technology can provide high-throughput data regarding CDR3 regions and, thus, enable TCR repertoire sequencing [4].
Breast cancer is known to be the most common cancer among women and 40,000 yearly deaths due to breast cancer are reported in the USA [5]. Although recent adjuvant and neoadjuvant systemic therapies have improved the patients’ survival rate, they continue to present resistance to those treatments. There is a significant correlation among survival outcome, clinicopathologic response to treatment, and a high TIL score in triple negative breast cancer (TNBC) in a neoadjuvant therapy setting [6, 7]. Per BIG 2-98, ECOG 2197, and ECOG 1199 clinical trials, TILs were also considered prognostic biomarkers in settings of TNBC adjuvant therapy [8, 9]. It has been known that the larger amounts of TILs are associated with the better survival of TNBC patients after adjuvant therapy. So far, we have actively studied regarding TILs and their associated immune system machineries using breast cancer samples. It would be considerably useful to characterize TCR sequences of TILs, which is the basis of selecting specific T cells against tumor antigens and targeting only tumor cells.
In our previous study, we successfully expanded TILs from breast cancer patients during the initial 2 weeks in most cases (177/182 cases), and utilized a rapid expansion protocol (REP) to obtain large amounts of TILs [10]. We also showed in vitro and in vivo anti-tumor activity of cultured TILs. Herein, we performed in-depth analysis of TCR repertoires of REP TILs obtained from nine breast cancer patients and compared them with recurrent CDR3s reported in The Cancer Genome Atlas (TCGA) as well as CDR3s known to be reactive with viral antigens reported in the VDJdb [11, 12]. Comparison of TCR repertoires between 2-week culture TILs and REP TILs and differences of TCR repertoires according to several clinicopathologic variables were also performed.
Materials and methods
Patients and tissue specimens
After obtaining informed consents, breast cancer tissue samples were harvested during surgery from nine female patients treated with or without neoadjuvant systemic chemotherapy at the Asan Medical Center, Seoul, Korea in 2016. This study was performed in accordance with the Declaration of Helsinki, and was approved by the Institutional Review Board of Asan Medical Center (IRB#2015–0438). For subtyping, cancer tissues were immunohistochemically stained with antibodies against estrogen receptor (ER, diluted 1:200; NCL-L-ER-6F11, Novocastra, Newcastle-upon-Tyne, UK), progesterone receptor (PR, diluted 1:200; NCL-L-PGR-312, Novocastra), and human epidermal growth factor receptor 2 (HER2, diluted 1:8; 88–4422, Ventana Medical Systems, Tucson, AZ, USA). ER and PR were evaluated via Allred score system in which their levels were regarded as positive if at least 1% of tumor nuclei were stained. Hormone receptor-positive (HR+) tumors were defined as those with positive ER and/or PR staining. Silver in situ hybridization for the HER2 gene was performed in cases with HER2 immunohistochemical staining result of 2+. The slides were evaluated for the levels of stromal TILs using full sections in 10% increments (if less than 10%, 0, 1, or 5% level criteria was used) [13].
Two week TIL culture and ex vivo expansion of TILs
Initial 2-week culture, ex vivo rapid expansion, and in vitro functional test of TILs obtained from breast cancer were done as previously described [10]. Briefly, tumor tissues were minced into 1-mm-diameter pieces. The minced fragments were plated in a 24-well plate (four fragments/well) containing 2 mL of TIL culture medium (RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS), 1× ZellShield (Minerva Biolabs, Berlin, Germany), 1× 2-mercaptoethanol, and 1000 IU/mL IL-2 (Miltenyi Biotec, Auburn, CA, USA)). The plates were incubated at 37 °C for 14 days to allow TILs to be released from the tissue fragment and expand. The TILs were then counted and cryopreserved. For further expansion of the TILs, standard REP was employed as follows: 5 × 104 TILs were cultured at 37 °C in a T25 flask containing 3 mL of 50% RPMI 1640 and 50% AIM-V media supplemented with 10% FBS, 1× ZellShield, and 1000 IU/mL IL-2 as well as 30 ng/mL human anti-CD3 antibody (OKT3, Miltenyi Biotec) and 1 × 107 irradiated (50 Gy) allogeneic peripheral blood mononuclear cell feeders. After 14 days, TILs (post-REP TILs) were collected, counted, and cryopreserved. To assess the in vitro reactivity of TILs against autologous tumor cells, 1 × 105 post-REP TILs were stimulated with PMA (32.4 nM, Sigma-Aldrich) and ION (1 μg/mL, Sigma-Aldrich) in 96-well plates for 24 h. Subsequently, the TILs were co-cultured for 24 h with 1 × 105 autologous breast cancer cells in 96-well plates at effector:target cell ratios of 1:1, 2:1, and 4:1. The capabilities of interferon-γ production by the TILs were evaluated by ELISA test and compared. When TILs release at least twice interferon-γ against their autologous tumor cells than when there were TILs alone, the case was designated to have significant in vitro reactivity.
Phenotype analysis of TILs using flow cytometry
To analyze the T cell composition, post-REP TILs were analyzed for surface marker expression. Briefly, TILs were washed three times with FACS buffer (PBS with 2% serum) and stained at 4 °C in the dark for 30 min with the following antibodies: PE-Cy7–labeled anti-CD3 (OKT3; BioLegend, San Diego, CA, USA), PerCP-Cy5.5–labeled anti-CD8 (HIT8a; BioLegend), PE-labeled anti-CD4 (RPA-T4; BD Biosciences, San Jose, CA), PE-labeled anti-CD45RO (UCHL1; BioLegend), and FITC-labeled anti-PD-1 (EH12.2H7; BioLegend). After washing with FACS buffer, the cells were stained with DAPI. The stained cells were evaluated using FACSCanto II (BD Biosciences) and analyzed with FlowJo software (Tree Star, Ashland, OR, USA).
Total RNA extraction and TCR sequencing
Total RNA was purified from 1 × 105 post-REP TILs using the RNeasy micro kit (Qiagen, Valencia, Ca, USA). Integrity of the total RNA was checked using a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, Ca, USA), and all RNA samples showed RIN numbers greater than 9.0. The TCRα/β sequencing library was generated via semi-nested PCR using the SMARTer human TCR α/β profiling kit (Clontech Laboratories, CA, USA). Briefly, TCR cDNA was synthesized from 100 μg of total RNA using the TCR dT primer and MMLV-derived SMARTScribe Reverse Transcriptase. For the first PCR step, cDNA templates were pre-amplified using SMART (Switching Mechanism at 5′ End of RNA Template) primer 1 and TCRα/β human primer 1, which is complementary to the constant region of TCR α/β. In this step, twice the amount of TCRα human primer 1 was used compared to the TCRβ human primer per the kit manual. The PCR conditions were denaturation at 95 °C for 1 min, 21 cycles of 94 °C for 1 min, 53 °C for 1 min, and 68 °C for 1 min, and then extension at 72 °C for 10 min. For the second PCR step, 1 μL of the first PCR product was amplified using TCR primer 2 and TCRα/β human primer 2. The resultant PCR products were size-selected and purified. The TCRα/β sequencing library was sequenced on the Illumina MiSeq platform (paired-end 300 bp).
TCR repertoire analysis
Read counts, CDR3 amino acid sequences, and V, D, and J segments were generated using MiXCR-version 1.8.1. [14] The TCRs with the same V gene, J gene, and CDR3 amino acid sequences were defined as one clonotype [15]. A productive clonotype was specified as a clonotype with no out-of-frame or stop codon in the CDR3 amino acid sequences. Only the CDR3 amino acid sequences starting with cysteine (C) and ending with phenylalanine (F) or tryptophan (W) were included. For calculation of TCR diversity, we used the following formula to assess the Shannon diversity index: \( H=-{\sum}_{i=1}^s\left( Pi\ast \ln Pi\right) \). Where H = the Shannon diversity index; ∑ = sum from clone 1 to clone S; S = numbers of clones; Pi = proportion of the sequence of our interest in the entire population. Statistical analyses with Mann Whitney test were carried out using SPSS version 20 (SPSS, Chicago, IL). Plots were generated using GraphPad Prism 7 (GraphPad Software Inc., La Jolla, CA, USA).
Comparison to CDR3s in TCGA data and VDJdb data
To identify recurrent CDR3s found in cancers, we downloaded processed data from a recently published study which generated a list of recurrent CDR3s identified via TCGA RNA sequencing data analysis of 8143 tumors representing 33 diverse cancer types [11]. We also downloaded TCR sequences with known antigen specificities from the VDJdb [12]. We compared 16,159 CDR3s known to be reactive to viral antigens reported in the VDJdb with our CDR3 list.
Correlation between characteristics of TCR repertoires and phenotypes of REP TILs
We executed Spearman correlation test to investigate whether TCR repertoires of ex vivo–expanded TILs were significantly associated with certain phenotypes of TILs. Representative variables including the proportion that the most common clonotype covers, Shannon’s diversity index, the number of total clonotypes, and that of productive clonotypes. Statistical analyses were carried out using SPSS version 20 (SPSS, Chicago, IL) as mentioned above.
Results
Study population and composition of REP samples
Nine invasive ductal carcinoma tissue samples from cases of primary tumors, lymph node metastasis, and metastatic breast cancer in the liver were included in this study. Among them, six samples were triple-negative breast cancer (TNBC) and three were HR+. Five patients received neoadjuvant chemotherapy. Three of the cases had analyzed results from TILs obtained after the initial 2-week culture.
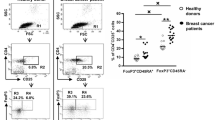
We phenotypically analyzed the T cells in post-REP samples. The samples were mostly composed of CD3+ T cells (Table 1). Composition of CD4+ and CD8+ T cells varied from 1:9 to 7.5:2.5. Over half of the CD3+ T cells showed CD45RO positivity (range, 55.7–99.2%). The frequency of PD-1+ cells, which are considered to be reactive to tumor, among the CD8+ cells was up to 40% (Supplementary Fig. 1) [16]. The amount of post-REP TILs, however, were unfortunately not enough to be analyzed for additional flow cytometry to elucidate the proportion of gammadelta T cells.
Basic TCR statistics
Following amplification of CDRs in the TCRα/β of T cells using semi-nested PCR, NGS analysis was performed to distinguish different repertoires of post-REP TILs from breast cancers. The results were summarized in Table 2. An average of 717,921 reads for the TCRα chain and 156,308 for TCRβ were sequenced in each sample. The productive read counts accounted for 89.3% and 95.8% of total TCRα and TCRβ, respectively. An average of 2404 clonotypes of TCRα and 1725 clonotypes of TCRβ were identified, which were productive. Additionally, the average frequency of the most common clonotype was 24.8% for TCRα and 25.6% for TCRβ. On average, the median read count in all the clonotypes was eight for TCRα and five for TCRβ.
Same basic information regarding TCRα/β of TILs harvested after the initial 2-week culture (BC16092, BC16096, and BC16126) was also investigated and tabulated in Table 2. An average of 725,234 and 163,560 total read counts were sequenced in each sample for the TCRα and TCRβ chain, respectively. An average of total 10,271 clonotypes of TCRα and total 6175 clonotypes of TCRβ were identified, but the number of clonotypes as well as the frequency of the most common clonotypes considerably varied among the three cases. Averagely, the median read count in all the clonotypes was three for both TCRα and TCRβ in those samples.
In addition, association between some representative characteristics of TCR repertoires and phenotypes of post-REP TILs was analyzed by Spearman’s rank correlation. Shannon’s diversity index and CD45RO+/CD8+ ratio were shown to have statistically significant negative correlation (rho = − 1.000, p = 0.010) in both alpha and beta chain (Supplementary Table 1).
The frequency and diversity index of clonotypes
The frequency of different clones in each sample is shown in Fig. 1A, which demonstrates the heterogeneity and complexity of T cell repertoires in intratumoral sites. The frequency of the most expanded clones in each REP ranged from 5.7 to 57.0% for TCRα and from 7.8 to 44.5% for TCRβ. The frequency of the second most expanded clones showed a narrower range, from 3.2 to 16.6% for TCRα and from 4.3 to 21.1% for TCRβ. Some REP samples (BC16096 and BC16158) were composed of several highly expanded clones, and the accumulative proportion of the top ten clones was more than 90% (Fig. 1B). On the contrary, other REP samples (BC16126, BC16127, and BC16143) were composed of low-frequency clones and the accumulative proportion of the top 300 clones was only about 80%. These cases showed a higher Shannon diversity index, representing more diversity. The proportion of the most expanded clones in TCRα and β showed similar patterns, but they were not identical (Fig. 1A).
The proportion of clonotypes in ex-vivo rapid expanded (REP) TILs from patients. A The 100% stacked cumulative column chart of the top 300 TCR clones is displayed in each sample. The top 50 clones commonly represent an abundance of more than 50% of total, whereas heterogeneity of TCR repertoire among the patients are well demonstrated. B For clear depiction of difference in clonality, the top 300 clones in each sample are shown in separate stacked line graphs
Three among the entire nine cases, TCR repertoires of TILs after the initial culture during 2 weeks were additionally analyzed and compared to those of TILs that underwent rapid expansion procedure. Shannon diversity index showed that unevenness of TCR sequences obtained after REP samples was consistently increased when compared to that of the initial 2-week culture samples (Fig. 2). Frequency of the most common clonotype is accordingly shown to be increased in all cases after REP.
Comparison of TCR repertoires between TILs from 2-week culture samples and those obtained after rapid expansion protocol (REP). In all three paired cases, Shannon diversity indices of TCR repertoires are consistently decreased after REP (A). Accordingly, proportion of the most common clonotype in each sample tends to be increased after REP (B)
V and J usage and VJ recombination in REP TILs
After evaluating the diversity, heterogeneity, and frequency of clonotypes along with their read counts, we analyzed V and J segment usage in TCRα/β. There was distinctive usage pattern for each V and J segment (Fig. 3). Of all the 49 identified TRAV segments, TRAV13-1 (5.4%), TRAV12-2 (5.2%), TRAV17 (4.5%), TRAV29DV5 (4.3%), and TRAV9-2 (4.0%) were the most frequent genes in the clonotypes of all patients. The most abundant segments in the 55 TRAJ genes in the clonotypes were TRAJ49 (3.8%), TRAJ42 (3.2%), TRAJ40 (3.1%), and TRAJ43 (3.1%). Among the 59 TRBV segments, TRBV20-1 (8.8%), TRBV28 (5.8%), TRBV5-1 (5.0%), TRBV7-9 (4.6%), TRBV2 (4.5%), and TRBV27 (4.3%) were the most frequent genes in the clonotypes. In addition, the most abundant segments in the 13 TRBJ genes were TRBJ2-7 (19.2%), TRBJ2-1 (17.3%), TRBJ2-3 (10.7%), and TRBJ1-1 (10.0%).
Comparison of individual V and J gene usage in clonotypes and read counts of post-rapid expansion protocol (REP) TILs. Proportions of each segment of the clonotypes show a narrow distribution, while those of actual read counts are widely distributed (Mann Whitney test, *p < 0.05, **p < 0.01, ***p < 0.001)
We compared V and J segment usage regarding clonotypes and read counts. The proportion range for each gene in the clonotypes was narrow among the nine patients, while the read count proportion for each gene was widely distributed (Fig. 3). These results showed that V and J segment usage in the clonotypes were similar in different breast cancer patients, but expansion of clonotypes with those segments was variable among them.
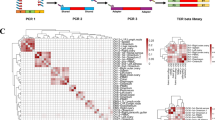
To define the substantial diversity of the TCR repertoire, we characterized the VJ recombination of each TCR clone (Fig. 4). Of 2695 possible VJ combinations for TCRα (49 TRAV and 55 TRAJ genes), 2007 VJ combinations were identified. For TCRβ, 595 VJ combinations were present (767 possible VJ combinations in 59 TRBV and 13 TRBJ genes). On average, the five most frequent VJ combinations among the total clonotypes in TCRα were TRAV12-3/TRAJ17 (0.37%), TRAV26-2/TRAJ49 (0.29%), TRAV13-1/TRAJ6 (0.28%), TRAV1-2/TRAJ33 (0.27%), and TRAV38-1/TRAJ41 (0.26%). The top 50% of the most frequently observed VJ combinations in TCRα were present in 82.6% of all the clonotypes. On the other hand, the five most frequent VJ combinations in TCRβ were TRBV20-1/TRBJ2-7 (1.7%), TRBV20-1/TRBJ2-1 (1.5%), TRBV28/TRBJ2-7 (1.3%), TRBV7-2/TRBJ2-7 (1.0%), and TRBV2/TRBJ2-7 (1.0%). In TCRβ, 89.3% of all the clonotypes consisted of the top 50% of the most frequently observed VJ combinations.
The shared CDR3s with public data
Next, we compared our TCR sequencing results with recurrent CDR3s identified via TCGA RNA sequencing data analysis of 8143 tumors representing 33 diverse cancer types [11]. Thorsson et al. found 16,971 recurrent CDR3s in TCRα and 12,079 recurrent CDR3s in TCRβ in TCGA cohorts, defined as being present in two or more patients. Recurrent CDR3s were present in up to 329 patients and the average number of read counts for recurrent CDR3s was two per patient in TCRα. In TCRβ, recurrent CDR3s were found in up to 95 patients and the average number of read counts per patient was 1.5. The average CDR3 length of these recurrent CDR3s in TCGA data was 12.4 amino acids in TCRα and 12.7 in TCRβ. When we compared our data to TCGA data, we found that 3346 recurrent CDR3s in TCRα and 590 CDR3s in TCRβ from the TCGA data were also present in our REP TIL repertoires. For each sample, 14.5–16.5% of TCRα clonotypes included recurrent CDR3s in the TCGA data, while their presence in actual read counts was 1.5–34.3%. For TCRβ, 3.3–4.3% of clonotypes included recurrent CDR3s in the TCGA data, while they comprised 0.2–17.8% of actual read counts. Therefore, each sample contained a similar amount of CDR3s which were recurrently present in TCGA RNA sequencing data, but the expansion of each clonotype was variable.
Because shared TCRs are known to have shorter CDR3 length, we compared CDR3 length between clonotypes present in the TCGA recurrent CDR3 data and those not present in the TCGA data [15, 17]. When we analyzed the length of amino acids in the CDR3 of our nine patients, the average length of the CDR3 sequence in the clonotypes was 13.8 amino acids in TCRα and 14.6 in TCRβ, respectively. The average CDR3 length in clonotypes present in the TCGA recurrent CDR3 data was 12.5 for TCRα and 12.7 for TCRβ, while that in clonotypes not present in the TCGA data was 14.0 for TCRα and 14.6 for TCRβ (Fig. 5; Mann Whitney test, p < 0.001 for both TCRα and TCRβ).
Comparison of CDR3 amino acid length distribution between shared CDR3s with TCGA data and not shared ones. The median CDR3 length in TCRα and TCRβ is 14 amino acids in total REP TILs. Shared TCR clonotypes show significantly shorter CDR3 lengths than both not shared ones and all (Mann Whitney test, p < 0.001 for both TCRα and TCRβ). The average CDR3 length in clonotypes present in the TCGA recurrent CDR3 data was 12.5 for TCRα and 12.7 for TCRβ, while that in clonotypes not present in the TCGA data was 14.0 for TCRα and 14.6 for TCRβ
CDR3s associated with microbial antigens
Next, we analyzed the presence of CDR3s showing reactivity to viral antigens by comparing data from the VDJdb which curates TCR and antigen specificity data from published articles. We identified 171 distinct CDR3s reactive to viral antigens in total (94 in TCRα and 77 in TCRβ). Among them, 52 CDR3s were known to be reactive to cytomegalovirus (CMV), 41 to Influenza A virus, 38 to yellow fever virus, 19 to Epstein Barr virus (EBV), and the rest to hepatitis C virus (HCV), human immunodeficiency virus-1 (HIV-1), human T lymphotropic virus (HTLV-1), and dengue virus (DENV). On average, 0.6% and 0.5% of the clonotypes and 0.3% each of the total read counts for TCRα and TCRβ included these CDR3s. Of 171 CDR3s known to be reactive to viral antigens, 112 were also present in the recurrent CDR3 set in the TCGA data.
The presence of mucosal-associated invariant T cells (MAIT cells) that are well known to have high specificity for microbial riboflavin–associated antigen was also analyzed. Our data from the nine patients showed the proportion of TRAV1-2/TRAJ33 as well as TRAJ12 and TRAJ20 corresponding to MAIT cells was 0.47% of total clonotypes, and the frequency ranged from 0.14 to 0.93% in each sample. Assumed proportion of MAIT cells out of entire productive read count ranged < 0.001 to 0.545%.
In case of invariant natural killer T (iNKT) cells which consistently express TRAV10-TRAJ18 in humans, only 14 clonotypes (0.02%) were identified in total. There were some samples without detected iNKT cells in contrast to MAIT cells.
Correlation of characteristics of TCR repertoires with clinicopathologic variables
In addition, we divided the patients by several clinicopathologic factors and compared their TCR repertoires. First of all, there were three cases of HR+ breast cancer and six of TNBCs among a total of nine patients. Not only TCR repertoires including total/productive read counts and TCR clonotypes but also Shannon’s diversity index showed no difference between the two subgroups by Mann-Whitney test.
Next, we divided the six TNBC patients into four cases with in vitro reactivity and the others without it. TILs which showed significant interferon-γ production against their autologous tumor cells (more than twice that in TILs alone) at ELISA test were defined to have in vitro reactivity. The cases which demonstrated in vitro reactivity (range of fold change 3.1–135.8) seemed to have lower Shannon’s diversity index in both TCR alpha and beta chains, and accordingly, higher proportion of the most common clonotype (Fig. 6). This result may represent that effective clonal expansion of tumor-specific T cells was elicited during REP in the cases showing in vitro reactivity. However, their p values were not statistically significant by Mann-Whitney test, either.
Comparison of characteristics of TCR repertoires according to clinicopathologic variables. Six patients with triple-negative breast cancer were divided into two groups by in vitro reactivity (present vs absent), and characteristics of ex vivo expanded TILs were compared. TILs which showed significant interferon-γ production against their autologous tumor cells at ELISA test were defined to have in vitro reactivity. The cases with in vitro reactivity (n = 4) tend to have fewer number of clonotypes, lower Shannon’s diversity index, and higher proportion of the most common clonotypes in both TCR alpha and beta chain
In terms of neoadjuvant chemotherapy, five patients were treated with therapeutic agent. One case obtained from metastatic lesion in the liver was excluded for the analysis, and the others were naive to chemotherapy. Neither neoadjuvant treatment nor lymph node metastasis status affected basic composition of TCR sequences.
In addition, we analyzed correlation coefficient between several characteristics of TCR repertoires and phenotypes of post-REP TILs via Spearman’s test (Supplementary Table 1). Interestingly, the proportion out of CD8+ T cells which CD45RO+ cells accounted for was negatively correlated with Shannon’s diversity index for both TCRα and TCRβ chains (p = 0.010). It might suggest that larger amount of memory T cells is associated with lower Shannon’s H which represents higher clonality. In alpha chain, accordingly, the proportion of CD45RO+ cells out of CD8+ T cells was positively correlated with frequency of the most common clonotype, whereas it was not in beta chain.
Discussion
The consensus that TILs are the key mediators triggering immune responses against tumors has attracted significant attention in immunotherapy of cancer. Therefore, investigating the nature of T cell clonotypes in the tumor microenvironment might be helpful in developing cancer immunotherapy. For deeper comprehension of the interaction between T cells (representing the adaptive immune system) and antigens of cancer, we need to characterize these clonotypes. The present report outlines the diversity of T cell clonotypes in post-REP samples from breast cancer patients.
Our present study suggests that TILs in the tumor microenvironment would be highly heterogeneous and polyclonal. Although there were clonotypes occurring at high frequency, most of the clones in each sample were present at a low frequency. The highly expanded clones are most likely indicative of T cells exposed to antigens [18]. In addition, TCR repertoires analyzed in TILs after REP showed lower Shannon diversity index and higher proportion of most frequent TCR clonotype which mean increased unevenness compared to those analyzed in TILs obtained after the initial 2-week culture. It could be assumed to be the result from clonal expansion of TILs which have TCRs specific to the tumor surface antigen.
However, most of the T cells in REP samples showed CD45RO positivity and we have previously found that freshly dissociated TILs from breast cancer contained more than 80% of CD45RO+ T cells, which are associated with effector and memory phenotypes [10]. Therefore, clonotypes occurring at a low frequency might also be reactive to tumor antigens, but they might be suppressed and down-regulated in the tumor niche. To evaluate this possibility, further studies are needed to explore the anti-tumor reactivity of REP TILs.
Because we analyzed TCRα repertoires in addition to the TCRβ repertoires usually solely assessed in most studies, we could compare the frequency of the most expanded clonotypes of TCRα and TCRβ. They showed a similar pattern of frequency, but some cases showed large differences. These might have been caused by different total read counts for TCRα and TCRβ, as we added twice the amount of primers for PCR-based quantification of TCRα detection than for TCRβ detection following the instructions. As a result, the productive read count for TCRα was more than three times higher than that for TCRβ (range, 3.0–6.9 times). However, productive clonotypes of TCRα were only 1.3 times higher than those of TCRβ (range, 1.3–2.1 times). Another possible explanation of these discrepancies could be differential allelic exclusion of TCRα and TCRβ during TCR gene rearrangement [19, 20]. Allelic exclusion in TCRα is less strict than TCRβ resulting in two different VJ recombination events in up to 30% of T cells [21, 22]. Therefore, if the most expanded clonotypes possess two rearranged TCRα chains, frequencies of the most expanded TCRα and TCRβ clonotypes may be different [23].
We found that basic V and J gene usage for generating distinct T cell clones were similar among the individuals. However, read counts of V and J segments were variable among the patients suggesting different levels of expansion in each T cell repertoire. Differential expression of V and J segments was reported previously based on MHC locus in individuals, differentiation status of T cells, and CDR3 length [17, 24, 25]. An attempt to correlate VJ usage of TILs and prognosis of cancer patients has also been reported [26]. Studies analyzing clinical significance of TCR V or J gene usage in cancer patients would provide a basis for effective immunotherapy.
In comparison with TCGA data from 8143 tumors representing 33 cancer types, we found a similar amount of recurrent CDR3s in each of our nine breast cancer REP TILs (14.5–16.5% of productive clonotypes for TCRα and 3.3–4.3% for TCRβ). However, the actual read counts were significantly variable between patients. Therefore, the REP TILs contained a certain amount of sequences occurring in multiple people, and some of them were reactive to breast cancer epitopes and expanded. These results are consistent with a previous analysis of early-stage breast cancer [15]. Beausang et al. compared CDR3s in 16 invasive breast carcinoma patients (tumor, normal breast, and blood) with TCR repertoires of 585 healthy donors. They found that shared CDR3 sequences in tumors of breast cancer patients were commonly present in healthy donor repertoires (more than 30%), while tumor-enriched CDR3s were much less frequently present (less than 5%). The average length of CDR3 sequences we found, 13.8 amino acids in TCRα and 14.6 in TCRβ, was consistent with previously reported CDR3 length (14 amino acids in TCRα and 15 in TCRβ) in an analysis of 20 breast cancer cases [27]. We also found significantly shorter CDR3 lengths in clonotypes recurrently present in the TCGA dataset in line with a previous study [17]. A shorter CDR3 region in shared, public TCRs have been explained based on a more efficient recombination process [28, 29].
All TILs are not necessarily reactive to tumor antigens, and viral antigen-reactive TILs have been identified [30, 31]. Using multiplex MHC tetramer staining, Simoni et al. found viral (EBV, CMV, and influenza virus) antigen-specific MHC tetramer positive cells in nine of 24 lung cancer TILs and 21 of 42 colorectal cancer TILs [30]. Less than 3.3% of all the CD8+ TILs were found to be reactive to each epitope. We compared our data to a set of curated TCRs reactive to viral antigens [12]. On average, less than 1% of clonotypes in our data were identified in the VDJdb. These clonotypes comprised less than 0.5% of the total read count, indicating that the clonotypes were not expanded in the REP TILs. However, we could not exclude the possibility that more specific clonotypes with viral antigen were present in the REP TILs, because limited data regarding T cell specificity and clonotypes are available for now.
In the present study, it seemed that presence of in vitro reactivity was associated with lower Shannon’s diversity index in both TCR alpha and beta chains, and higher frequency of the most common clonotype. It may represent that clonal expansion of tumor-specific T cells accomplished during REP results in in vitro reactivity. Other obvious relationship between clinical information and TCR repertoires could not be elicited here. Clinical significance of TCR repertoires has been recently studied, and some studies presented the results showing T cell clonality or intratumoral TCR heterogeneity was related in some cancer types [32, 33].
Limitations of this study include its small sample size and lack of TCR analysis obtained from primary tumor before ex vivo expansion. Koning et al. found that T cell repertoires of antigen-specific CD8+ T cells were clearly altered through in vitro stimulation and expansion [34]. Therefore, further studies including comparison of TCRs among TILs immediately extracted after tumor tissue dissociation, expanded TILs, and T cells from peripheral blood, as well as evaluation of TCR repertoires of CD4+, CD8+, or various T cell differentiation states are warranted.
Conclusions
In conclusion, we identified a heterogeneous pattern of expanded clones in REP TILs from breast cancer patients; some REP samples were composed of several highly expanded clones and showed lower diversity, while others had higher diversity. REP TILs in breast cancer patients showed similar TCR V and J gene segment usage in the clonotypes, but variable expansion of T cell clones resulted in different V/J usage values in the actual read counts. Further large-scale studies analyzing the clinical significance of TCR characteristics are needed.
Funding statement
This study was supported by the Korean Health Technology R&D Project, Ministry of Health & Welfare, Republic of Korea (HI15C0708 and HI17C0337).
References
Chae YK, Oh MS, Giles FJ. Molecular biomarkers of primary and acquired resistance to T-cell-mediated immunotherapy in cancer: landscape, clinical implications, and future directions. Oncologist. 2017;23:410–21. https://doi.org/10.1634/theoncologist.2017-0354.
Yang JC, Rosenberg SA. Adoptive T-cell therapy for cancer. Adv Immunol. 2016;130:279–94. https://doi.org/10.1016/bs.ai.2015.12.006.
Bobisse S, Genolet R, Roberti A, Tanyi JL, Racle J, Stevenson BJ, et al. Sensitive and frequent identification of high avidity neo-epitope specific CD8 (+) T cells in immunotherapy-naive ovarian cancer. Nat Commun. 2018;9(1):1092. https://doi.org/10.1038/s41467-018-03301-0.
Mahe E, Pugh T, Kamel-Reid S. T cell clonality assessment: past, present and future. J Clin Pathol. 2018;71(3):195–200. https://doi.org/10.1136/jclinpath-2017-204761.
Shachar SS, Mashiach T, Fried G, Drumea K, Shafran N, Muss HB, et al. Biopsy of breast cancer metastases: patient characteristics and survival. BMC Cancer. 2017;17(1):7. https://doi.org/10.1186/s12885-016-3014-6.
Denkert C, von Minckwitz G, Brase JC, Sinn BV, Gade S, Kronenwett R, et al. Tumor-infiltrating lymphocytes and response to neoadjuvant chemotherapy with or without carboplatin in human epidermal growth factor receptor 2-positive and triple-negative primary breast cancers. J Clin Oncol. 2015;33(9):983–91. https://doi.org/10.1200/JCO.2014.58.1967.
Dieci MV, Criscitiello C, Goubar A, Viale G, Conte P, Guarneri V, et al. Prognostic value of tumor-infiltrating lymphocytes on residual disease after primary chemotherapy for triple-negative breast cancer: a retrospective multicenter study. Ann Oncol. 2014;25(3):611–8. https://doi.org/10.1093/annonc/mdt556.
Adams S, Gray RJ, Demaria S, Goldstein L, Perez EA, Shulman LN, et al. Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin Oncol. 2014;32(27):2959–66. https://doi.org/10.1200/JCO.2013.55.0491.
Loi S, Sirtaine N, Piette F, Salgado R, Viale G, Van Eenoo F, et al. Prognostic and predictive value of tumor-infiltrating lymphocytes in a phase III randomized adjuvant breast cancer trial in node-positive breast cancer comparing the addition of docetaxel to doxorubicin with doxorubicin-based chemotherapy: BIG 02-98. J Clin Oncol. 2013;31(7):860–7. https://doi.org/10.1200/JCO.2011.41.0902.
Lee HJ, Kim YA, Sim CK, Heo SH, Song IH, Park HS, et al. Expansion of tumor-infiltrating lymphocytes and their potential for application as adoptive cell transfer therapy in human breast cancer. Oncotarget. 2017;8(69):113345–59. https://doi.org/10.18632/oncotarget.23007.
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, et al. The immune landscape of cancer. Immunity. 2018;48(4):812–30 e814. https://doi.org/10.1016/j.immuni.2018.03.023.
Shugay M, Bagaev DV, Zvyagin IV, Vroomans RM, Crawford JC, Dolton G, et al. VDJdb: a curated database of T-cell receptor sequences with known antigen specificity. Nucleic Acids Res. 2018;46(D1):D419–27. https://doi.org/10.1093/nar/gkx760.
Hendry S, Salgado R, Gevaert T, Russell PA, John T, Thapa B, et al. Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the International Immunooncology Biomarkers Working Group: part 1: assessing the host immune response, TILs in invasive breast carcinoma and ductal carcinoma in situ, metastatic tumor deposits and areas for further research. Adv Anat Pathol. 2017;24(5):235–51. https://doi.org/10.1097/PAP.0000000000000162.
Bolotin DA, Poslavsky S, Mitrophanov I, Shugay M, Mamedov IZ, Putintseva EV, et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat Methods. 2015;12(5):380–1. https://doi.org/10.1038/nmeth.3364.
Beausang JF, Wheeler AJ, Chan NH, Hanft VR, Dirbas FM, Jeffrey SS, et al. T cell receptor sequencing of early-stage breast cancer tumors identifies altered clonal structure of the T cell repertoire. Proc Natl Acad Sci U S A. 2017;114(48):E10409–17. https://doi.org/10.1073/pnas.1713863114.
Gros A, Robbins PF, Yao X, Li YF, Turcotte S, Tran E, et al. PD-1 identifies the patient-specific CD8(+) tumor-reactive repertoire infiltrating human tumors. J Clin Invest. 2014;124(5):2246–59. https://doi.org/10.1172/JCI73639.
Hou X, Wang M, Lu C, Xie Q, Cui G, Chen J, et al. Analysis of the repertoire features of TCR Beta chain CDR3 in human by high-throughput sequencing. Cell Physiol Biochem. 2016;39(2):651–67. https://doi.org/10.1159/000445656.
Pasetto A, Gros A, Robbins PF, Deniger DC, Prickett TD, Matus-Nicodemos R, et al. Tumor- and neoantigen-reactive T-cell receptors can be identified based on their frequency in fresh tumor. Cancer Immunol Res. 2016;4(9):734–43. https://doi.org/10.1158/2326-6066.CIR-16-0001.
Rybakin V, Westernberg L, Fu G, Kim HO, Ampudia J, Sauer K, et al. Allelic exclusion of TCR alpha-chains upon severe restriction of V alpha repertoire. PLoS One. 2014;9(12):e114320. https://doi.org/10.1371/journal.pone.0114320.
Skok JA, Gisler R, Novatchkova M, Farmer D, de Laat W, Busslinger M. Reversible contraction by looping of the Tcra and Tcrb loci in rearranging thymocytes. Nat Immunol. 2007;8(4):378–87. https://doi.org/10.1038/ni1448.
Padovan E, Casorati G, Dellabona P, Meyer S, Brockhaus M, Lanzavecchia A. Expression of two T cell receptor alpha chains: dual receptor T cells. Science. 1993;262(5132):422–4.
Kim JY, Heo SH, Song IH, Park IA, Kim YA, Gong G, et al. Activation of the PERK-eIF2alpha pathway is associated with tumor-infiltrating lymphocytes in HER2-positive breast cancer. Anticancer Res. 2016;36(6):2705–11.
Krangel MS. Mechanics of T cell receptor gene rearrangement. Curr Opin Immunol. 2009;21(2):133–9. https://doi.org/10.1016/j.coi.2009.03.009.
Sharon E, Sibener LV, Battle A, Fraser HB, Garcia KC, Pritchard JK. Genetic variation in MHC proteins is associated with T cell receptor expression biases. Nat Genet. 2016;48(9):995–1002. https://doi.org/10.1038/ng.3625.
Afik S, Yates KB, Bi K, Darko S, Godec J, Gerdemann U, et al. Targeted reconstruction of T cell receptor sequence from single cell RNA-seq links CDR3 length to T cell differentiation state. Nucleic Acids Res. 2017;45(16):e148. https://doi.org/10.1093/nar/gkx615.
Callahan BM, Yavorski JM, Tu YN, Tong WL, Kinskey JC, Clark KR, et al. T-cell receptor-beta V and J usage, in combination with particular HLA class I and class II alleles, correlates with cancer survival patterns. Cancer Immunol Immunother. 2018;67:885–92. https://doi.org/10.1007/s00262-018-2139-7.
Munson DJ, Egelston CA, Chiotti KE, Parra ZE, Bruno TC, Moore BL, et al. Identification of shared TCR sequences from T cells in human breast cancer using emulsion RT-PCR. Proc Natl Acad Sci U S A. 2016;113(29):8272–7. https://doi.org/10.1073/pnas.1606994113.
Venturi V, Quigley MF, Greenaway HY, Ng PC, Ende ZS, McIntosh T, et al. A mechanism for TCR sharing between T cell subsets and individuals revealed by pyrosequencing. J Immunol. 2011;186(7):4285–94. https://doi.org/10.4049/jimmunol.1003898.
Robins HS, Srivastava SK, Campregher PV, Turtle CJ, Andriesen J, Riddell SR, et al. Overlap and effective size of the human CD8+ T cell receptor repertoire. Sci Transl Med. 2010;2(47):47ra64. https://doi.org/10.1126/scitranslmed.3001442.
Simoni Y, Becht E, Fehlings M, Loh CY, Koo SL, Teng KWW, et al. Bystander CD8(+) T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature. 2018;557:575–9. https://doi.org/10.1038/s41586-018-0130-2.
Baitsch L, Baumgaertner P, Devevre E, Raghav SK, Legat A, Barba L, et al. Exhaustion of tumor-specific CD8(+) T cells in metastases from melanoma patients. J Clin Invest. 2011;121(6):2350–60. https://doi.org/10.1172/JCI46102.
Miao Y, Xiong H, Lu Z, Ikeda Y, Kiyotani K, Yew PY, et al. Clinical significance of T cell clonality and expression levels of immune-related genes in endometrial cancer. Int J Cancer. 2017;37(5):2603–10. https://doi.org/10.1002/ijc.3176010.3892/or.2017.5536.
Zhang C, Ding H, Huang H, Palashati H (2018) TCR repertoire intratumor heterogeneity of CD4(+) and CD8(+) T cells in centers and margins of localized lung adenocarcinomas. https://doi.org/10.1002/ijc.31760.
Koning D, Costa AI, Hasrat R, Grady BP, Spijkers S, Nanlohy N, et al. In vitro expansion of antigen-specific CD8(+) T cells distorts the T-cell repertoire. J Immunol Methods. 2014;405:199–203. https://doi.org/10.1016/j.jim.2014.01.013.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Précis
· Despite of intricate heterogeneity of T cell receptor clonotypes, usage of V/J segments in ex vivo–expanded tumor-infiltrating lymphocytes from breast cancer patients is stable.
· Diversity of T cell receptor clonotypes seems to be decreased during ex vivo rapid expansion protocol.
Rights and permissions
About this article
Cite this article
Park, I.A., Rajaei, H., Kim, YA. et al. T cell receptor repertoires of ex vivo–expanded tumor-infiltrating lymphocytes from breast cancer patients. Immunol Res 68, 233–245 (2020). https://doi.org/10.1007/s12026-020-09150-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-020-09150-8