Abstract
Background
Human embryonic stem cell (hESC)-derived endothelial cells (ECs) possess therapeutic potential in many diseases. Cytokine supplementation induction and transcription factor overexpression have become two mainstream methods of hESC-EC induction. Single-cell RNA-seq technology has been widely used to analyse dynamic processes during hESC-EC induction and components of induced endothelial cells. However, studies that used single-cell RNA-seq are mainly based on cytokine supplementation methods. In this study, we used a high-efficiency human embryonic stem cell-endothelial cell line (hESC-EC) called the “FLI1-PKC system” as a research model and employed single-cell RNA sequencing (scRNA-seq) to investigate the transcriptional landscape and cellular dynamics.
Methods
The high-efficiency hESC-EC induction (FLI1-PKC) system was established in our previous study. We applied single-cell RNA sequencing (scRNA-seq) of the differentiated cells at different time points and investigated the gene expression profiles.
Results
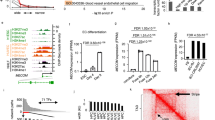
The FLI1-PKC induction system can directionally differentiate hESCs into mature endothelial cells with all the requisite functions. Unlike other hES-EC induction protocols, the FLI1-PKC method follows a different induction route; nonetheless, the transcriptome of induced endothelial cells (iECs) remains the same. The elevated number of activated transcription factors may explain why the FLI1-PKC system is more effective than other hES-EC protocols.
Conclusion
Our study has presented a single-cell transcriptional overview of a high-efficiency hESC-EC induction system, which can be used as a model and reference for further improvement in other hESC induction systems.
Graphical Abstract







Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- hESC :
-
Human embryonic cells
- ECs :
-
Endothelial cells
- iEC :
-
Induced endothelial cells
- TF :
-
Transcriptional factor
- UMAP :
-
Uniform Manifold Approximation and Projection
References
Yamanaka, S. (2020). Pluripotent stem cell-based cell therapy-promise and challenges. Cell Stem Cell, 27(4), 523–531.
Balboa, D., et al. (2022). Functional, metabolic and transcriptional maturation of human pancreatic islets derived from stem cells. Nature Biotechnology, 40(7), 1042–1055.
Liu, M., et al. (2020). A hepatocyte differentiation model reveals two subtypes of liver cancer with different oncofetal properties and therapeutic targets. Proceedings of the National Academy of Sciences of the United States of America, 117(11), 6103–6113.
Patsch, C., et al. (2015). Generation of vascular endothelial and smooth muscle cells from human pluripotent stem cells. Nature Cell Biology, 17(8), 994–1003.
Van Gelder, R. N., et al. (2022). Regenerative and restorative medicine for eye disease. Nature Medicine, 28(6), 1149–1156.
Lancaster, M., et al. (2022). Anniversary reflections: Inspiring discoveries and the future of the field. Cell Stem Cell, 29(6), 879–881.
Bibli, S. I., et al. (2021). Mapping the endothelial cell s-sulfhydrome highlights the crucial role of integrin sulfhydration in vascular function. Circulation, 143(9), 935–948.
Colliva, A., et al. (2020). Endothelial cell-cardiomyocyte crosstalk in heart development and disease. Journal of Physiology, 598(14), 2923–2939.
Liang, J., et al. (2017). Inhibition of microRNA-495 enhances therapeutic angiogenesis of human induced pluripotent stem cells. Stem Cells, 35(2), 337–350.
Zhang, F., et al. (2017). Optimizing mesoderm progenitor selection and three-dimensional microniche culture allows highly efficient endothelial differentiation and ischemic tissue repair from human pluripotent stem cells. Stem Cell Research & Therapy, 8(1), 6.
Gao, L., et al. (2017). Myocardial tissue engineering with cells derived from human-induced pluripotent stem cells and a native-like, high-resolution, 3-dimensionally printed scaffold. Circulation Research, 120(8), 1318–1325.
Du, C., et al. (2014). Induced pluripotent stem cell-derived hepatocytes and endothelial cells in multi-component hydrogel fibers for liver tissue engineering. Biomaterials, 35(23), 6006–6014.
Abaci, H. E., et al. (2016). Human skin constructs with spatially controlled vasculature using primary and iPSC-derived endothelial cells. Advanced Healthcare Materials, 5(14), 1800–1807.
Huang, N. F., et al. (2010). Embryonic stem cell-derived endothelial cells engraft into the ischemic hindlimb and restore perfusion. Arteriosclerosis, Thrombosis, and Vascular Biology, 30(5), 984–991.
Umeda, K., et al. (2006). Identification and characterization of hemoangiogenic progenitors during cynomolgus monkey embryonic stem cell differentiation. Stem Cells, 24(5), 1348–1358.
Rufaihah, A. J., et al. (2011). Endothelial cells derived from human iPSCS increase capillary density and improve perfusion in a mouse model of peripheral arterial disease. Arteriosclerosis, Thrombosis, and Vascular Biology, 31(11), e72–e79.
Harding, A., et al. (2017). Highly efficient differentiation of endothelial cells from pluripotent stem cells requires the MAPK and the PI3K pathways. Stem Cells, 35(4), 909–919.
Zhao, H., et al. (2018). FLI1 and PKC co-activation promote highly efficient differentiation of human embryonic stem cells into endothelial-like cells. Cell Death & Disease, 9(2), 131.
Spevak, C. C., et al. (2020). Hematopoietic stem and progenitor cells exhibit stage-specific translational programs via mTOR- and CDK1-dependent mechanisms. Cell Stem Cell, 26(5), 755-765.e7.
Stark, R., Grzelak, M., & Hadfield, J. (2019). RNA sequencing: The teenage years. Nature Reviews Genetics, 20(11), 631–656.
Ranzoni, A. M., et al. (2021). Integrative single-cell RNA-Seq and ATAC-Seq analysis of human developmental hematopoiesis. Cell Stem Cell, 28(3), 472-487.e7.
McCracken, I. R., et al. (2020). Transcriptional dynamics of pluripotent stem cell-derived endothelial cell differentiation revealed by single-cell RNA sequencing. European Heart Journal, 41(9), 1024–1036.
Wang, M., et al. (2018). Single-cell RNA sequencing analysis reveals sequential cell fate transition during human spermatogenesis. Cell Stem Cell, 23(4), 599-614.e4.
Lang, C., et al. (2019). Single-cell sequencing of iPSC-dopamine neurons reconstructs disease progression and identifies HDAC4 as a regulator of Parkinson cell phenotypes. Cell Stem Cell, 24(1), 93-106.e6.
Wu, T. D., & Nacu, S. (2010). Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics, 26(7), 873–881.
Lun, A. T., Bach, K., & Marioni, J. C. (2016). Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biology, 17, 75.
MacAskill, M. G., et al. (2018). Robust revascularization in models of limb ischemia using a clinically translatable human stem cell-derived endothelial cell product. Molecular Therapy, 26(7), 1669–1684.
Zhao, H., et al. (2020). VEGF promotes endothelial cell differentiation from human embryonic stem cells mainly through PKC-ɛ/η pathway. Stem Cells and Development, 29(2), 90–99.
Lambert, S. A., et al. (2018). The human transcription factors. Cell, 172(4), 650–665.
Han, M., & Zhou, B. (2020). Sox17 and coronary arteriogenesis in development. Circulation Research, 127(11), 1381–1383.
Yang, Y., et al. (2022). Endothelial MEKK3-KLF2/4 signaling integrates inflammatory and hemodynamic signals during definitive hematopoiesis. Blood, 139(19), 2942–2957.
Kong, Y., et al. (2020). circNFIB1 inhibits lymphangiogenesis and lymphatic metastasis via the miR-486-5p/PIK3R1/VEGF-C axis in pancreatic cancer. Molecular Cancer, 19(1), 82.
Bucher, K., et al. (2000). The T cell oncogene Tal2 is necessary for normal development of the mouse brain. Developmental Biology, 227(2), 533–544.
Wang, R., Guo, S., & Yang, L. (2023). Tal2 is required for generation of GABAergic neurons in the zebrafish midbrain. Developmental Dynamics, 252(2), 263–275.
Mori, S., et al. (1999). The leukemic oncogene tal-2 is expressed in the developing mouse brain. Brain Research Molecular Brain Research, 64(2), 199–210.
Yang, L., et al. (2021). 3D genome alterations associated with dysregulated HOXA13 expression in high-risk T-lineage acute lymphoblastic leukemia. Nature Communications, 12(1), 3708.
Murthi, P., et al. (2008). Novel homeobox genes are differentially expressed in placental microvascular endothelial cells compared with macrovascular cells. Placenta, 29(7), 624–630.
Fish, J. E., et al. (2017). Dynamic regulation of VEGF-inducible genes by an ERK/ERG/p300 transcriptional network. Development, 144(13), 2428–2444.
Testori, J., et al. (2011). The VEGF-regulated transcription factor HLX controls the expression of guidance cues and negatively regulates sprouting of endothelial cells. Blood, 117(9), 2735–2744.
Kawahara, M., et al. (2012). H2.0-like homeobox regulates early hematopoiesis and promotes acute myeloid leukemia. Cancer Cell, 22(2), 194–208.
Bonifer, C., et al. (2017). Runx1 structure and function in blood cell development. Advances in Experimental Medicine and Biology, 962, 65–81.
Collin, M., Dickinson, R., & Bigley, V. (2015). Haematopoietic and immune defects associated with GATA2 mutation. British Journal of Haematology, 169(2), 173–187.
Daly, M. E. (2017). Transcription factor defects causing platelet disorders. Blood Reviews, 31(1), 1–10.
Spitz, F., & Furlong, E. E. (2012). Transcription factors: From enhancer binding to developmental control. Nature Reviews Genetics, 13(9), 613–626.
Thoms, J. A. I., et al. (2021). Disruption of a GATA2-TAL1-ERG regulatory circuit promotes erythroid transition in healthy and leukemic stem cells. Blood, 138(16), 1441–1455.
Zhou, G., et al. (2015). Zbtb20 regulates the terminal differentiation of hypertrophic chondrocytes via repression of Sox9. Development, 142(2), 385–393.
Cioffi, S., et al. (2014). Tbx1 regulates brain vascularization. Human Molecular Genetics, 23(1), 78–89.
Ji, Z., et al. (2021). Oct4-dependent FoxC1 activation improves the survival and neovascularization of mesenchymal stem cells under myocardial ischemia. Stem Cell Research & Therapy, 12(1), 483.
Cooley, B. C., et al. (2014). TGF-β signaling mediates endothelial-to-mesenchymal transition (EndMT) during vein graft remodeling. Science Translational Medicine, 6(227), 227ra34.
Wernig, M., et al. (2007). In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature, 448(7151), 318–324.
Hochedlinger, K. (2010). From MYOD1 to iPS cells. Nature Reviews Molecular Cell Biology, 11(12), 817.
Acknowledgements
We thank Genergy Bio-Technology (Shanghai) Co. for providing sequencing service and helping with data analyses.
Funding
This research was supported by grants from the National Natural Science Foundation of China (Grant number: 81873478) and Hunan Provincial Grant for Innovative Province Construction (Grant number: 2019SK4012). The funding body played no role in the design of the study and collection, analysis, and interpretation of data.
Author information
Authors and Affiliations
Contributions
LH designed the study and revised the manuscript. XWX performed most of the experiments. HZ build the FLI1-PKC hES-EC induction system and provide technical support for the experiments. GL revised the manuscript. JRC analyzed the single-cell sequencing data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
(1) Title of the approved project: Stem cell bank of the institution of National Engineering and Research Center of Human Stem Cell; (2) Name of the institutional approval committee or unit: ethical committee of Reproductive and Genetic Hospital of CITIC-Xiangya; (3) Approval number: LL-SC-2022–031; (4) Date of approval: 8 October 2022.
Consent for Publication
Not applicable.
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.



Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, X., Chen, J., Zhao, H. et al. Single-Cell RNA-seq Analysis of a Human Embryonic Stem Cell to Endothelial Cell System Based on Transcription Factor Overexpression. Stem Cell Rev and Rep 19, 2497–2509 (2023). https://doi.org/10.1007/s12015-023-10598-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-023-10598-y