Abstract
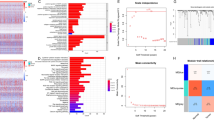
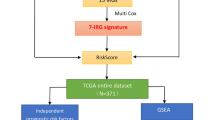
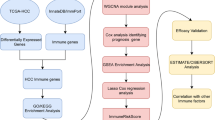
Though patients with hepatocellular carcinoma (HCC) benefit from the treatment of immune checkpoint inhibitor (ICB), it is still of vital significance to develop more effective drugs and predict patients’ response to ICB therapy. Herein, we utilized single sample gene set enrichment analysis (ssGSEA) to score the downloaded tumor samples from TCGA-LIHC based on 29 immune gene sets, thus reflecting the immunologic competence of samples. Then samples were classified into high, moderate, and low immunity groups. Additionally, we utilized survival analysis and ESTIMATE score to verify the reliability of the immunity grouping. We then performed differential expression analysis on the samples in these two groups and obtained 716 differentially expressed genes (DEGs). Next, the DEGs mentioned above were subjected to GO and KEGG analyses. The outcomes demonstrated that these DEGs were mostly correlated with the immune-related biological functions. To further verify biological processes in which DEGs might be involved, we constructed a protein–protein interaction network. Afterward, we used MCODE plugin to conduct subnetwork analysis. Thereafter, KEGG enrichment analysis was performed on two genes with the highest score in the subnetwork. The results exhibited that these genes were gathered in pathways such as Th1 and Th2 cell differentiation and NF-κB. Finally, we utilized Connectivity Map to find possible drugs for the treatment of HCC and obtained complex methyl-angolensate. The above results may contribute to distinguishing HCC patients who are eligible for immunotherapy and providing the foundations for the development of therapeutic drugs for HCC.





Similar content being viewed by others
Data Availability
The data used to support the findings of this study are included within the article. The data and materials in the current study are available from the corresponding author on reasonable request.
References
Siegel, R. L., Miller, K. D. & Jemal, A. (2020). Cancer statistics. CA: A Cancer Journal For Clinicians, 70, 7–30. https://doi.org/10.3322/caac.21590.
Cong, Y. et al. (2018) Dual drug backboned shattering polymeric theranostic nanomedicine for synergistic eradication of patient-derived lung cancer. Advanced Materials. 30, https://doi.org/10.1002/adma.201706220.
Hou, J., Zhang, H., Sun, B., & Karin, M. (2020). The immunobiology of hepatocellular carcinoma in humans and mice: basic concepts and therapeutic implications. Journal of Hepatology, 72, 167–182. https://doi.org/10.1016/j.jhep.2019.08.014.
Binnewies, M., et al. (2018). Understanding the tumor immune microenvironment (TIME) for effective therapy. Nature Medicine, 24, 541–550. https://doi.org/10.1038/s41591-018-0014-x.
Smyth, M. J., Ngiow, S. F., Ribas, A., & Teng, M. W. (2016). Combination cancer immunotherapies tailored to the tumour microenvironment. Nature Reviews Clinical Oncology, 13, 143–158. https://doi.org/10.1038/nrclinonc.2015.209.
Racanelli, V., & Rehermann, B. (2006). The liver as an immunological organ. Hepatology, 43, S54–62. https://doi.org/10.1002/hep.21060.
Prieto, J., Melero, I., & Sangro, B. (2015). Immunological landscape and immunotherapy of hepatocellular carcinoma. Nature Reviews Gastroenterology Hepatology, 12, 681–700. https://doi.org/10.1038/nrgastro.2015.173.
Cariani, E., & Missale, G. (2019). Immune landscape of hepatocellular carcinoma microenvironment: implications for prognosis and therapeutic applications. Liver International, 39, 1608–1621. https://doi.org/10.1111/liv.14192.
Rui, X., Shao, S., Wang, L., & Leng, J. (2019). Identification of recurrence marker associated with immune infiltration in prostate cancer with radical resection and build prognostic nomogram. BMC Cancer, 19, 1179 https://doi.org/10.1186/s12885-019-6391-9.
Bao, X., et al. (2019). Immune landscape of invasive ductal carcinoma tumor microenvironment identifies a prognostic and immunotherapeutically relevant gene signature. Frontiers in Oncology, 9, 903 https://doi.org/10.3389/fonc.2019.00903.
Subramanian, A., et al. (2017). A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell, 171, 1437–1452 e1417. https://doi.org/10.1016/j.cell.2017.10.049.
Hanzelmann, S., Castelo, R., & Guinney, J. (2013). GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics, 14, 7. https://doi.org/10.1186/1471-2105-14-7.
Zhou, C., et al. (2020). High RPS3A expression correlates with low tumor immune cell infiltration and unfavorable prognosis in hepatocellular carcinoma patients. American Journal of Cancer Research, 10, 2768–2784.
Ringelhan, M., Pfister, D., O’Connor, T., Pikarsky, E., & Heikenwalder, M. (2018). The immunology of hepatocellular carcinoma. Nature Immunology, 19, 222–232. https://doi.org/10.1038/s41590-018-0044-z.
Hu, Y., Sun, H., Zhang, H., & Wang, X. (2020). An Immunogram for an individualized assessment of the antitumor immune response in patients with hepatocellular carcinoma. Frontiers in Oncology, 10, 1189 https://doi.org/10.3389/fonc.2020.01189.
Mlecnik, B., et al. (2016). Integrative analyses of colorectal cancer show immunoscore is a stronger predictor of patient survival than microsatellite instability. Immunity, 44, 698–711. https://doi.org/10.1016/j.immuni.2016.02.025.
Sun, C., et al. (2015). The predictive value of centre tumour CD8(+) T cells in patients with hepatocellular carcinoma: comparison with Immunoscore. Oncotarget, 6, 35602–35615. https://doi.org/10.18632/oncotarget.5801.
Yao, Q., et al. (2017). Prognostic value of immunoscore to identify mortality outcomes in adults with HBV-related primary hepatocellular carcinoma. Medicine (Baltimore), 96, e6735 https://doi.org/10.1097/MD.0000000000006735.
Ng, H. H. M. et al. Immunohistochemical scoring of CD38 in the tumor microenvironment predicts responsiveness to anti-PD-1/PD-L1 immunotherapy in hepatocellular carcinoma. J Immunother Cancer 8, https://doi.org/10.1136/jitc-2020-000987 (2020).
Owen, K. L., Brockwell, N. K. & Parker, B. S. (2019) JAK-STAT signaling: a double-edged sword of immune regulation and cancer progression. Cancers (Basel) 11, https://doi.org/10.3390/cancers11122002.
Lu, X., et al. (2018). The anti-inflammatory NHE-06 restores antitumor immunity by targeting NF-kappaB/IL-6/STAT3 signaling in hepatocellular carcinoma. Biomedicine & Pharmacotherapy, 102, 420–427. https://doi.org/10.1016/j.biopha.2018.03.099.
Chiruvella, K. K., Panjamurthy, K., Choudhary, B., Joy, O., & Raghavan, S. C. (2010). Methyl angolensate from callus of Indian redwood induces cytotoxicity in human breast cancer cells. International Journal of Biomedical Science, 6, 182–194.
Chiruvella, K. K., & Raghavan, S. C. (2011). A natural compound, methyl angolensate, induces mitochondrial pathway of apoptosis in Daudi cells. Investigational New Drugs, 29, 583–592. https://doi.org/10.1007/s10637-010-9393-7.
Funding
This study received support from Fuzhou Key Subject Project (No. 201912002), the Key Project of the Guidance of Fujian Province (No. 2019Y0068) and the Surface Project of Natural Science Foundation of Fujian Province (No. 2020J011144). The funders have no role in any process of this study.
Author information
Authors and Affiliations
Contributions
XH: conceptualization, methodology, software. HH: data curation, writing- original draft preparation. JS: visualization, investigation. XZ: supervision. YJ: software, validation. LL: writing- reviewing and editing. SD: writing - review & editing, project administration. All the authors have approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Huang, X., Hu, H., Liu, J. et al. Immune Analysis and Small Molecule Drug Prediction of Hepatocellular Carcinoma Based on Single Sample Gene Set Enrichment Analysis. Cell Biochem Biophys 80, 427–434 (2022). https://doi.org/10.1007/s12013-022-01070-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-022-01070-8