Abstract
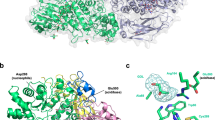
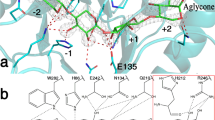
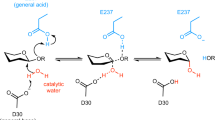
We report the X-ray crystal structure of a glycoside hydrolase family 43 β-xylosidase, RS223BX, which is strongly activated by the addition of divalent metal cations. The 2.69 Å structure reveals that the Ca2+ cation is located at the back of the active-site pocket. The Ca2+ is held in the active site by the carboxylate of D85, an “extra” acid residue in comparison to other GH43 active sites. The Ca2+ is in close contact with a histidine imidazole, which in turn is in contact with the catalytic base (D15) thus providing a mechanism for stabilizing the carboxylate anion of the base and achieve metal activation. The active-site pocket is mirrored by an “inactive-site” pocket of unknown function that resides on the opposite side of the monomer.





Similar content being viewed by others
Abbreviations
- 4NPX:
-
4-nitrophenyl-β-D-xylopyranoside
- GH43:
-
glycoside hydrolase family 43
- M2+ :
-
divalent metal cation
- RS223BX:
-
β-xylosidase of this study
- X2:
-
xylobiose
- X3:
-
xylotriose
- X4:
-
xylotetraose
References
Jordan, D. B., Lee, C. C., Wagschal, K., & Braker, J. D. (2013). Activation of a GH43 β-xylosidase by divalent metal cations: slow binding of divalent metal and high substrate specificity. Archives of Biochemistry and Biophysics, 533, 79–87.
Lee, C. C., Braker, J. D., Grigorescu, A. A., Wagschal, K., & Jordan, D. B. (2013). Divalent metal activation of a GH43 β-xylosidase. Enzyme and Microbial Technology, 52, 84–90.
Jordan, D. B., Wagschal, K., Grigorescu, A. A., & Braker, J. D. (2013). Highly active β-xylosidases of glycoside hydrolase family 43 operating on natural and artificial substrates. Applied Microbiology and Biotechnology, 97, 4415–4428.
Brunzelle, J. S., Jordan, D. B., McCaslin, D. R., Olczak, A., & Wawrzak, Z. (2008). Structure of the two-subsite β-D-xylosidase from Selenomonas ruminantium in complex with 1,3-bis[tris(hydroxymethyl)methylamino]propane. Archives of Biochemistry and Biophysics, 474, 157–166.
Brüx, C., Ben-David, A., Shallom-Shezifi, D., Leon, M., Niefind, K., Shoham, G., et al. (2006). The structure of an inverting GH43 β-xylosidase from Geobacillus stearothermophilus with its substrate reveals the role of the three catalytic residues. Journal of Molecular Biology, 359, 97–109.
Otwinowski, Z., & Minor, W. (1997). Processing of X-ray diffraction data collected in oscillation mode. In C. W. Carter Jr. (Ed.), Methods in enzymology (pp. 307–326). Academic Press.
Keegan, R. M., & Winn, M. D. (2007). Automated search-model discovery and preparation for structure solution by molecular replacement. Acta Crystallographica Section D, 63, 447–457.
Vagin, A., & Teplyakov, A. (1997). MOLREP: an automated program for molecular replacement. Journal of Applied Crystallography, 30, 1022–1025.
Murshudov, G. N., Vagin, A. A., & Dodson, E. J. (1997). Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallographica Section D, 53, 240–255.
Winn, M. D., Ballard, C. C., Cowtan, K. D., Dodson, E. J., Emsley, P., Evans, P. R., et al. (2011). Overview of the CCP4 suite and current developments. Acta Crystallographica Section D, 67, 235–242.
Cartmell, A., McKee, L. S., Peña, M. J., Larsbrink, J., Brumer, H., Kaneko, S., et al. (2011). The structure and function of an arabinan-specific α-1,2-arabinofuranosidase identified from screening the activities of bacterial GH43 glycoside hydrolases. Journal of Biological Chemistry, 286, 15483–15495.
Emsley, P., Lohkamp, B., Scott, W. G., & Cowtan, K. (2010). Features and development of coot. Acta Crystallographica Section D, 66, 486–501.
Adams, P. D., Afonine, P. V., Bunkoczi, G., Chen, V. B., Davis, I. W., Echols, N., et al. (2010). PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallographica Section D, 66, 213–221.
Durette, P.L., & Horton, D. (1971). Conformational analysis of sugars and their derivatives. In R.S. Tipson, H. Derek (Eds.), Adv Carbohydr Chem Biochem (pp. 49–125). Academic Press.
Jordan, D. B. (2008). β-D-Xylosidase from Selenomonas ruminantium: catalyzed reactions with natural and artificial substrates. Applied Biochemistry and Biotechnology, 146, 137–149.
Rajan, S. S., Yang, X., Collart, F., Yip, V. L., Withers, S. G., Varrot, A., et al. (2004). Novel catalytic mechanism of glycoside hydrolysis based on the structure of an NAD+/Mn2+ -dependent phospho-α-glucosidase from Bacillus subtilis. Structure, 12, 1619–1629.
Varrot, A., Yip, V. L., Li, Y., Rajan, S. S., Yang, X., Anderson, W. F., et al. (2005). NAD+ and metal-ion dependent hydrolysis by family 4 glycosidases: structural insight into specificity for phospho-β-D-glucosides. Journal of Molecular Biology, 346, 423–435.
Yang, W., Lee, H. W., Hellinga, H., & Yang, J. J. (2002). Structural analysis, identification, and design of calcium-binding sites in proteins. Proteins: Structure, Function, and Bioinformatics, 47, 344–356.
de Sanctis, D., Inácio, J. M., Lindley, P. F., de Sá-Nogueira, I., & Bento, I. (2010). New evidence for the role of calcium in the glycosidase reaction of GH43 arabinanases. FEBS Journal, 277, 4562–4574.
Acknowledgments
This work was supported in part by the United States Department of Agriculture CRIS 3620-41000-133-00D (D.B.J. and J.D.B.) and CRIS 5325-41000-049-00 (K.W, C.C.L, and V.J.C). Mention of trade names or commercial products in this report is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
A figure of some of the xylotetraose contacts within 4 Å of RS223BX, table of closest residue-residue pair contacts between subunits that are not part of the same dimer and could help form a tetramer, electron density map of a portion of the RS223BX active site, a 3D overlay of the RS223BX (this work) and the arabinanase (PDB ID 2X8F) structures, and an alignment of amino acid sequences of the four GH43 enzymes discussed in this work. (DOCX 1898 kb)
Rights and permissions
About this article
Cite this article
Jordan, D.B., Braker, J.D., Wagschal, K. et al. X-ray Crystal Structure of Divalent Metal-Activated β-xylosidase, RS223BX. Appl Biochem Biotechnol 177, 637–648 (2015). https://doi.org/10.1007/s12010-015-1767-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1767-z