Abstract
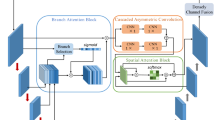
Accurate segmentation and detection of leaf area is of great significance for automatic plant growth monitoring and is one of the most effective strategies to monitor the sustainability of plant growth, yield, width, and height of plants for agricultural production. The leaf segmentation in carrot plants is a challenging task because of secondary leaflets, which introduces complex variability in leaf shape. In this paper, a novel AWUNet (Attention-gated Wavelet pooled UNet) integrating the concepts of wavelet pooling to reduce the size of the feature map and attention gate module which focuses the semantic content in the feature map is proposed. The skip connections in existing UNet (U-shaped network) model are remodeled using attention gate mechanism for saliency improvement, and pooling layers utilize wavelet function for compression. The effectiveness of hyper parameters is investigated to enhance the proposed AWUNet model’s accuracy. The proposed model was evaluated and compared with existing methods such as UNet + + , Inception UNet, SAUNet, and INCSA UNet, as well as pre-trained deep learning architectures like Visual Geometry Group models such as VGG16-UNet, VGG19-UNet, and residual neural network (ResNet) UNet. The highest segmentation IoU (intersection of union) score of 94.81% was achieved by the proposed AWUNet model, whereas the state-of-the-art methods UNet, UNet + + , Inception UNet, SAUNet, INCSA UNet,VGG16 UNet,VGG19 UNet, and ResNet UNet achieved IoU score of 86.29%, 63.37%, 45.26%, 72.39%, 89.30%, 82.99%, 85.16%, and 75.50%, respectively. The findings showed that the proposed AWUNet model outperforms than other models in segmenting leaf area in plants and can be implemented in the agriculture sector for crop monitoring.












Similar content being viewed by others
Data availability
The datasets used in the study are publically available in the repository: https://github.com/cwfid/dataset
Change history
28 February 2023
A Correction to this paper has been published: https://doi.org/10.1007/s11760-023-02508-z
References
Pieruschka, R., Schurr, U.: Plant phenotyping: past, present, and future. Plant Phenomics. (2019). https://doi.org/10.1155/2019/7507131
Singh, V., Misra, A.K.: Detection of plant leaf diseases using image segmentation and soft computing techniques. Inf. Process. Agricult. (2017). https://doi.org/10.1016/j.inpa.2016.10.005
Minervini, M., Giuffrida, M.V., Perata, P., Tsaftaris, S.A.: Phenotiki: an open software and hardware platform for affordable and easy image-based phenotyping of rosette-shaped plants. Plant J. (2017). https://doi.org/10.1111/tpj.13472
Castillo-Martínez, M., Funes, F., Carvajal-Gámez, B., Urriolagoitia-Sosa, G., Rosales, A.: Color index based thresholding method for background and foreground segmentation of plant images. Comput. Electron. Agric. (2020). https://doi.org/10.1016/j.compag.2020.105783
Sengar, N., Dutta, M., Travieso, C.: Computer vision based technique for identification and quantification of powdery mildew disease in cherry leaves. Computing (2018). https://doi.org/10.5555/3288338.3288342
Riehle, D., Reiser, D., Griepentrog, H.W.: Robust index-based semantic plant/background segmentation for RGB- images. Comput. Electron. Agric. 169, 1–12 (2020). https://doi.org/10.1016/j.compag.2019.105201
Kumar, J.P., Domnic, S.: Image based Leaf segmentation and counting in Rosette plants. Inf. Process. Agricul (2019). https://doi.org/10.1016/j.inpa.2018.09.005
Wang, Z., Wang, K., Yang, F., Pan, S., Han, H.: Image segmentation of overlapping leaves based on Chan-vese model and sobel operator. Information Processing in Agriculture (2018). https://doi.org/10.1016/j.inpa.2017.09.005
Ye, M., Cao, Z.-G., Yu, Z., Bai, X.D.: Crop feature extraction from images with probabilistic superpixel Markov random field. Comput. Electron. Agric. (2015). https://doi.org/10.1016/j.compag.2015.04.010
Zhou, J., Zhou, J.: Automated segmentation of soybean plants from 3D point cloud using machine learning. Comput. Electron. Agric. (2019). https://doi.org/10.1016/j.compag.2019.04.014
Pravin Kumar, S.K., Sumithra, M.G., Saranya, N.: Artificial bee colony-based fuzzy c means (ABC-FCM) segmentation algorithm and dimensionality reduction for leaf disease detection in bioinformatics. J Supercomput (2019). https://doi.org/10.1007/s11227-019-02999-z
Smith, A.G., Petersen, J., Selvan, R., et al.: Segmentation of roots in soil with UNet. Plant Methods 16, 1–15 (2020). https://doi.org/10.1186/s13007-020-0563-0
Minaee, S., Boykov, Y.Y., Porikli, F., Plaza, A.J., Kehtarnavaz, N., Terzopoulos, D.: Image segmentation using deep learning: a survey. IEEE Trans. Pattern Anal. Mach. Intell. (2021). https://doi.org/10.1109/TPAMI.2021.3059968
Zhou, Z., Rahman Siddiquee, M.M., Tajbakhsh, N. and Liang, J.: UNet++: a nested unet architecture for medical image segmentation. In: Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Lecture Notes in Computer Science, Springer Cham, pp. 3-11 (2018). doi:https://doi.org/10.1007/978-3-030-00889-5_1
Punn, N., Agarwal, S.: Inception U-net architecture for semantic segmentation to identify nuclei in microscopy cell images. ACM Trans. Multimed. Comput. Commun. Appl. 16, 1–15 (2020). https://doi.org/10.1145/3376922
Guo, C., Szemenyei, M., Pei, Y., Yi, Y., Zhou, W.: SD-Unet: A Structured Dropout U-Net for Retinal Vessel Segmentation, pp 439–444 (2019). https://doi.org/10.1109/BIBE.2019.00085.
Delibasoğlu, İ: INCSA-UNET: Spatial attention inception UNET for aerial images segmentation. Comput. Inf. 40(6), 1244–1262 (2022). https://doi.org/10.31577/cai_2021_6_1244
Agarwal, M., Gupta, S.K., Biswas, K.K.: Plant leaf disease segmentation using compressed UNet architecture. In: Gupta, M., Ramakrishnan, G. (eds.) Trends and Applications in Knowledge Discovery and Data Mining PAKDD. Lecture Notes in Computer Science, pp. 9–14. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-75015-2_2
Williams, T., Li, R.: Advanced Image Classification Using Wavelets and Convolutional Neural Networks. https://doi.org/10.1109/ICMLA. (2016)
Allison, R., Wenjin Z.: Improving classification with CNNs using wavelet pooling with Nesterov-accelerated Adam, In: Proceedings of 11th International Conference on Bioinformatics and Computational Biology, EPiC Series in Computing, pp. 84–93 (2019), https://doi.org/10.29007/9c5j
Ozan, O., Jo, S., Bernhard, K., Ben, G.: Attention UNet: Learning Where to Look for the Pancreas, (2018). https://doi.org/10.48550/arXiv.1804.03999.
Kang, J., Liu, L., Zhang, F., Shen, C., Wang, N., Shao, L.: Semantic segmentation model of cotton roots in-situ image based on attention mechanism. Comput. Electron. Agric. (2021). https://doi.org/10.1016/j.compag.2021.106370
Jo, S., Ozan, O., Michiel, S., Mattias, H., Bernhard, K., Ben, G., Daniel, R.: Attention gated networks: Learning to leverage salient regions in medical images. Med. Image Anal. (2019). https://doi.org/10.1016/j.media.2019.01.012
Sambasivam, G., Opiyo, G.D.: A predictive machine learning application in agriculture: Cassava disease detection and classification with imbalanced dataset using Convolutional neural networks. Egyptian Inf. J. (2021). https://doi.org/10.1016/j.eij.2020.02.007
Jalal Uddin, Md., Yubin Li, Md., Sattar, A., Most, Z., Nasrin, C.L.: Effects of learning rates and optimization algorithms on forecasting accuracy of hourly typhoon rainfall: experiments with convolutional neural network. Earth Space Sci. (2022). https://doi.org/10.1029/2021EA002168
Ronneberger, O., Fischer, P., Brox, T.: UNet: Convolutional Networks for Biomedical Image Segmentation, pp.234–241 (2015). https://doi.org/10.1007/978-3-319-24574-4_28.
Bai, Y., Guo, Y., Zhang, Q., Cao, B., Zhang, B.: Multi-network fusion algorithm with transfer learning for green cucumber segmentation and recognition under complex natural environment. Comput. Electron. Agricult. (2022). https://doi.org/10.1016/j.compag.2022.106789
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. (2014) http://arxiv.org/abs/1409.1556.
Ghosh, S., Chaki, A., Santosh, K.: Improved UNet architecture with VGG-16 for brain tumor segmentation. Phys. Eng. Sci. Med. 44(703–712), 919–995 (2021). https://doi.org/10.1007/s13246-021-01019-w
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition, pp. 770–778 (2016). https://doi.org/10.1109/CVPR.2016.90.
Mushtaq, M., Akram, M.U., Alghamdi, N.S., Fatima, J., Masood, R.F.: Localization and edge-based segmentation of lumbar spine vertebrae to identify the deformities using deep learning models. Sensors (Basel) 22(4), 1547 (2022). https://doi.org/10.3390/s22041547
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
All authors carried out the literature review. Ms A. Shamim Banu conducted the experiments. Dr. S. Deivalakshmi reviewed the manuscript and approved the final version of manuscript after peer review. All the authors approved to submit the manuscript in the Journal tilted “Signal, Image and Video Processing.”
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Banu, A.S., Deivalakshmi, S. AWUNet: leaf area segmentation based on attention gate and wavelet pooling mechanism. SIViP 17, 1915–1924 (2023). https://doi.org/10.1007/s11760-022-02403-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11760-022-02403-z