Abstract
No approved vaccine exists for Klebsiella pneumoniae yet. Outer membrane protein-K17 (OMPK17) is involved in K. pneumoniae pathogenesis. No information has been found about OMPK17 dominant epitopes in the literature. Therefore, this study aimed to predict both T cell and B cell epitopes of K. pneumoniae OMPK17 via immunoinformatics approaches. Both T cell (class-I and II) and B cell (linear and discontinuous) epitopes of OMPK17 were predicted. Several screening analyses were performed including clustering, immunogenicity, human similarity, toxicity, allergenicity, conservancy, docking, and structural/physicochemical suitability. The results showed that some regions of OMPK17 have more potential as epitopes. The most possible epitopes were found via several analyses including the selection of higher-scoring epitopes, the epitopes predicted with more tools, more immunogenic epitopes, the epitopes capable of producing interferon-gamma, the epitopes with more dissimilarity to human peptides, and non-toxic and non-allergenic epitopes. By comparing the best T cell and B cell epitopes, we reached a 25-mer peptide containing both T cell (class-I and class-II) and B cell (linear) epitopes and comprising appropriate physicochemical characteristics that are required for K. pneumoniae vaccine development. The in vitro/in vivo study of this peptide is recommended to clarify its actual efficiency and efficacy.
Similar content being viewed by others
Introduction
Klebsiella pneumoniae is a Gram-negative, encapsulated bacterium often found in hospital settings, especially in intensive care, pediatric and surgical wards (Piperaki et al. 2017). This pathogen is primarily an opportunistic pathogen that colonizes human mucous membranes, including the gastrointestinal tract and oropharynx. From these sites, it can invade other tissues and cause pneumonia, sepsis, liver abscesses, meningitis, bacteremia, urinary tract infections, and many other diseases (Piperaki et al. 2017).
In recent years, due to the long-term use of antibacterial drugs, the incidence of multidrug-resistant K. pneumoniae, especially carbapenem-resistant K. pneumoniae, has been increasing (Bassetti et al. 2018; Dai and Hu 2022; Uzairue et al. 2022). The development process of new antibiotics is very slow and cannot reach the development of drug-resistant K. pneumoniae. Therefore, antibacterial programs that bypass the mechanisms of bacterial drug resistance are of particular importance. Vaccines are effective measures to protect against drug-resistant bacterial infections. The vaccines can significantly reduce antibiotic use, slowing down bacterial drug resistance growth (Piperaki et al. 2017; Malachowa et al. 2019; Assoni et al. 2021).
Despite much research to produce an effective vaccine for K. pneumoniae, no vaccine for human use has yet been approved, and efforts are ongoing to find an effective antigen as a vaccine (Pletz et al. 2016; Opoku-Temeng et al. 2022).
Outer membrane proteins (OMPs) are surface trimmers proteins acting as water-filled channels that allow hydrophilic molecules across the membrane. K. pneumoniae OMPs are considered safe subunit vaccines that have been shown to induce host-specific antibodies and have been effective in animal models (Pletz et al. 2016; Zhu et al. 2021).
Outer membrane protein-K17 (OMPK17, also called OMPX) is a member of the OMP family that is involved in K. pneumoniae pathogenesis (Climent et al. 1997). It has been shown that K. pneumoniae OMPK17 can be detected by immune serum from patients with acute K. pneumoniae infection (Kurupati et al. 2006). It has been also shown that this protein can evoke protection against K. pneumoniae in mice, mainly by inducing T helper type 2 immune response, although its real effect on protecting animal models needs to be confirmed (Hussein et al. 2018). Despite these attempts, no information has been found about OMPK17 dominant immunogenic regions (epitopes) in the literature. Immunoinformatics methods are useful in predicting immunogenic epitopes as vaccine candidates and are widely accepted in many scientific fields (De Groot et al. 2001; He et al. 2010; Rahmat Ullah et al. 2021).
This study aimed to predict both T cell and B cell epitopes of K. pneumoniae OMPK17 via immunoinformatics approaches. Also, applying different analyses an antigenic 25-mer peptide containing both T cell and B cell epitopes was introduced for possible use as a vaccine against K. pneumoniae.
Methods
Retrieving of OMPK17 sequence
The one-letter amino acid sequence of K. pneumoniae OMPK17 was obtained from the NCBI protein database (https://www.ncbi.nlm.nih.gov/protein), with the accession number AAA97932.1. The sequence was as follows: “MNKIARLSALAVVLAASVGTTAFAATSTVTGGYAQSDMQGKANKAGGFNLKYRYEQDNNPLGVIGSFTYTEKDNNSNGTYNKGQYYGITAGPAYRLNDWASIYGVVGVGYGKFQNNNYPHKSDMSDYGFSYGAGLQFNPIENVALDFSYEQSRIRNVDVGTWIAGVGYRF”. Domain search using InterPro (https://www.ebi.ac.uk/interpro/) showed that this protein contained one domain (from amino acid no. 11 to no.170), belonging to OMP proteins namely OMP_b-brl (InterPro code: IPR027385).
BLASTing of this sequence by the BLASTP (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins) server showed that it is highly conserved among K. pneumoniae serotypes as well as other Klebsiella species such as K. oxytoca, K. quasipneumoniae, K. michiganensis, K. grimontii, K. variicola, and K. aerogenes. The NCBI conserved domain search (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) showed that our used sequence is the conserved domain of outer membrane protein OmpX belonging to the OM-Channels superfamily.
Prediction of T cell epitopes
Class-I (CD8 + T cell-specific) and class-II (CD4 + T cell-specific) T cell epitopes of OMPK17 were predicted using highly frequent human leukocyte antigen (HLA) alleles (HLA-I for class-I and HLA-II for class-II epitope prediction). These HLA alleles are highly frequent in the world population and have been previously used by us for epitope prediction of other K. pneumoniae antigens (Rostamian et al. 2020; Zargaran et al. 2021; Nemati Zargaran et al., 2021) (Table S1).
The T cell epitopes prediction was done as in our previous works (Akya et al. 2019; Rostamian et al. 2020) using three prediction tools including IEDB (http://tools.iedb.org, (Fleri et al. 2017), SYFPEITHI (http://www.syfpeithi.de/bin/MHCServer.dll/EpitopePrediction.htm, (Schuler et al. 2007)), and ProPred (ProPred-I for class-I and ProPred-II class-II epitope prediction) (http://crdd.osdd.net/raghava/propred/, (Singh and Raghava 2003)). The score of ≥ 10 for SYFPEITHI and ProPred-I, was applied as the thresholds. In ProPred-II, the predicted epitopes were listed without reporting their scores, and all of these epitopes were chosen. In IEDB, the percentile rank of ≤ 1 for class-I and ≤ 10 for class-II epitopes were used as the thresholds. Only epitopes with 9-mer amino acids for class-I and 15-mer length for class-II epitopes were selected to be predicted. Of the predicted epitopes, the sequence similarity was determined and the epitopes were clustered by the IEDB Epitope Cluster Analysis tool (http://tools.iedb.org/cluster/). One epitope in each cluster was chosen for further analyses, which was the epitope with the highest prediction score. For class-I epitopes, the immunogenicity was checked by IEDB Class I immunogenicity tool (http://tools.iedb.org/immunogenicity/) (Calis et al. 2013). For class-II epitopes, the ability in interferon-gamma (IFN-γ) production was checked by the IFNepitope server (http://crdd.osdd.net/raghava/ifnepitope/).
Finding class II/class I windows and selecting final T cell epitopes
Regarding that 9-mer class-I may be placed inside 15-mer class-II epitopes, class II/class I windows were found using the Epitope Cluster Analysis tool. The cut-off was set at ≥ 70% which allows the tool to put the epitopes with ≥ 70% similarity in a unique cluster. Those class-II T cell epitopes that contained at least one class-I T cell epitope were selected.
For selecting the final T cell epitopes, the following criteria were applied: 1- The T cell epitopes containing both class-I and class-II epitopes, 2- The epitopes with higher prediction scores, 3- Being positive in IFN-γ production (for class-II epitopes) or have more immunogenicity score (for class-I epitopes), and 4- The epitopes which were predicted by more prediction tools.
Prediction of B cell epitopes
For predicting B cell linear epitopes seven tools of the IEDB server (http://tools.iedb.org/main/bcell/) provided for B cell linear epitope prediction as well as the ABCpred server (http://crdd.osdd.net/raghava/abcpred/, (Saha and Raghava 2006)) was applied. The linear B cell epitope prediction was done as in our previous work (Zargaran et al. 2021). Briefly, by using the tools’ defaults parameters, the provided charts by each IEDB tool were compared and the epitopes that were predicted by at least three (out of seven) tools were selected. The threshold scores of the tools were as follow: Chou & Fasman Beta-Turn: 1.058, Emini Surface Accessibility: 1.000, Karplus & Schulz Flexibility: 0.998, Kolaskar & Tongaonkar Antigenicity: 1.010, Parker Hydrophilicity: 2.014, Bepipred 1.0: 0.321, Bepipred 2.0: 0.500, and ABCpred: 0.510.
Using the Epitope Cluster Analysis tool, the selected epitopes were then compared and clustered with those predicted by the ABCpred server. The epitopes that were predicted by more tools and/or had higher prediction scores were selected.
For the prediction of discontinuous B cell epitopes, since the crystallographic structure of OMPK17 was not available, it was modeled using the SWISS-MODEL server (https://swissmodel.expasy.org/, (Waterhouse et al. 2018)). The structure of E.coli OMPX (PDB ID: 1QJ8) with 85.62% sequence identity with OMPK17 was used as a template. The quality of the created model was checked by qualitative model energy analysis (QMEAN) scoring and Ramachandran plot generating using the MolProbity server (http://molprobity.biochem.duke.edu/, (Chen et al. 2010)).
The discontinuous B cell epitopes were predicted by two prediction tools provided by IEDB including ElliPro (Ponomarenko et al. 2008) and DiscoTope 2.0 (Kringelum et al. 2012). Using the default parameters, the created model of OMPK17 was presented to the tools and the predicted discontinuous epitopes by both tools were taken as more probable discontinuous B cell epitopes.
Allergenicity, toxicity, and human similarity
All selected T cell-, linear B cell-, and major segments of discontinuous B cell epitopes were evaluated for allergenicity toxicity and human similarity. The allergenicity was checked by the structural database of allergenic proteins (SDAP) (Ivanciuc 2003) and AllerCatPro (Maurer-Stroh et al. 2019) using the tools’ defaults.
The toxicity of the epitopes was checked by the ToxinPred server (https://webs.iiitd.edu.in/raghava/toxinpred/algo.php) (Gupta et al. 2013) remaining with the server parameters as defaults.
The possible human similarity was evaluated by putting the FASTA sequence of the epitopes in the BLASTP server and using the human proteome (taxid 9606) as the organism. Epitopes with more than 90% similarity (both coverage and identity indices) with human peptides were omitted from further analyses.
Detecting the epitopic regions comprising both T cell and linear B cell epitopes
The final selected T cell epitopes (both class-I and class-II T cell epitopes) were compared with linear B cell epitopes by the Epitope Cluster Analysis tool and those containing both T cell and B cell epitopes were selected.
The characteristics of the final epitopic region
The physicochemical characteristics of the final peptide were estimated by ProtParam from Expasy (https://web.expasy.org/protparam/) (Adhikari et al. 2018). The physicochemical characteristics studied included hydrophobicity, molecular weight, grand average of hydropathicity index (GRAVY), net charge, instability index, isoelectric pH (pI), and the estimated half-life. The secondary structure of the final 25-mer peptide was calculated using the Stride Web interface (http://webclu.bio.wzw.tum.de/cgi-bin/stride/stridecgi.py).
The conservancy of the 25-mer epitopic peptide was assessed by NCBI conserved domain search and BLASTP server. The Web3DMol server (http://web3dmol.net/) was applied to draw the structure of OMPK17 and demonstrate the location of the final epitope.
Molecular docking
For further study of HLA-T cell epitope binding, molecular docking studies were also done on T cell epitopes (both class-I and II) of the final 25-mer epitopic peptide using HADDOCK 2.2 (http://haddock.science.uu.nl/services/HADDOCK2.2). The 3D structure of the epitopes was modeled by the PEP-FOLD server (Shen et al. 2014) and validated by generating a Ramachandran plot using the MolProbity server. The 3D structure of HLA alleles was obtained from the PDB databank as follows: HLA-B*35:01 (ID: 2CIK) and HLA-DRB1*15:01 (ID: 1BX2). To exact prediction of residues incorporated in the HLA-epitope binding, consensus prediction of interface residues in transient complexes (CPORT) (https://milou.science.uu.nl/services/CPORT/) was used (de Vries and Bonvin 2011). Following these residues collection, the HADDOCK server ran the docking and delivered the results as HADDOCK scores (as the main result), alongside some other scores (van Zundert et al. 2016). The original ligands of HLA PDB files, previously characterized by crystallography, were used as control.
Results
T cell epitope prediction
Using highly frequent HLA alleles, potential T cell epitopes were predicted by three different tools. The predicted epitopes were evaluated with other analyses to choose the best ones. These analyses include 1- Omitting the duplicate epitopes and selecting one from several similar epitopes, 2- Selecting those with high prediction scores or those predicted by more tools, 3- Considering higher immunogenicity score (for class-I epitopes) or being positive for IFN-γ production (for class-II epitopes) as a favored criterion, and 4- Omitting toxic epitopes, allergens, and those with > 90% human similarity. By applying these criteria, the T cell epitopes were selected and their class II/class I windows were found using the cluster analysis tool. The final three class II/class I windows of T cell epitopes and their analysis results are presented in Table 1.
Linear B epitope prediction
Linear B cell epitope prediction by IEDB tools provided charts indicating regions with scores either above (positive) or below the threshold (negative) (Fig. S1). The positive regions were compared and the regions that had been positive in more tools were chosen as more potential epitopes. Also, the linear B cell epitopes were found by ABCpred and its results were compared and clustered with the epitopes found by IEDB tools. The epitope that was predicted by more tools or had the highest prediction score was chosen. Similar to that was mentioned for T cell epitopes, the linear B cell epitopes were also evaluated for possible toxicity, allergenicity, and human similarity. The top seven linear B cell epitopes and their analysis results are presented in Table 2.
Discontinuous B cell epitope prediction
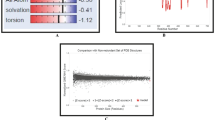
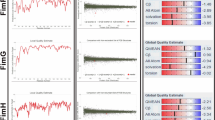
To predict the discontinuous B cell epitopes, initially, OMPK17 was modeled. The model quality was checked by generating Ramachandran and scoring functions provided by the SWISS-MODEL server. Both methods indicated the high quality of the modeling (Fig. 1).
The quality assessment of OMPK17 modeling. The quality of OMPK17 modeling was assessed by scoring functions provided by the SWISS-MODEL server including Global Quality Estimate (GQE) (A), Local Quality Estimate (LQE) (B), and Comparison with non-redundant PDB structures (C) as well as by generating the Ramachandran plot (D). In the GQE plot, the Z-score of several terms (as indicated in the picture) for the model was around zero which shows the better agreement between the model and the experimental structures. In the LQE plot, the majority of the model residues have a score of > 0.6 which means an acceptable quality of modeling. In the comparison plot, the model (indicated by a red star) was close to the grey areas, suggesting a high quality of modeling. Generating the Ramachandran plot (summarized in a table) showed that a high percentage of the model residues are in favored and allowed regions, indicating a high quality of the modeling
DiscoTope 2.0 predicted a discontinuous B cell epitope composed of nine segments of amino acid residues (Table S2). ElliPro tool predicted three discontinuous B cell epitopes, each containing several segments of amino acid residues (Table S2). The epitopic segments were compared and the most dominant segments (those that were longer than four amino acid residues) were presented in Table 2. These segments were also checked for potential toxicity, allergenicity, and human similarity.
The final B/T cell epitopic region
Comparing and clustering T cell and linear B cell epitopes indicated one specific 25-mer region that contained both class-I and class-II T cell epitopes as well as a linear B cell epitope (Table 3). The docking score (HADDOCK score) of the class I epitope-HLA (DMSDYGFSY-HLA B*35:01) was − 63.8±-14.8, which is more negative than the score of the control (class I HLA-original peptide) (-57.3±-10.0).
The HADDOCK score of class II epitope-HLA (HKSDMSDYGFSYGAG-HLA DRB*15:01) was − 27.9±-13.9, which is higher than what was observed for the control (class II HLA-original peptide) (-89.6±-13.9).
The allergenicity and toxicity prediction showed that the peptide is not toxic and not an allergen. The peptide had no ≥ 90% similarity with the human proteome.
The physicochemical characteristics of the final epitopic region (25-mer peptide) are presented in Table 3.
The peptide had a pI equal to 6.74 and a molecular weight of 2848.01 Dalton. Its estimated half-life was 2 min in E. coli, in vivo, 2.8 h in mammalian reticulocytes, in vitro, and 10 min in yeast, in vivo. The peptide is hydrophilic and stable. It is composed of a mixture of strands, helixes, and turns (Table 3) and is located on the lateral surface of the OMPK17 tertiary structure (Fig. 2). The BLASTP results showed that the final 25-mer peptide has a significant sequence identity among Klebsiella species (Table S3). The conservancy assessment showed that the 25-mer peptide is a member of the highly conserved OM-channels (porin) superfamily in Gram-negative bacteria (Fig. S2).
Discussion
The lack of an approved vaccine for K. pneumoniae, along with the therapeutic limitations of this pathogen, led us to find new candidates to be used as vaccines. OMPs are among the most prominent vaccine candidates for Gram-negative bacteria including K. pneumoniae. Previous studies on OMPs, mainly on OMPA, have shown that these proteins can induce protection against K. pneumoniae infections mediated by both types of immune responses including humoral (IgA, IgG1, and IgG2) and cell-mediated immunity (TNFa, nitric oxide, IFNγ, and several interleukins) (Malachowa et al. 2019; Assoni et al. 2021).
One member of the OMPs family that has been less studied but whose immunogenic potential has been previously reported (Hussein et al. 2018) is the OMPK17 protein. This protein was able to induce a Th2-type immune response and a significant IgG1 production which led to protection against K. pneumoniae lethal challenge (Hussein et al. 2018). However, the epitopes of this protein have not been identified clearly so far, so in the current design, we decided to find the B cell and T cell epitopes of the OMPK17 protein using various immunoinformatics methods. The results showed that many regions may play the role of T cell and/or B cell epitopes. However, some of these regions have more potential as epitopes. That’s why we did several analyses to reduce the number of these predicted regions and select the best ones. These analyses included the selection of higher-scoring epitopes, the epitopes predicted with more tools, more immunogenic epitopes, the epitopes capable of producing interferon-gamma, the epitopes with more dissimilarity to human peptides, and non-toxic and non-allergenic epitopes. We also used several tools (instead of one tool) to make sure that the selected regions were epitopes, as it has previously been reported that the use of multiple prediction tools increases the likelihood of selecting epitopes correctly (Trost et al. 2007).
Previous reports have shown that both cellular and humoral immunity are necessary for the complete control of K. pneumoniae (Campbell et al. 1996; Ten Hagen et al. 1998; Pletz et al. 2016). Therefore, if we can design a peptide that contains both type T cell epitope and linear B cell epitope, we can expect this peptide to activate both types of immune responses (Lee et al. 2015). Therefore, in the present study, by comparing the predicted T cell and B cell epitopes, we reached a 25-mer peptide that appears to be able to elicit both types of immune responses. It should be noted that although we predicted the discontinuous B cell epitopes of OMPK17, they were not used for the selection of the final 25-mer epitopic region, because the discontinuous B cell epitopes are dependent on the three-dimensional shape of the protein (Chyau Liang 1998), hence, they are not easily comparable to linear B cell or T cell epitopes.
For a vaccine candidate peptide to be able to pass the in vitro and in vivo experiments successfully and, more importantly, to function properly and produce a proper immune response in humans, it must have a stable structure and a series of appropriate physicochemical properties in addition to being immunogenic. The bioinformatics studies showed that the 25-mer peptide found in the present study is very stable, has a suitable pI, and has an acceptable half-life (about 3 h) in the human body. It is noteworthy that if its immunogenicity is confirmed via studies in animal models and humans, the stability and half-life of this peptide can be increased via various chemical/biochemical methods. The docking studies showed that HLA bind to the class-I epitope of our final 25-mer peptide with higher affinity than the original peptide, demonstrating its suitability to bind the HLA and subsequently elicit immune responses. In contrast, the class-II epitope of our final 25-mer peptide had a low docking score, which could be considered a negative point for eliciting immune responses. However, it should be noted that the control original peptide used here was a peptide of 14 amino acids in length while our class-II peptide had 15 amino acids. This little difference in size may cause the difference between the docking scores of our peptide compared to the original peptide. More docking studies with more appropriate controls seem required in this case.
The index pI is an important factor for the selection of a peptide vaccine and should not be in the range of the body tissues’ pH (pH 7.2 to 7.6) (Wang et al. 2018; Campos-Pinto et al. 2019).
In our 25-mer peptide, the pI was 6.74, which is far from the pH range of body tissues.
The hydrophobicity ratio analysis and the GRAVY index both showed that the 25-mer peptide is more hydrophilic. The negative GRAVY, as seen for our peptide, means being more hydrophilic (Kyte and Doolittle 1982; Adhikari et al. 2018).
Moreover, the instability index of the 25-mer peptide estimated that our epitopic peptide was stable. All these parameters also suggested the peptide as an appropriate vaccine candidate for more in vitro and in vivo experiments.
The conservancy assessment of the 25-mer peptide indicated that it belongs to OM_channels porin) superfamily that is highly conserved among Gram-negative bacteria. Therefore, if this 25-mer peptide is developed as a vaccine, it may elicit immune responses against other Gram-negative bacteria as well, although its real effect required much more studies.
Epitope-based vaccines, such as our proposed 25-mer epitopic peptide, are associated with limited immunity. This limitation may be due to several reasons including the easy degradation of these small-size molecules by proteases in the body, and the difficulty of their recognition by the receptors on the surface of immune cells. One effective way to overcome this limitation is conjugating them to traditional adjuvants and/or new built-in adjuvants such as new biomaterials or carriers. Some of the built-in adjuvants that are proposed for conjugation with epitope-based vaccines include virus-like particle carriers, ligands of pattern recognition receptors, bacterial toxins, nanoparticles, etc. (Lei et al. 2019). Another limitation of our proposed peptide as K. pneumoniae vaccine is that the target population of this vaccine will be mainly elderly patients suffering from K. pneumoniae infections with multiple comorbidities. Accordingly, this population may not be able to evoke strong immune responses (Weinberger et al. 2008). In addition, another issue that is neglected and hence should be assumed as a limitation of our study is that K. pneumoniae vaccines should not cross-react with the intestinal normal flora (Ahmad et al. 2012). Finally, an important issue that was neglected in the present study is that K. pneumoniae has several serotypes, so vaccines against one serotype may be ineffective against others. This issue is recommended to be considered in future research.
Altogether, here we found the high immunogenic class-I/class-II T cell and linear/discontinuous B cell epitopes of K. pneumoniae OMPK17 that can be used in future research. Moreover, the present study introduced a high immunogenic 25-mer peptide containing both B cell and T cell epitopes and comprising appropriate physicochemical characteristics that are required for K. pneumoniae vaccine development. The in vitro/in vivo study of this peptide is recommended to clarify its actual efficiency and efficacy.
Data availability
All the data used for this study are presented in the text and/or figures and tables of the article.
Abbreviations
- CPORT:
-
Consensus prediction of interface residues in transient complexes
- GQE:
-
Global Quality Estimate
- GRAVY:
-
Grand average of hydropathicity index
- HLA:
-
Human leukocyte antigen
- IFN-γ:
-
Inte
rferon-gamma
- LQE:
-
Local Quality Estimate
- OMP:
-
Outer membrane protein
- OMPK17:
-
Outer
membrane protein-K17
- pI:
-
Isoelectric pH
- QMEAN:
-
Qualitative model energy analysis
- SDAP:
-
Structural database of allergenic proteins
References
Adhikari UK, Tayebi M, Rahman MM (2018) Immunoinformatics approach for epitope-based peptide vaccine design and active site prediction against polyprotein of emerging oropouche virus. J Immunol Res 2018:6718083. https://doi.org/10.1155/2018/6718083
Ahmad TA, El-Sayed LH, Haroun M, Hussein AA, El Ashry ESH (2012) Development of immunization trials against Klebsiella pneumoniae Vaccine 30:2411–2420. https://doi.org/10.1016/j.vaccine.2011.11.027
Akya A, Farasat A, Ghadiri K, Rostamian M (2019) Identification of HLA-I restricted epitopes in six vaccine candidates of Leishmania tropica using immunoinformatics and molecular dynamics simulation approaches. Infect Genet Evol 75:103953. https://doi.org/10.1016/j.meegid.2019.103953
Assoni L, Girardello R, Converso TR, Darrieux M (2021) Current stage in the development of Klebsiella pneumoniae vaccines. Infect Dis Ther 10:2157–2175. https://doi.org/10.1007/s40121-021-00533-4
Bassetti M, Righi E, Carnelutti A, Graziano E, Russo A (2018) Multidrug-resistant Klebsiella pneumoniae: challenges for treatment, prevention and infection control. Expert Rev Anti Infect Ther 16:749–761. https://doi.org/10.1080/14787210.2018.1522249
Calis JJ, Maybeno M, Greenbaum JA, Weiskopf D, De Silva AD, Sette A, Kesmir C, Peters B (2013) Properties of MHC class I presented peptides that enhance immunogenicity. PLoS Comput Biol 9:e1003266. https://doi.org/10.1371/journal.pcbi.1003266
Campbell WN, Hendrix E, Cryz S, Cross AS (1996) Immunogenicity of a 24-Valent Klebsiella Capsular Polysaccharide Vaccine and an Eight-Valent Pseudomonas O-Polysaccharide conjugate vaccine administered to victims of acute trauma. Clin Infect Dis 23:179–181. https://doi.org/10.1093/clinids/23.1.179
Campos-Pinto I, Méndez L, Schouten J, Wilkins J, Fedorova M, Pitt AR, Davis P, Spickett CM (2019) Epitope mapping and characterization of 4-hydroxy-2-nonenal modified-human serum albumin using two different polyclonal antibodies. Free Radic Biol Med 144:234–244. https://doi.org/10.1016/j.freeradbiomed.2019.05.008
Chen VB, Arendall WB, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Murray LW, Richardson JS, Richardson DC (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr Sect D Biol Crystallogr 66:12–21. https://doi.org/10.1107/S0907444909042073
Chyau Liang T (1998) Encyclopedia of Immunology. 825–827 pp. https://doi.org/10.1006/rwei.1999.0219
Climent N, Ferrer S, Rubires X, Merino S, Tomás JM, Regué M (1997) Molecular characterization of a 17-kDa outer-membrane protein from Klebsiella pneumoniae Res Microbiol 148:133–143. https://doi.org/10.1016/S0923-2508(97)87644-9
Dai P, Hu D (2022) The making of hypervirulent Klebsiella pneumoniae J Clin Lab Anal 36:e24743. https://doi.org/10.1002/jcla.24743
De Groot AS, Bosma A, Chinai N, Frost J, Jesdale BM, Gonzalez MA, Martin W, Saint-Aubin C (2001) From genome to vaccine: in silico predictions, ex vivo verification. Vaccine 19:4385–4395. https://doi.org/10.1016/s0264-410x(01)00145-1
de Vries SJ, Bonvin AMJJ (2011) CPORT: a consensus interface predictor and its performance in prediction-driven docking with HADDOCK. PLoS ONE 6:e17695. https://doi.org/10.1371/journal.pone.0017695
Fleri W, Paul S, Dhanda SK, Mahajan S, Xu X, Peters B, Sette A (2017) The immune epitope database and analysis resource in epitope discovery and synthetic vaccine design. Front Immunol 8:278. https://doi.org/10.3389/fimmu.2017.00278
Gupta S, Kapoor P, Chaudhary K, Gautam A, Kumar R, Raghava GPS (2013) In Silico Approach for Predicting toxicity of peptides and proteins. PLoS ONE 8:e73957. https://doi.org/10.1371/journal.pone.0073957
He Y, Rappuoli R, De Groot AS, Chen RT (2010) Emerging vaccine informatics. J Biomed Biotechnol 2010:1–26. https://doi.org/10.1155/2010/218590
Hussein KE, Bahey-El-Din M, Sheweita SA (2018) Immunization with the outer membrane proteins OmpK17 and OmpK36 elicits protection against Klebsiella pneumoniae in the murine infection model. Microb Pathog 119:12–18. https://doi.org/10.1016/j.micpath.2018.04.004
Ivanciuc O (2003) SDAP: database and computational tools for allergenic proteins. Nucleic Acids Res 31:359–362. https://doi.org/10.1093/nar/gkg010
Kringelum JV, Lundegaard C, Lund O, Nielsen M (2012) Reliable B cell epitope predictions: impacts of method development and improved benchmarking. PLoS Comput Biol 8:e1002829. https://doi.org/10.1371/journal.pcbi.1002829
Kurupati P, Teh BK, Kumarasinghe G, Poh CL (2006) Identification of vaccine candidate antigens of an ESBL producing Klebsiella pneumoniae clinical strain by immunoproteome analysis. Proteomics 6:836–844. https://doi.org/10.1002/pmic.200500214
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132. https://doi.org/10.1016/0022-2836(82)90515-0
Lee WH, Choi H, Il, Hong SW, Kim KS, Gho YS, Jeon SG (2015) Vaccination with Klebsiella pneumoniae-derived extracellular vesicles protects against bacteria-induced lethality via both humoral and cellular immunity. Exp Mol Med 47:e183. https://doi.org/10.1038/emm.2015.59
Lei Y, Zhao F, Shao J, Li Y, Li S, Chang H, Zhang Y (2019) Application of built-in adjuvants for epitope-based vaccines. PeerJ 6:e6185. https://doi.org/10.7717/peerj.6185
Malachowa N, Kobayashi SD, Porter AR, Freedman B, Hanley PW, Lovaglio J, Saturday GA, Gardner DJ, Scott DP, Griffin A, Cordova K, Long D, Rosenke R, Sturdevant DE, Bruno D, Martens C, Kreiswirth BN, DeLeo FR (2019) Vaccine protection against multidrug-resistant Klebsiella pneumoniae in a nonhuman primate model of severe lower respiratory tract infection. MBio 10:e02994-19. https://doi.org/10.1128/mBio.02994-19
Maurer-Stroh S, Krutz NL, Kern PS, Gunalan V, Nguyen MN, Limviphuvadh V, Eisenhaber F, Gerberick GF (2019) AllerCatPro—prediction of protein allergenicity potential from the protein sequence. Bioinformatics 35:3020–3027. https://doi.org/10.1093/bioinformatics/btz029
Nemati Zargaran F, Akya A, Ghadiri K, Ranjbarian P, Rostamian M (2021) Detecting the dominant T and B epitopes of Klebsiella pneumoniae Ferric Enterobactin protein (FepA) and introducing a single epitopic peptide as vaccine candidate. Int J Pept Res Ther 27:2209–2221. https://doi.org/10.1007/s10989-021-10247-3
Opoku-Temeng C, Malachowa N, Kobayashi SD, DeLeo FR (2022) Innate host defense against Klebsiella pneumoniae and the outlook for development of immunotherapies. J Innate Immun 14:167–181. https://doi.org/10.1159/000518679
Piperaki ET, Syrogiannopoulos GA, Tzouvelekis LS, Daikos GL (2017) Klebsiella pneumoniae: virulence, biofilm and antimicrobial resistance. Pediatr Infect Dis J 36:1002–1005. https://doi.org/10.1097/INF.0000000000001675
Pletz MW, Uebele J, Gotz K, Hagel S, Bekeredjian-Ding I (2016) Vaccines against major ICU pathogens: where do we stand? Curr Opin Crit Care 22:470–476. https://doi.org/10.1097/MCC.0000000000000338
Ponomarenko J, Bui HH, Li W, Fusseder N, Bourne PE, Sette A, Peters B (2008) ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinf 9:514. https://doi.org/10.1186/1471-2105-9-514
Rahmat Ullah S, Majid M, Rashid MI, Mehmood K, Andleeb S (2021) Immunoinformatics driven prediction of multiepitopic vaccine against Klebsiella pneumoniae and Mycobacterium tuberculosis coinfection and its validation via in silico expression. Int J Pept Res Ther 27:987–999. https://doi.org/10.1007/s10989-020-10144-1
Rostamian M, Farasat A, Chegene Lorestani R, Nemati Zargaran F, Ghadiri K, Akya A (2020) Immunoinformatics and molecular dynamics studies to predict T-cell-specific epitopes of four Klebsiella pneumoniae fimbriae antigens. J Biomol Struct Dyn 40:166–176. https://doi.org/10.1080/07391102.2020.1810126
Saha S, Raghava GPS (2006) Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins Struct Funct Genet 65:40–48. https://doi.org/10.1002/prot.21078
Schuler MM, Nastke MD, Stevanovikc S (2007) SYFPEITHI: database for searching and T-cell epitope prediction. Methods Mol Biol 409:75–93. https://doi.org/10.1007/978-1-60327-118-9_5
Shen Y, Maupetit J, Derreumaux P, Tuffery P (2014) Improved PEP-FOLD approach for peptide and miniprotein structure prediction. J Chem Theory Comput 10:4745–4758. https://doi.org/10.1021/ct500592m
Singh H, Raghava GPS (2003) ProPred1: prediction of promiscuous MHC Class-I binding sites. Bioinformatics 19:1009–1014. https://doi.org/10.1093/bioinformatics/btg108
Ten Hagen TLM, Van Vianen W, Savelkoul HFJ, Heremans H, Buurman WA, Bakker-Woudenberg IAJM (1998) Involvement of T cells in enhanced resistance to Klebsiella pneumoniae septicemia in mice treated with liposome-encapsulated muramyl tripeptide phosphatidylethanolamine or gamma interferon. Infect Immun 66:1962–1967. https://doi.org/10.1128/IAI.66.5.1962-1967.1998
Trost B, Bickis M, Kusalik A (2007) Strength in numbers: achieving greater accuracy in MHC-I binding prediction by combining the results from multiple prediction tools. Immunome Res 3:5. https://doi.org/10.1186/1745-7580-3-5
Uzairue LI, Rabaan AA, Adewumi FA, Okolie OJ, Folorunso JB, Bakhrebah MA, Garout M, Alfouzan WA, Halwani MA, Alamri AA, Halawani SA, Alshahrani FS, Hasan A, Mutair A, Al, Alhumaid S, Etafo J, Utip I, Odoh IM, Uwaezuoke NS (2022) Global prevalence of colistin resistance in Klebsiella pneumoniae from bloodstream infection: a systematic review and meta-analysis. Pathogens 11:1092. https://doi.org/10.3390/pathogens11101092
van Zundert GCP, Rodrigues JPGLM, Trellet M, Schmitz C, Kastritis PL, Karaca E, Melquiond ASJ, van Dijk M, de Vries SJ, Bonvin AMJJ (2016) The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol 428:720–725. https://doi.org/10.1016/j.jmb.2015.09.014
Wang M, Zhu D, Zhu J, Nussinov R, Ma B (2018) Local and global anatomy of antibody-protein antigen recognition. J Mol Recognit 31:e2693. https://doi.org/10.1002/jmr.2693
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296–W303. https://doi.org/10.1093/nar/gky427
Weinberger B, Herndler-Brandstetter D, Schwanninger A, Weiskopf D, Grubeck‐Loebenstein B (2008) Biology of immune responses to vaccines in elderly persons. Clin Infect Dis 46:1078–1084. https://doi.org/10.1086/529197
Zargaran FN, Akya A, Rezaeian S, Ghadiri K, Lorestani RC, Madanchi H, Safaei S, Rostamian M (2021) B cell epitopes of four fimbriae antigens of Klebsiella pneumoniae: a comprehensive in silico study for vaccine development. Int J Pept Res Ther 27:875–886. https://doi.org/10.1007/s10989-020-10134-3
Zhu J, Wang T, Chen L, Du H (2021) Virulence factors in hypervirulent Klebsiella pneumoniae Front Microbiol 12:642484. https://doi.org/10.3389/fmicb.2021.642484
Acknowledgements
The support of this work by the Infectious Diseases Research Center of Kermanshah University of Medical Sciences is highly appreciated.
Funding
This work was supported by the Kermanshah University of Medical Sciences as a student thesis.
Author information
Authors and Affiliations
Contributions
Parivash Ranjbarian and Mosayeb Rostamian contributed to the study’s conception and design. Data collection was done by Parivash Ranjbarian, Farjam Goudarzi, Alisha Akya, and Hana Heidarinia. Data analysis was performed by Alireza Farasat and Mosayeb Rostamian. Funding acquisition and resource preparation were done by Alisha Akya and Mosayeb Rostamian. The first draft of the manuscript was written by Mosayeb Rostamian. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Fig. S1
Charts of linear B cell epitopes prediction for OMPK17. For each prediction tool, a chart was supplied in which residues with higher scores (compared to the threshold) are more possible to be epitopes (shown in yellow). (JPG 1.60 MB)
Fig. S2
The conservancy of the 25-mer epitopic peptide of OMPK17. NCBI conserved domain search indicated that the 25-mer peptide is a member of the highly conserved OM-channels (porin) superfamily in Gram-negative bacteria. (JPG 210 KB)
Table S1
(XLS 19.0 KB)
Table S2
(XLS 21.0 KB)
Table S3
(XLS 402 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ranjbarian, P., Goudarzi, F., Akya, A. et al. Finding epitopes of Klebsiella pneumoniae outer membrane protein-K17 (OMPK17) and introducing a 25-mer peptide of it as a vaccine candidate. Biologia 78, 2261–2271 (2023). https://doi.org/10.1007/s11756-023-01371-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01371-0