Abstract
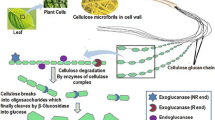
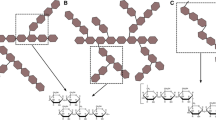
Sugar chains are biopolymers that play various important roles for organisms, and their structures are complex and diverse. Enzymes acting on sugar chains have undergone a wide variety of molecular evolutions. Nevertheless, the limited availability of sugar chains often prevents the investigation of related enzymes, even among enzymes that act on sugar chains comprising only glucose, an essential monosaccharide in nature. Glucans such as starch and cellulose represent a large amount of biomass, and enzymes acting on these glucans have been investigated extensively. In contrast, there is much less known about enzymes acting on glucans with β-1,2-glucosidic linkages (β-1,2-glucans), especially those enzymes involved in degrading β-1,2-glucans. Research on such degrading enzymes has only been undertaken for approximately a decade. In this review, β-1,2-glucan-associated enzymes will be introduced, especially β-1,2-glucan-degrading enzymes such as a glycoside phosphorylase acting on β-1,2-glucooligosaccharides and the identification of two novel β-1,2-glucanases and related enzymes, along with the distribution and physiological role of β-1,2-glucans in nature.








Similar content being viewed by others
Abbreviations
- ACP:
-
Acyl carrier protein
- BGL:
-
β-Glucosidase
- CBP:
-
Cellobiose phosphorylase
- CβG:
-
Cyclic β-1,2-glucan
- CGS:
-
Cyclic β-1,2-glucan synthase
- CpSGL:
-
SGL from Chitinophaga pinensis
- DP:
-
Degree of polymerization
- G1P:
-
α-Glucose 1-phosphate
- GH:
-
Glycoside hydrolase family
- IaSGT:
-
IALB_1185 from Ignavibacterium album
- Lam2 :
-
Laminaribiose
- LiBGL:
-
BGL from Listeria innocua
- LiSO-BP:
-
SO-BP from L. innocua
- LiSOGP:
-
SOGP from L. innocua
- LpSOGP:
-
SOGP from Lachnochlostridium phytofermentans
- OPG:
-
Osmoregulated periplasmic glucans
- PdSGL:
-
SGL from Parabacteroides distasonis
- SGL:
-
β-1,2-Glucanase
- SO-BP:
-
Sopn-binding protein
- SOGP:
-
1,2-β-Oligoglucan phosphorylase
- Sopn :
-
β-1,2-Glucooligosaccharide with DP of n
- TfSGL:
-
SGL from Talaromyces funiculosus
References
Abe K, Nakajima M, Kitaoka M, Toyoizumi H, Takahashi Y, Sugimoto N, Nakai H, Taguchi H (2015) Large-scale preparation of 1,2-β-glucan using 1,2-β-oligoglucan phosphorylase. J Appl Glycosci 62:47–52. https://doi.org/10.5458/jag.jag.JAG-2014_011
Abe K, Nakajima M, Yamashita T, Matsunaga H, Kamisuki S, Nihira T, Takahashi Y, Sugimoto N, Miyanaga A, Nakai H, Arakawa T, Fushinobu S, Taguchi H (2017) Biochemical and structural analyses of a bacterial endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem 292:7487–7506. https://doi.org/10.1074/jbc.M116.762724
Abe K, Sunagawa N, Terada T, Takahashi Y, Arakawa T, Igarashi K, Samejima M, Nakai H, Taguchi H, Nakajima M, Fushinobu S (2018) Structural and thermodynamic insights into β-1,2-glucooligosaccharide capture by a solute-binding protein in Listeria innocua. J Biol Chem 293:8812–8828. https://doi.org/10.1074/jbc.RA117.001536
Alzari PM, Souchen H, Dominguez R (1996) The crystal structure of endoglucanase CelA, a family 8 glycosyl hydrolase from Clostridium thermocellum. Structure 4:265–275. https://doi.org/10.1107/S2053230X16017891
Amemura A, Hisamatsu M, Mitani H, Harada T (1983) Cyclic (1→2)-β-d-glucan and the octasaccharide repeating-units of extracellular acidic polysaccharides produced by Rhizobium. Carbohydr Res 114:277–285. https://doi.org/10.1016/0008-6215(83)88194-4
Amemura A, Cabrera-Crespo J (1986) Extracellular oligosaccharides and Low-M, polysaccharides containing (1→2)-β-d-glucosidic linkages from strains of Xanthornonas, Escherichia coli and Klebsiella pneumoniae. J Gen Microbiol 132:2443–2452. https://doi.org/10.1099/00221287-132-9-2443
Arellano-Reynoso B, Lapaque N, Salcedo S, Briones G, Ciocchini AE, Ugalde R, Moreno E, Moriyón I, Gorvel JP (2005) Cyclic β-1,2-glucan is a brucella virulence factor required for intracellular survival. Nat Immunol 6:618–625. https://doi.org/10.1038/ni1202
Astua-monge G, Freitas-astua J, Bacocina G, Roncoletta J, Machado MA (2005) Expression profiling of virulence and pathogenicity genes of Xanthomonas axonopodis pv. citri. J Bacteriol 187:1201–1205. https://doi.org/10.1128/JB.187.3.1201
Barsanti L, Vismara R, Passarelli V, Gualtieri P (2001) Paramylon (β-1,3-glucan) content in wild type and WZSL mutant of Euglena gracilis. Effects of growth conditions. J Appl Phycol 13:59–65. https://doi.org/10.1023/A:1008105416065
Barsanti L, Gualtieri P (2019) Paramylon, a potent immunomodulator from WZSL mutant of Euglena gracilis. Molecules 24:3114. https://doi.org/10.3390/molecules24173114
Bauer S, Vasu P, Persson S, Mort AJ, Somerville CR (2006) Development and application of a suite of polysaccharide-degrading enzymes for analyzing plant cell walls. Proc Natl Acad Sci U S A 103:11417–11422. https://doi.org/10.1073/pnas.0604632103
Becker S, Scheffel A, Polz MF, Hehemann JH (2017) Accurate quantification of laminarin in marine organic matter with enzymes from marine microbes. Appl Environ Microbiol 83:1–14. https://doi.org/10.1128/AEM.03389-16
Bohin JP (2000) Osmoregulated periplasmic glucans in Proteobacteria. FEMS Microbiol Lett 186:11–19. https://doi.org/10.1111/j.1574-6968.2000.tb09075.x
Bontemps-Gallo S, Cogez V, Robbe-Masselot C, Quintard K, Dondeyne J, Madec E, Lacroix JM (2013) Biosynthesis of osmoregulated periplasmic glucans in Escherichia coli: The phosphoethanolamine transferase is encoded by opgE. Biomed Res Int 2013:371429. https://doi.org/10.1155/2013/371429
Bontemps-Gallo S, Madec E, Robbe-Masselot C, Souche E, Dondeyne J, Lacroix JM (2016) The opgC gene is required for OPGs succinylation and is osmoregulated through RcsCDB and EnvZ/OmpR in the phytopathogen Dickeya dadantii. Sci Rep 6:srep19619. https://doi.org/10.1038/srep19619
Bontemps-Gallo S, Bohin JP, Lacroix JM (2017) Osmoregulated periplasmic glucans. EcoSal Plus 7:1–17. https://doi.org/10.1128/ecosalplus.ESP-0001-2017
Brahmachari G, Mandal LC, Roy R, Sadhan M, Brahmachari AK (2011) Stevioside and related compounds – molecules of pharmaceutical promise: A critical overview. Arch Pharm Chem Life Sci 344:5–19. https://doi.org/10.1002/ardp.201000181
Breedveld MW, Zevenhuizen LP, Zehnder AJ (1990) Excessive excretion of cyclic β-(1,2)-glucan by Rhizobium trifolii TA-1. Appl Env Microbiol 56:2080–2086. https://doi.org/10.1128/aem.56.7.2080-2086.1990
Brown DG, Allen C (2004) Ralstonia solanacearum genes induced during growth in tomato: An inside view of bacterial wilt. Mol Microbiol 53:1641–1660. https://doi.org/10.1111/j.1365-2958.2004.04237.x
Bundle DR, Cherwonogrodzky JW, Perry MB (1988) Characterization of Brucella polysaccharide B. Infect Immun 56:1101–1106. https://doi.org/10.1128/iai.56.5.1101-1106.1988
Cangelosi GA, Martinetti G, Leigh JA, Lee CC, Theines C, Nester EW (1989) Role for Agrobacterium tumefaciens ChvA protein in export of β-1,2-glucan. J Bacteriol 171:1609–1615. https://doi.org/10.1128/jb.171.3.1609-1615.1989
Castro OA, Zorreguieta A, Ielmini V, Vega G, Ielpi L (1996) Cyclic β-(1,2)-glucan synthesis in Rhizobiaceae: roles of the 319-kilodalton protein intermediate. J Bacteriol 178:6043–6048. https://doi.org/10.1128/jb.178.20.6043-6048.1996
CAZypedia Consortium (2018) Ten years of CAZypedia: a living encyclopedia of carbohydrate-active enzymes. Glycobiology 28(1):3–8. https://doi.org/10.1093/glycob/cwx089
Celli J, Gorvel J-P (2004) Organelle robbery: Brucella interactions with the endoplasmic reticulum. Curr Opin Microbiol 7:93–97. https://doi.org/10.1016/j.mib.2003.11.001
Cheng W, Wang L, Jiang YL, Bai XH, Chu J, Li Q, Yu G, Liang QL, Zhou CZ, Chen Y (2012) Structural insights into the substrate specificity of Streptococcus pneumoniae β(1,3)-galactosidase BgaC. J Biol Chem 287:22910–22918. https://doi.org/10.1074/jbc.M112.367128
Ciocchini AE, Roset MS, De Iñó N, Ugalde RA (2004) Membrane topology analysis of cyclic glucan synthase, a virulence determinant of Brucella abortus. J Bacteriol 186:7205–7213. https://doi.org/10.1128/JB.186.21.7205-7213.2004
Ciocchini AE, Roset MS, Briones G, Iñón de Iannino N, Ugalde RA (2006) Identification of active site residues of the inverting glycosyltransferase Cgs required for the synthesis of cyclic β-1,2-glucan, a Brucella abortus virulence factor. Glycobiology 16:679–691. https://doi.org/10.1093/glycob/cwj113
Ciocchini AE, Guidolin LS, Casabuono AC, Couto AS, de Iannino NI, Ugalde RA (2007) A glycosyltransferase with a length-controlling activity as a mechanism to regulate the size of polysaccharides. Proc Natl Acad Sci U S A 104:16492–16497. https://doi.org/10.1073/pnas.0708025104
Claus S, Van Bogaert INA (2017) Sophorolipid production by yeasts: a critical review of the literature and suggestions for future research. Appl Microbiol Biotechnol 101:7811–7821. https://doi.org/10.1007/s00253-017-8519-7
Dartigalongue C, Missiakas D, Raina S (2001) Characterization of the Escherichia coli σE regulon. J Biol Chem 276:20866–20875. https://doi.org/10.1074/jbc.M100464200
de Iannino NI, Ugalde RA (1989) Biochemical characterization of avirulent Agrobacterium tumefaciens chvA mutants: synthesis and excretion of β-(1–2)glucan. J Bacteriol 171:2842–2849. https://doi.org/10.1128/jb.171.5.2842-2849.1989
De Souza AA, Takita MA, Coletta-Filho HD, Caldana C, Yanai GM, Muto NH, De Oliveira RC, Nunes LR, Machado MA (2004) Gene expression profile of the plant pathogen Xylella fastidiosa during biofilm formation in vitro. FEMS Microbiol Lett 237:341–353. https://doi.org/10.1016/j.femsle.2004.06.055
Debarbieux L, Bohin A, Bohin JP (1997) Topological analysis of the membrane-bound glucosyltransferase, MdoH, required for osmoregulated periplasmic glucan synthesis in Escherichia coli. J Bacteriol 179:6692–6698. https://doi.org/10.1128/jb.179.21.6692-6698.1997
Dell A, York WS, McNeil M, Darvill AG, Albersheim P (1983) The cyclic structure of β-d-(1→2)-linked d-glucans secreted by Rhizobia and Agrobacteria. Carbohydr Res 117:185–200. https://doi.org/10.1016/0008-6215(83)88086-0
Drula E, Garron ML, Dogan S, Lombard V, Henrissat B, Terrapon N (2022) The carbohydrate-active enzyme database: Functions and literature. Nucleic Acids Res 50:D571–D577. https://doi.org/10.1093/nar/gkab1045
Dylan T, Ielpi L, Stanfield S (1986) Rhizobium meliloti genes required for nodule development are related to chromosomal virulence genes in Agrobacterium tumefaciens. Proc Natl Acad Sci U S A 83:4403–4407. https://doi.org/10.1073/pnas.83.12.4403
Dylan T, Helinski DR, Ditta GS (1990) Hypoosmotic adaptation in Rhizobium meliloti requires β-(1→2)-glucan. J Bacteriol 172:1400–1408. https://doi.org/10.1128/jb.172.3.1400-1408.1990
Felse PA, Shah V, Chan J, Rao KJ, Gross RA (2007) Sophorolipid biosynthesis by Candida bombicola from industrial fatty acid residues. Enzyme Microb Technol 40:316–323. https://doi.org/10.1016/j.enzmictec.2006.04.013
Ferreres F, Valentão P, Llorach R, Pinheiro C, Cardoso L, Pereira JA, Sousa C, Seabra RM, Andrade PB (2005) Phenolic compounds in external leaves of tronchuda cabbage (Brassica oleracea L. var. costata DC). J Agric Food Chem 53:2901–2907. https://doi.org/10.1021/jf040441s
Fiedler W, Rotering H (1985) Characterization of an Escherichia coli mdoB mutant strain unable to transfer sn-1-phosphoglycerol to membrane-derived oligosaccharides. J Biol Chem 260:4799–4806. https://doi.org/10.1016/s0021-9258(18)89142-6
Gay-Fraret J, Ardissone S, Kambara K, Broughton WJ, Deakin WJ, Le Quéré A (2012) Cyclic-β-glucans of Rhizobium (Sinorhizobium) sp. strain NGR234 are required for hypo-osmotic adaptation, motility, and efficient symbiosis with host plants. FEMS Microbiol Lett 333:28–36. https://doi.org/10.1111/j.1574-6968.2012.02595.x
Golisch B, Lei Z, Tamura K, Brumer H (2021) Configured for the human gut microbiota: molecular mechanisms of dietary β-glucan utilization. ACS Chem Biol 16:2087–2102. https://doi.org/10.1021/acschembio.1c00563
Goulas T, Goulas A, Tzortzis G, Gibson GR (2009) Comparative analysis of four β-galactosidases from Bifidobacterium bifidum NCIMB41171: Purification and biochemical characterisation. Appl Microbiol Biotechnol 82:1079–1088. https://doi.org/10.1007/s00253-008-1795-5
Gregg KJ, Suits MDL, Deng L, Vocadlo DJ, Boraston AB (2015) Structural analysis of a family 101 glycoside hydrolase in complex with carbohydrates reveals insights into its mechanism. J Biol Chem 290:25657–25669. https://doi.org/10.1074/jbc.M115.680470
Guidolin LS, Ciocchini AE, De Iannino NI, Ugalde RA (2009) Functional mapping of Brucella abortus cyclic β-1,2-glucan synthase: Identification of the protein domain required for cyclization. J Bacteriol 191:1230–1238. https://doi.org/10.1128/JB.01108-08
Guidolin LS, Morrone Seijo SM, Guaimas FF, Comerci DJ, Ciocchinia AE (2015) Interaction network and localization of Brucella abortus membrane proteins involved in the synthesis, transport, and succinylation of cyclic β-1,2-glucans. J Bacteriol 197:1640–1648. https://doi.org/10.1128/JB.00068-15
Hanoulle X, Rollet E, Clantin B, Landrieu I, Ödberg-Ferragut C, Lippens G, Bohin JP, Villeret V (2004) Structural analysis of Escherichia coli OpgG, a protein required for the biosynthesis of osmoregulated periplasmic glucans. J Mol Biol 342:195–205. https://doi.org/10.1016/j.jmb.2004.07.004
Helbert W, Poulet L, Drouillard S, Mathieu S, Loiodice M, Couturier M, Lombard V, Terrapon N, Turchetto J, Vincentelli R, Henrissat B (2019) Discovery of novel carbohydrate-active enzymes through the rational exploration of the protein sequences space. Proc Natl Acad Sci U S A 116:6063–6068. https://doi.org/10.1073/pnas.1815791116
Henrissat B, Vegetales M, Grenoble F (1991) A classification of glycosyl hydrolases based sequence similarities amino acid. Biochem J 280:309–316. https://doi.org/10.1007/s007920050009
Henrissat B, Sulzenbacher G, Bourne Y (2008) Glycosyltransferases, glycoside hydrolases: surprise, surprise! Curr Opin Struct Biol 18:527–533. https://doi.org/10.1016/j.sbi.2008.09.003
Hidaka M, Honda Y, Kitaoka M, Nirasawa S, Hayashi K, Wakagi T, Shoun H, Fushinobu S (2004) Chitobiose phosphorylase from Vibrio proteolyticus, a member of glycosyl transferase family 36, has a clan GH-L-like (α/α)6 barrel fold. Structure 12:937–947. https://doi.org/10.1016/j.str.2004.03.027
Hidaka M, Kitaoka M, Hayashi K, Wakagi T, Shoun H, Fushinobu S (2006) Structural dissection of the reaction mechanism of cellobiose phosphorylase. Biochem J 398:37–43. https://doi.org/10.1042/BJ20060274
Higashiura T, Ikeda M, Okubo M, Hisamatsu M, Amemura A, Harada T (1985) An improved method for preparation of cyclic (1→2)-β-d-glucan using an acidic polysaccharide-negative mutant of Rhizobium phaseoli AHU1133. Agric Biol Chem 49:1865–1866. https://doi.org/10.1271/bbb1961.49.1865
Hill NS, Buske PJ, Shi Y, Levin PA (2013) A moonlighting enzyme links Escherichia coli cell size with central metabolism. PLOS Genet 9:e1003663. https://doi.org/10.1371/journal.pgen.1003663
Hiraishi M, Igarashi K, Kimura S, Wada M, Kitaoka M, Samejima M (2009) Synthesis of highly ordered cellulose II in vitro using cellodextrin phosphorylase. Carbohydr Res 344:2468–2473. https://doi.org/10.1016/j.carres.2009.10.002
Hisamatsu M, Amemura A, Matsuo T, Matsuda H, Harada T (1982) Cyclic (1→2)-β-d-glucan and the octasaccharide repeating-unit of succinoglycan produced by Agrobactenium. J Gen Microbiol 128:1873–1879
Hisamatsu M (1992) Cyclic (1→2)-β-d-glucans (cyclosophorans) produced by Agrobacterium and Rhizobium species. Carbohydr Res 231:137–146. https://doi.org/10.1016/0008-6215(92)84014-J
Holm L (2020) DALI and the persistence of protein shape. Protein Sci 29:128–140. https://doi.org/10.1002/pro.3749
Honda Y, Kitaoka M, Hayashi K (2004) Reaction mechanism of chitobiose phosphorylase from Vibrio proteolyticus: identification of family 36 glycosyltransferase in Vibrio. Biochem J 377:225–232. https://doi.org/10.1042/BJ20031171
Hori C, Ishida T et al (2014) Analysis of the Phlebiopsis gigantea genome, transcriptome and secretome provides insight into its pioneer colonization strategies of wood. PLOS Genet 10:e1004759. https://doi.org/10.1371/journal.pgen.1004759
Hrmova M, De Gori R, Smith BJ, Fairweather JK, Driguez H, Varghese JN, Fincher GB (2002) Structural basis for broad substrate specificity in higher plant β-d-glucan glucohydrolases. Plant Cell 14:1033–1052. https://doi.org/10.1105/tpc.010442
Iñón de Iannino N, Ugalde RA (1989) Biochemical characterization of avirulent Agrobacterium tumefaciens chvA mutants: synthesis and excretion of β-(1–2)glucan. J Bacteriol 171:2842–2849. https://doi.org/10.1128/jb.171.5.2842-2849.1989
Iñón de Iannino N, Briones G, Tolmasky M, Ugalde RA (1998) Molecular cloning and characterization of cgs, the Brucella abortus cyclic β(1–2) glucan synthetase gene: genetic complementation of Rhizobium meliloti ndvB and Agrobacterium tumefaciens chvB mutants. J Bacteriol 180:4392–4400. https://doi.org/10.1128/JB.180.17.4392-4400.1998
Ishida T, Fushinobu S, Kawai R, Kitaoka M, Igarashi K, Samejima M (2009) Crystal structure of glycoside hydrolase family 55 β-1,3-glucanase from the basidiomycete Phanerochaete chrysosporium. J Biol Chem 284:10100–10109. https://doi.org/10.1074/jbc.M808122200
Ishiguro R, Tanaka N, Abe K, Nakajima M, Maeda T, Miyanaga A, Takahashi Y, Sugimoto N, Nakai H, Taguchi H (2017) Function and structure relationships of a β-1,2-glucooligosaccharide-degrading β-glucosidase. FEBS Lett 591:3926–3936. https://doi.org/10.1002/1873-3468.12911
Javvadi S, Pandey SS, Mishra A, Pradhan BB, Chatterjee S (2018) Bacterial cyclic β-(1,2)-glucans sequester iron to protect against iron-induced toxicity. EMBO Rep 19:172–186. https://doi.org/10.15252/embr.201744650
Kanaoka M, Akao T, Kobashi K (1994) Metabolism of ginseng saponins, ginsenosides, by human intestinal flora. J Tradit Med 11:241–245
Kennedy EP (1982) Osmotic regulation and the biosynthesis of membrane derived oligosaccharides in Escherichia coli. Proc Natl Acad Sci U S A 79:1092–1095. https://doi.org/10.1073/pnas.79.4.1092
Kim TH, Ku SK, Lee IC, Bae JS (2012) Anti-inflammatory effects of kaempferol-3-O-sophoroside in human endothelial cells. Inflamm Res 61:217–224. https://doi.org/10.1007/s00011-011-0403-9
Kim YK, Kitaoka M, Krishnareddy M, Mori Y, Hayashi K (2002) Kinetic studies of a recombinant cellobiose phosphorylase (CBP) of the Clostridium thermocellum YM4 strain expressed in Escherichia coli. J Biochem 132:197–203. https://doi.org/10.1093/oxfordjournals.jbchem.a003210
Kitahata S, Edagawa S (1987) Cyclic (1→2)-β-d-glucan-hydrolyzing enzymes from Acremonium sp. 15: Purification and some properties of endo-(1→2)-β-d-glucanase and β-d-glucosidase. Agric Biol Chem 51:2701–2708. https://doi.org/10.1271/bbb1961.51.2701
Kitaoka M, Matsuoka Y, Mori K, Nishimoto M, Hayashi K (2012) Characterization of a bacterial laminaribiose phosphorylase. Biosci Biotechnol Biochem 76:343–348. https://doi.org/10.1271/bbb.110772
Kobayashi K, Aramasa H, Nakai H, Nakajima M, Taguchi H (2018) Colorimetric determination of β-1,2-glucooligosaccharides for an enzymatic assay using 3-methyl-2-benzothiazolinonehydrazone. Anal Biochem 560:1–6. https://doi.org/10.1016/j.ab.2018.08.021
Kobayashi K, Nakajima M, Aramasa H, Kimura S, Iwata T, Nakai H, Taguchi H (2019) Large-scale preparation of β-1,2-glucan using quite a small amount of sophorose. Biosci Biotechnol Biochem 83:1867–1874. https://doi.org/10.1080/09168451.2019.1630257
Kobayashi K, Shimizu H, Tanaka N, Kuramochi K, Nakai H, Nakajima M, Taguchi H (2022) Characterization and structural analyses of a novel glycosyltransferase acting on the β-1,2-glucosidic linkages. J Biol Chem 298:101606. https://doi.org/10.1016/j.jbc.2022.101606
Koivula A, Ruohonen L, Wohlfahrt G, Reinikainen T, Teeri TT, Piens K, Claeyssens M, Weber M, Vasella A, Becker D, Sinnott ML, Zou J, Kleywegt GJ, Szardenings M, Ståhlberg J, Jones TA (2002) The active site of cellobiohydrolase Cel6A from Trichoderma reesei: The roles of aspartic acids D221 and D175. J Am Chem Soc 124:10015–10024. https://doi.org/10.1021/ja012659q
Koizumi K, Okada Y, Utamura T (1984) Further studies on the separation of cyclic (1→2)-β-d-glucans (cyclosophoraoses) produced by Rhizobium meliloti IFO 13336, and determination of their degrees of polymerization by high-performance liquid chromatography. J Chromatogr 299:215–224. https://doi.org/10.1016/S0021-9673(01)97833-1
Kondo T, Nishimura Y, Matsuyama K, Ishimaru M, Nakazawa M, Ueda M, Sakamoto T (2020) Characterization of three GH35 β-galactosidases, enzymes able to shave galactosyl residues linked to rhamnogalacturonan in pectin, from Penicillium chrysogenum 31B. Appl Microbiol Biotechnol 104:1135–1148. https://doi.org/10.1007/s00253-019-10299-y
Kondo Y, Kato A, Hojo H, Nozoe S, Takeuchi M, Ochi K (1992) Cytokine-related immunopotentiation activities of paramylon, a β-(1→3)-d-glucan from Euglena gracilis. J Pharmacobio-Dyn 15:617–621. https://doi.org/10.1248/bpb1978.15.617
Kony DB, Damm W, Stoll S, Van Gunsteren WF, Hünenberger PH (2007) Explicit-solvent molecular dynamics simulations of the polysaccharide schizophyllan in water. Biophys J 93:442–455. https://doi.org/10.1529/biophysj.106.086116
Kremer C, Pettolino F, Bacic A, Drinnan A (2004) Distribution of cell wall components in Sphagnum hyaline cells and in liverwort and hornwort elaters. Planta 219:1023–1035. https://doi.org/10.1007/s00425-004-1308-4
Kuhaudomlarp S, Pergolizzi G, Patron NJ, Henrissat B, Field RA (2019) Unraveling the subtleties of β-(1→3)-glucan phosphorylase specificity in the GH94, GH149, and GH161 glycoside hydrolase families. J Biol Chem 294:6483–6493. https://doi.org/10.1074/jbc.RA119.007712
Lacroix JM, Tempête M, Menichi B, Bohin JP (1989) Molecular cloning and expression of a locus (mdoA) implicated in the biosynthesis of membrane-derived oligosaccharides in Escherichia coli. Mol Microbiol 3:1173–1182. https://doi.org/10.1111/j.1365-2958.1989.tb00267.x
Lacroix JM, Loubens I, Tempête M, Menichi B, Bohin JP (1991) The mdoA locus of Escherichia coli consists of an operon under osmotic control. Mol Microbiol 5:1745–1753. https://doi.org/10.1111/j.1365-2958.1991.tb01924.x
Lacroix JM, Lanfroy E, Cogez V, Lequette Y, Bohin A, Bohin JP (1999) The mdoC gene of Escherichia coli encodes a membrane protein that is required for succinylation of osmoregulated periplasmic glucans. J Bacteriol 181:3626–3631. https://doi.org/10.1128/jb.181.12.3626-3631.1999
Laine RA (1994) A calculation of all possible oligosaccharide isomers both branched and linear yield 1.05 × 1012 structures for a reducing hexasaccharides: the Isomer Barrier to development of single-method saccharide sequencing or synthesis systems. Glycobiology 4:759–767. https://doi.org/10.1093/glycob/4.6.759
Lee S, Seo DH, Kim HW, Jung S (2001) Investigation of inclusion complexation of paclitaxel by cyclohenicosakis-(1→2)-(β-d-glucopyranosyl), by cyclic-(1→2)-β-d-glucans (cyclosophoraoses), and by cyclomaltoheptaoses (β-cyclodextrins). Carbohydr Res 334:119–126. https://doi.org/10.1016/S0008-6215(01)00178-1
Lee S, Jung S (2003a) Enantioseparation using cyclosophoraoses as a novel chiral additive in capillary electrophoresis. Carbohydr Res 338:1143–1146. https://doi.org/10.1016/S0008-6215(03)00083-1
Lee S, Seo DH, Park HL, Choi Y, Jung S (2003b) Solubility enhancement of a hydrophobic flavonoid, luteolin by the complexation with cyclosophoraoses isolated from Rhizobium meliloti. Antonie Van Leeuwenhoek 84:201–207. https://doi.org/10.1023/A:1026075215921
Lee S, Jung S (2004) Cyclosophoraose as a catalytic carbohydrate for methanolysis. Carbohydr Res 339:461–468. https://doi.org/10.1016/j.carres.2003.11.004
Lequette Y, Rollet E, Delangle A, Greenberg EP, Bohin JP (2007) Linear osmoregulated periplasmic glucans are encoded by the opgGH locus of Pseudomonas aeruginosa. Microbiology 153:3255–3263. https://doi.org/10.1099/mic.0.2007/008953-0
Lesage G, Bussey H (2006) Cell wall assembly in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 70:317–343. https://doi.org/10.1128/mmbr.00038-05
Levasseur A, Drula E, Lombard V, Coutinho PM, Henrissat B (2013) Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes. Biotechnol Biofuels 6:1. https://doi.org/10.1186/1754-6834-6-41
Li YK, Chir J, Tanaka S, Chen FY (2002) Identification of the general acid/base catalyst of a family 3 β-glucosidase from Flavobacterium meningosepticum. Biochemistry 41:2751–2759. https://doi.org/10.1021/bi016049e
Liberato MV, Prates ET, Gonçalves TA, Bernardes A, Vilela N, Fattori J, Ematsu GC, Chinaglia M, Gomes ERM, Figueira ACM, Damasio A, Polikarpov I, Skaf MS, Squina FM (2021) Insights into the dual cleavage activity of the GH16 laminarinase enzyme class on β-1,3 and β-1,4 glycosidic bonds. J Biol Chem 296:100385. https://doi.org/10.1016/j.jbc.2021.100385
LoScalzo R, Genna A, Branca F, Chedin M, Chassaignec H (2008) Anthocyanin composition of cauliflower (Brassica oleracea L. var. botrytis) and cabbage (B. oleracea L. var. capitata) and its stability in relation to thermal treatments. Food Chem 107:136–144. https://doi.org/10.1016/j.foodchem.2007.07.072
Loubens I, Debarbieux L, Bohin A, Lacroix JM, Bohin JP (1993) Homology between a genetic locus (mdoA) involved in the osmoregulated biosynthesis of periplasmic glucans in Escherichia coli and a genetic locus (hrpM) controlling pathogenicity of Pseudomonas syringae. Mol Microbiol 10:329–340. https://doi.org/10.1111/j.1365-2958.1993.tb01959.x
Macdonald SS, Pereira JH, Liu F, Tegl G, DeGiovanni A, Wardman JF, Deutsch S, Yoshikuni Y, Adams PD, Withers SG (2022) A synthetic gene library yields a previously unknown glycoside phosphorylase that degrades and assembles poly-β-1,3-GlcNAc, completing the suite of β-linked GlcNAc polysaccharides. ACS Cent Sci 8:430–440. https://doi.org/10.1021/acscentsci.1c01570
Mao B, Pear MR, Mccammonj JA, Quiocho FA (1982) Hinge-bending in l-arabinose-binding protein. the “Venus’s-flytrap” model. J Biol Chem 257:1131–1134. https://doi.org/10.1016/S0021-9258(19)68161-5
McAndrew RP, Park JI, Heins RA, Reindl W, Friedland GD, D’haeseleer P, Northen T, Sale KL, Simmons BA, Adams PD (2013) From soil to structure, a novel dimeric β-glucosidase belonging to glycoside hydrolase family 3 isolated from compost using metagenomic analysis. J Biol Chem 288:14985–14992. https://doi.org/10.1074/jbc.M113.458356
McCready RM, Hassid WZ (1944) The Preparation and purification of glucose 1-phosphate by the aid of ion exchange adsorbents. J Am Chem Soc 66:560–563. https://doi.org/10.1021/ja01232a019
McIntire FC, Peterson WH, Riker AJ (1940) Factors influencing the carbon metabolism of the crown gall organism. J Agric Res 61:313–320
McIntire FC, Peterson WH, Riker AJ (1942) A polysaccharide produced by the crown-gall organism. J Biol Chem 143:491–496. https://doi.org/10.1016/s0021-9258(18)72637-9
Medrano FJ, De Souza CS, Romero A, Balan A (2014) Structure determination of a sugar-binding protein from the phytopathogenic bacterium Xanthomonas citri. Acta Crystallogr Sect F 70:564–571. https://doi.org/10.1107/S2053230X14006578
Mendoza NS, Amemura A (1983) (1,2)-β-d-Glucan-hydrolyzing enzymes in Cytophaga arvensicola: partial purification and some properties of endo-(1,2)-β-glucanase and β-d-glucosidase specific for (1,2)-and (1,3)-linkages. J Ferment Technol 61:473–481
Meng J, Huang C, Huang X, Liu D, Han B, Chen J (2020) Osmoregulated periplasmic glucans transmit external signals through Rcs phosphorelay pathway in Yersinia enterocolitica. Front Microbiol 11:122. https://doi.org/10.3389/fmicb.2020.00122
Meng J, Young G, Chen J (2021) The Rcs system in Enterobacteriaceae: envelope stress responses and virulence regulation. Front Microbiol 12:627104. https://doi.org/10.3389/fmicb.2021.627104
Miller KJ, Kennedy EP, Reinhold VN (1986) Osmotic adaptation by gram-negative bacteria: possible role for periplasmic oligosaccharides. Science 231:48–51. https://doi.org/10.1126/science.3941890
Mine S, Watanabe M (2019) Structural insights into the molecular evolution of the archaeal exo-β-d-glucosaminidase. Int J Mol Sci 20:2460. https://doi.org/10.3390/ijms20102460
Moreno E (2014) Retrospective and prospective perspectives on zoonotic brucellosis. Front Microbiol 5:213. https://doi.org/10.3389/fmicb.2014.00213
Murakami K, Nasu H, Fujiwara T, Takatsu N, Yoshida N, Furuta K, Kaito C (2021) The absence of osmoregulated periplasmic glucan confers antimicrobial resistance and increases virulence in Escherichia coli. J Bacteriol 203:e00515–e00520. https://doi.org/10.1128/JB.00515-20
Nakai H, Kitaoka M, Svensson B, Ohtsubo K (2013) Recent development of phosphorylases possessing large potential for oligosaccharide synthesis. Curr Opin Chem Biol 17:301–309. https://doi.org/10.1016/j.cbpa.2013.01.006
Nakajima M, Toyoizumi H, Abe K, Nakai H, Taguchi H, Kitaoka M (2014) 1,2-β-oligoglucan phosphorylase from Listeria innocua. PLoS ONE 9:e92353. https://doi.org/10.1371/journal.pone.0092353
Nakajima M, Tanaka N, Furukawa N, Nihira T, Kodutsumi Y, Takahashi Y, Sugimoto N, Miyanaga A, Fushinobu S, Taguchi H, Nakai H (2017) Mechanistic insight into the substrate specificity of 1,2-β-oligoglucan phosphorylase from Lachnoclostridium phytofermentans. Sci Rep 7:42671. https://doi.org/10.1038/srep42671
Nakajima M, Tanaka N, Kobayashi K, Nakai H, Kimura S, Iwata T, Taguchi H (2021) Enzymatic control and evaluation of degrees of polymerization of β-(1→2)-glucans. Anal Biochem 632:114366. https://doi.org/10.1016/j.ab.2021.114366
Nakano T, Oka K, Hanba K, Morita S (1996) Intratumoral administration of sizofiran activates Langerhans cell and T-cell infiltration in cervical cancer. Clin Immunol Immunopathol 79:79–86. https://doi.org/10.1006/clin.1996.0053
Nakashima A, Yamada K, Iwata O, Sugimoto R, Atsuji K, Ogawa T, Ishibashi-Ohgo N, Suzuki K (2018) β-Glucan in foods and its physiological functions. J Nutr Sci Vitaminol 64:8–17. https://doi.org/10.3177/jnsv.64.8
Nam YW, Nihira T, Arakawa T, Saito Y, Kitaoka M, Nakai H, Fushinobu S (2015) Crystal structure and substrate recognition of cellobionic acid phosphorylase, which plays a key role in oxidative cellulose degradation by microbes. J Biol Chem 290:18281–18292. https://doi.org/10.1074/jbc.M115.664664
Nelson TE, Lewis BA (1974) Separation and characterization of the soluble and insoluble components of insoluble laminaran. Carbohydr Res 33:63–74. https://doi.org/10.1016/S0008-6215(00)82940-7
Nihira T, Saito Y, Kitaoka M, Nishimoto M, Otsubo K, Nakai H (2012) Characterization of a laminaribiose phosphorylase from Acholeplasma laidlawii PG-8A and production of 1,3-β-d-glucosyl disaccharides. Carbohydr Res 361:49–54. https://doi.org/10.1016/j.carres.2012.08.006
Nihira T, Saito Y, Nishimoto M, Kitaoka M, Igarashi K, Ohtsubo K, Nakai H (2013) Discovery of cellobionic acid phosphorylase in cellulolytic bacteria and fungi. FEBS Lett 587:3556–3561. https://doi.org/10.1016/j.febslet.2013.09.014
Olafsdottir ES, Ingólfsdottir K (2001) Polysaccharides from lichens: structural characteristics and biological activity. Planta Med 67:199–208. https://doi.org/10.1055/s-2001-12012
Peters NK, Frost JW, Long SR (1986) A plant flavone, luteolin, induces expression of Rhizobium meliloti nodulation genes. Science 233:977–980. https://doi.org/10.1126/science.3738520
Piao J, Janga A, Choi Y, Tahira MN, Kim Y, Park S, Cho E, Jung S (2014) Solubility enhancement of α-naphthoflavone by synthesized hydroxypropyl cyclic-(1→2)-β-d-glucans (cyclosophoroases). Carbohydr Polym 101:733–740. https://doi.org/10.1016/j.carbpol.2013.09.104
Pluvinage B, Fillo A, Massel P, Boraston AB (2017) Structural analysis of a family 81 glycoside hydrolase implicates its recognition of β-1,3-glucan quaternary structure. Structure 25:1348–1359. https://doi.org/10.1016/j.str.2017.06.019
Poletaev VI (2006) New species of spiriferids (Brachiopoda) from the Devonian and Carboniferous of Eastern Europe. Paleontol J 40:507–517. https://doi.org/10.1134/S0031030106050054
Popper ZA, Michel G, Hervé C, Domozych DS, Willats WGT, Tuohy MG, Kloareg B, Stengel DB (2011) Evolution and diversity of plant cell walls: From algae to flowering plants. Annu Rev Plant Biol 62:567–590. https://doi.org/10.1146/annurev-arplant-042110-103809
Putman EW, Potter AL, Hodgson R, Hassid WZ (1950) The structure of crown-gall polysaccharide. I J Am Chem Soc 72:5024–5026. https://doi.org/10.1021/ja01167a055
Rajashekhara E, Kitaoka M, Kim YK, Hayashi K (2002) Characterization of a cellobiose phosphorylase from a hyperthermophilic eubacterium, Thermotoga maritima MSB8. Biosci Biotechnol Biochem 66:2578–2586. https://doi.org/10.1271/bbb.66.2578
Reese ET, Parrish FW, Mandels M (1961) β-d-1,2-Glucanases in fungi. Can J Microbiol 7:309–317. https://doi.org/10.1139/m61-038
Reichenbecher M, Lottspeich F, Bronnenmeier K (1997) Purification and properties of a cellobiose phosphorylase (CepA) and a cellodextrin phosphorylase (CepB) from the cellulolytic thermophile Clostridium stercorarium. Eur J Biochem 247:262–267. https://doi.org/10.1111/j.1432-1033.1997.00262.x
Rigano LA, Payette C, Brouillard G, Marano MR, Abramowicz L, Torres PS, Yun M, Castagnaro AP, Oirdi ME, Dufour V, Malamud F, Dow JM, Bouarab K, Vojnov AA (2007) Bacterial cyclic β-(1,2)-glucan acts in systemic suppression of plant immune responses. Plant Cell 19:2077–2089. https://doi.org/10.1105/tpc.106.047944
Roset MS, Ciocchini AE, Ugalde RA, Iñón de Iannino N (2004) Molecular cloning and characterization of cgt, the Brucella abortus cyclic β-1,2-glucan transporter gene, and its role in virulence. Infect Immun 72:2263–2271. https://doi.org/10.1128/IAI.72.4.2263-2271.2004
Roset MS, Ciocchini AE, Ugalde RA, Iñón de Iannino N (2006) The Brucella abortus cyclic β-1,2-glucan virulence factor is substituted with O-ester-linked succinyl residues. J Bacteriol 188:5003–5013. https://doi.org/10.1128/JB.00086-06
Roset MS, Ibãnez AE, De Souza Filho JA, Spera JM, Minatel L, Oliveira SC, Giambartolomei GH, Cassataro J, Briones G (2014) Brucella cyclic β-1,2-glucan plays a critical role in the induction of splenomegaly in mice. PLoS ONE 9:e101279. https://doi.org/10.1371/journal.pone.0101279
Ruiz-Herrera J, Ortiz-Castellanos L (2019) Cell wall glucans of fungi. A review. Cell Surf 5:100022. https://doi.org/10.1016/j.tcsw.2019.100022
Saburi W, Nihira T, Nakai H, Kitaoka M, Mori H (2022) Discovery of solabiose phosphorylase and its application for enzymatic synthesis of solabiose from sucrose and lactose. Sci Rep 12:259. https://doi.org/10.1038/s41598-021-04421-2
Sakamoto Y, Watanabe H, Nagai M, Nakade K, Takahashi M, Sato T (2006) Lentinula edodes tlg1 encodes a thaumatin-like protein that is involved in lentinan degradation and fruiting body senescence. Plant Physiol 141:793–801. https://doi.org/10.1104/pp.106.076679
Sakamoto Y, Nakade K, Konno N (2011) Endo-β-1,3-Glucanase GLU1, from the fruiting body of Lentinula edodes, belongs to a new glycoside hydrolase family. Appl Environ Microbiol 77:8350–8354. https://doi.org/10.1128/AEM.05581-11
Schneider JE, Reinhold V, Rumley MK, Kennedy EP (1979) Structural studies of the membrane-derived oligosaccharides of Escherichia coli. J Biol Chem 254:10135–10138. https://doi.org/10.1093/protein/3.3.193
Shen SH, Chretien P, Bastien L, Slilaty SN (1991) Primary sequence of the glucanase gene from Oerskovia xanthineolytica. Expression and purification of the enzyme from Escherichia coli. J Biol Chem 266:1058–1063. https://doi.org/10.1016/s0021-9258(17)35282-1
Shimizu H, Nakajima M, Miyanaga A, Takahashi Y, Tanaka N, Kobayashi K, Sugimoto N, Nakai H, Taguchi H (2018) Characterization and structural analysis of a novel exo-type enzyme acting on β-1,2-glucooligosaccharides from Parabacteroides distasonis. Biochemistry 57:3849–3860. https://doi.org/10.1021/acs.biochem.8b00385
Sierks MR, Ford C, Reilly PJ, Svensson B (1990) Catalytic mechanism of fungal glucoamylase as defined by mutagenesis of Asp176, Glu179 and Glu180 in the enzyme from Aspergillus awamori. Protein Eng 3:193–198. https://doi.org/10.1093/protein/3.3.193
Stanfield SW, Ielpi L, O’Brochta D, Helinski DR, Ditta GS (1988) The ndvA gene product of Rhizobium meliloti is required for β-(1→2)glucan production and has homology to the ATP-binding export protein HlyB. J Bacteriol 170:3523–3530. https://doi.org/10.1128/jb.170.8.3523-3530.1988
Starr T, Ng TW, Wehrly TD, Knodler LA, Celli J (2008) Brucella intracellular replication requires trafficking through the late endosomal/lysosomal compartment. Traffic 9:678–694. https://doi.org/10.1111/j.1600-0854.2008.00718.x
Talaga P, Fournet B, Bohin JP (1994) Periplasmic glucans of Pseudomonas syringae pv. syringae. J Bacteriol 176:6538–6544. https://doi.org/10.1128/jb.176.21.6538-6544.1994
Tanaka N, Nakajima M et al (2019) Identification, characterization, and structural analyses of a fungal endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem 294:7942–7965. https://doi.org/10.1074/jbc.RA118.007087
Tanaka T, Fukui T, Atomi H, Imanaka T (2003) Characterization of an exo-β-d-glucosaminidase involved in a novel chitinolytic pathway from the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J Bacteriol 185:5175–5181. https://doi.org/10.1128/JB.185.17.5175-5181.2003
Tang L, Weissborn AC, Kennedy EP (1997) Domains of Escherichia coli acyl carrier protein important for membrane- derived-oligosaccharide biosynthesis. J Bacteriol 179:3697–3705. https://doi.org/10.1128/jb.179.11.3697-3705.1997
Temple MJ, Cuskin F, Baslé A, Hickey N, Speciale G, Williams SJ, Gilbert HJ, Lowe EC (2017) A Bacteroidetes locus dedicated to fungal 1,6-β-glucan degradation: Unique substrate conformation drives specificity of the key endo-1,6-β-glucanase. J Biol Chem 292:10639–10650. https://doi.org/10.1074/jbc.M117.787606
Terrapon N, Lombard V, Drula É, Lapébie P, Al-Masaudi S, Gilbert HJ, Henrissat B (2018) PULDB: The expanded database of Polysaccharide Utilization Loci. Nucleic Acids Res 46:D677–D683. https://doi.org/10.1093/nar/gkx1022
Teufel F, Almagro Armenteros JJ, Johansen AR, Gíslason MH, Pihl SI, Tsirigos KD, Winther O, Brunak S, von Heijne G, Nielsen H (2022) SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat Biotechnol 40:1023–1025. https://doi.org/10.1038/s41587-021-01156-3
Therisod H, Weissborn AC, Kennedy EP (1986) An essential function for acyl carrier protein in the biosynthesis of membrane-derived oligosaccharides of Escherichia coli. Proc Natl Acad Sci U S A 83:7236–7240. https://doi.org/10.1073/pnas.83.19.7236
Therisod H, Kennedy EP (1987) The function of acyl carrier protein in the synthesis of membrane-derived oligosaccharides does not require its phosphopantetheine prosthetic group. Proc Natl Acad Sci U S A 84:8235–8238. https://doi.org/10.1073/pnas.84.23.8235
Ubiparip Z, De Doncker M, Beerens K, Franceus J, Desmet T (2021) β-Glucan phosphorylases in carbohydrate synthesis. Appl Microbiol Biotechnol 105:4073–4087. https://doi.org/10.1007/s00253-021-11320-z
Van Hoorebeke A, Stout J, Kyndt J, De Groeve M, Dix I, Desmet T, Soetaert W, Van Beeumen J, Savvides SN (2010) Crystallization and X-ray diffraction studies of cellobiose phosphorylase from Cellulomonas uda. Acta Crystallogr Sect F Struct Biol Cryst Commun 66:346–351. https://doi.org/10.1107/S1744309110002642
Venkatachalam G, Srinivasan D, Doble M (2013) Cyclic β-(1,2)-glucan production by Rhizobium meliloti MTCC 3402. Process Biochem 48:1848–1854. https://doi.org/10.1016/j.procbio.2013.08.024
Wang D, Kim DH, Seo N, Yun EJ, An HJ, Kim J-HK, Kim KH (2016) A novel glycoside hydrolase family 5 β-1,3 – 1,6-endoglucanase from Saccharophagus degradans 2-40T and its transglycosylase activity. Appl Environ Microbiol 82:4340–4349. https://doi.org/10.1128/AEM.00635-16
Warnick TA, Methé BA, Leschine SB (2002) Clostridium phytofermentans sp. nov., a cellulolytic mesophile from forest soil. Int J Syst Evol Microbiol 52:1155–1160. https://doi.org/10.1099/00207713-52-4-1155
Weissbornl AC, Kennedy P (1984) Biosynthesis of membrane-derived oligosaccharides. Novel glucosyltransferase system from Escherichia coli for the elongation of β 1→2-linked polyglucose chains. J Biol Chem 259:12644–12651. https://doi.org/10.1016/S0021-9258(18)90794-5
Wu CC, Zhang HT, Gao ZX, Qu JJ, Zhu L, Zhan XB (2022) Enhanced solubility of curcumin by complexation with fermented cyclic β-1,2-glucans. J Pharm Biomed Anal 211:114613. https://doi.org/10.1016/j.jpba.2022.114613
Xu C, Cao Q, Lan L (2021) Glucose-binding of periplasmic protein GltB activates GtrS-GltR two-component system in Pseudomonas aeruginosa. Microorganisms 9:447. https://doi.org/10.3390/microorganisms9020447
Zevenhuizen LPTM, Scholten-Koerselman HJ (1979) Surface carbohydrates of Rhizobium. Antonie Van Leeuwenhoek 45:165–175. https://doi.org/10.1007/BF00418581
Zhu L, Chen L (2019) Progress in research on paclitaxel and tumor immunotherapy. Cell Mol Biol Lett 24:1–11. https://doi.org/10.1186/s11658-019-0164-y
Zorreguieta A, Ugalde RA, Leloir LF (1985) An intermediate in cyclic β 1–2 glucan biosynthesis. Biochem Biophys Res Commun 126:352–357. https://doi.org/10.1016/0006-291x(85)90613-8
Zorreguieta A, Ugalde RA (1986) Formation in Rhizobium and Agrobacterium spp. of a 235-kilodalton protein intermediate in β-d(1–2) glucan synthesis. J Bacteriol 167:947–951. https://doi.org/10.1128/jb.167.3.947-951.1986
Acknowledgements
The author thanks Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This work was supported in part by JSPS KAKENHI Grant Number 20K05830. The author declares that there is no conflict of interest with the contents of this review article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nakajima, M. β-1,2-Glucans and associated enzymes. Biologia 78, 1741–1757 (2023). https://doi.org/10.1007/s11756-022-01205-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-022-01205-5