Abstract
Human immunodeficiency virus type 1 (HIV-1) is a major global health problem, with over 38 million people infected worldwide. Current anti-HIV-1 drugs are limited in their ability to prevent the virus from replicating inside host cells, making them less effective as preventive measures. In contrast, viral inhibitors that inactivate the virus before it can bind to a host cell have great potential as drugs. In this study, we aimed to design mutant peptides that could block the interaction between gp120 and the CD4 receptor on host cells, thus preventing HIV-1 infection. We designed a 20-amino-acid peptide that mimicked the amino acids of the CD4 binding site and docked it to gp120. Molecular dynamics simulations were performed to calculate the energy of MMPBSA (Poisson–Boltzmann Surface Area) for each residue of the peptide, and unfavorable energy residues were identified as potential mutation points. Using MAESTRO (Multi AgEnt STability pRedictiOn), we measured ΔΔG (change in the change in Gibbs free energy) for mutations and generated a library of 240 mutated peptides using OSPREY software. The peptides were then screened for allergenicity and binding affinity. Finally, molecular dynamics simulations (via GROMACS 2020.2) and control docking (via HADDOCK 2.4) were used to evaluate the ability of four selected peptides to inhibit HIV-1 infection. Three peptides, P3 (AHRQIRQWFLTRGPNRSLWQ), P4 (VHRQIRQWFLTRGPNRSLWQ), and P9 (AHRQIRQMFLTRGPNRSLWQ), showed practical and potential as HIV inhibitors, based on their binding affinity and ability to inhibit infection. These peptides have the ability to inactivate the virus before it can bind to a host cell, thus representing a promising approach to HIV-1 prevention. Our findings suggest that mutant peptides designed to block the interaction between gp120 and the CD4 receptor have potential as HIV-1 inhibitors. These peptides could be used as preventive measures against HIV-1 transmission, and further research is needed to evaluate their safety and efficacy in clinical settings.
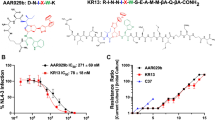
Graphical Abstract








Similar content being viewed by others
References
Dahl V (2013) Characterization of HIV-1 in the central nervous system during suppressive therapy. Karolinska Institutet, Sweden
Emuze BO, Jain MS, Luvsannyam E, Bhaya P, Vaquero C (2021) Central nervous system toxoplasmosis and cytomegalovirus colitis in an asymptomatic HIV positive patient. Cureus. https://doi.org/10.7759/cureus.17683
Bock PJ, Markovitz DM (2001) Infection with HIV-2. Aids 15(SUPPL. 5). https://doi.org/10.1097/00002030-200100005-00006
Laumaea A, Iii ABS, Sodroski J, Finzi A (2020) Opening the HIV envelope: potential of CD4 mimics as multifunctional HIV entry inhibitors. Current Opin HIV AIDS 15(5):300–308. https://doi.org/10.1097/COH.0000000000000637
Abbas AK, Lichtman AH, Pillai S, Baker DL, Baker A (2021) Cellular and molecular immunology E-book. Elsevier Health Sciences, United Kingdom. https://www.uk.elsevierhealth.com/cellular-and-molecular-immunology-9780323757485.html
Andrianov AM, Nikolaev GI, Kornoushenko YV, Xu W, Jiang S, Tuzikov A V (2019) In silico identification of novel aromatic compounds as potential HIV-1 entry inhibitors mimicking cellular receptor CD4. Viruses 11(8). https://doi.org/10.3390/v11080746
Sabzian-Molaei F, Hosseini S, Bolhasani A, Eskandari V, Norouzi S, Hadi A (2022) Antiviral effect of saffron compounds on the GP120 of HIV-1: an in silico study. Chem Select 7(47). https://doi.org/10.1002/slct.202203471
Kwong PD, Wyatt R, Robinson J, Sweet RW, Sodroski J, Hendrickson WA (1998) Structure of an HIV gp 120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393(6686):648–659. https://doi.org/10.1038/31405
U.S. Food and Drug Administration (2019) Human Immunodeficiency Virus-1 Infection: Developing Systemic Drug Products for Pre-Exposure Prophylaxis. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/human-immunodeficiency-virus-1-infection-developing-systemic-drug-products-pre-exposure-prophylaxis
De Clercq E (2005) New approaches toward anti-HIV chemotherapy. ChemInform 36(26). https://doi.org/10.1002/chin.200526245
Esté JA, Telenti A (2007) HIV entry inhibitors. Lancet 370(9581):81–88. https://doi.org/10.1016/S0140-6736(07)61052-6
Rusconi S, Scozzafava A, Mastrolorenzo A, Supuran CT (2007) An update in the development of HIV entry inhibitors. Curr Top Med Chem 7(13):1273–1289. https://doi.org/10.2174/156802607781212239
Ryser HJ-P, Flückiger R (2005) Progress in targeting HIV-1 entry. Drug Discov Today 10(16):1085–94. http://www.ncbi.nlm.nih.gov/pubmed/16182193
Adamson CS, Freed EO (2010) Novel approaches to inhibiting HIV-1 replication. Antiviral Res 85(1):119–141. https://doi.org/10.1016/j.antiviral.2009.09.009
Li W, Lu L, Li W, Jiang S (2017) Small-molecule HIV-1 entry inhibitors targeting gp120 and gp41: a patent review (2010–2015). Expert Opin Ther Pat 27(6):707–719. https://doi.org/10.1080/13543776.2017.1281249
Su S, Wang Q, Xu W, Yu F, Hua C, Zhu Y et al (2017) A novel HIV-1 gp41 tripartite model for rational design of HIV-1 fusion inhibitors with improved antiviral activity. AIDS 31(7):885–894. https://doi.org/10.1097/QAD.0000000000001415
Matthews T, Salgo M, Greenberg M, Chung J, DeMasi R, Bolognesi D (2004) Enfuvirtide: the first therapy to inhibit the entry of HIV-1 into host CD4 lymphocytes. Nat Rev Drug Discov 3(3):215–225. https://doi.org/10.1038/nrd1331
MacArthur RD, Novak RM (2008) Maraviroc: The first of a new class of antiretroviral agents. Clin Infect Dis 47(2):236–241. https://doi.org/10.1086/589289
Dawood R, Benjelloun F, Pin JJ, Kone A, Chanut B, Jospin F et al (2013) Generation of HIV-1 potent and broad neutralizing antibodies by immunization with postfusion HR1/HR2 complex. AIDS 27(5):717–730. https://doi.org/10.1097/QAD.0b013e32835cfca5
Su S, Ma Z, Hua C, Li W, Lu L, Jiang S (2017) Adding an artificial tail-anchor to a peptide-based HIV-1 fusion inhibitor for improvement of its potency and resistance profile. Molecules 22(11). https://doi.org/10.1097/QAD.0000000000001415
Su S, Rasquinha G, Du L, Wang Q, Xu W, Li W, et al (2019) A peptide-based HIV-1 fusion inhibitor with two tail-anchors and palmitic acid exhibits substantially improved in vitro and ex vivo anti-HIV-1 activity and prolonged in vivo half-life. Molecules 24(6). https://doi.org/10.3390/molecules24061134
Su S, Zhu Y, Ye S, Qi Q, Xia S, Ma Z, et al (2017) Creating an artificial tail anchor as a novel strategy to enhance the potency of peptide-based HIV fusion inhibitors. J Virol 91(1). https://doi.org/10.1128/jvi.01445-16
Ding X, Zhang X, Chong H, Zhu Y, Wei H, Wu X, et al (2017) Enfuvirtide (T20)-based lipopeptide is a potent HIV-1 cell fusion inhibitor: implications for viral entry and inhibition. J Virol 91(18). https://doi.org/10.1128/jvi.00831-17
Chong H, Zhu Y, Yu D, He Y (2018) Structural and functional characterization of membrane fusion inhibitors with extremely potent activity against human immunodeficiency virus type 1 (HIV-1), HIV-2, and simian immunodeficiency virus. J Virol 92(20). https://doi.org/10.1128/jvi.01088-18
Ingallinella P, Bianchi E, Ladwa NA, Wang YJ, Hrin R, Veneziano M et al (2009) Addition of a cholesterol group to an HIV-1 peptide fusion inhibitor dramatically increases its antiviral potency. Proc Natl Acad Sci U S A 106(14):5801–5806. https://doi.org/10.1073/pnas.0901007106
Louis JM, Bewley CA, Clore GM (2001) Design and properties of NCCG-gp41, a chimeric gp41 molecule with nanomolar HIV fusion inhibitory activity. J Biol Chem 276(31):29485–29489. https://doi.org/10.1074/jbc.C100317200
Tong P, Lu Z, Chen X, Wang Q, Yu F, Zou P et al (2013) An engineered HIV-1 gp41 trimeric coiled coil with increased stability and anti-HIV-1 activity: implication for developing anti-HIV microbicides. J Antimicrob Chemother 68(11):2533–2544. https://doi.org/10.1093/jac/dkt230
Ni L, Gao GF, Tien P (2005) Rational design of highly potent HIV-1 fusion inhibitory proteins: implication for developing antiviral therapeutics. Biochem Biophys Res Commun 332(3):831–836. https://doi.org/10.1016/j.bbrc.2005.05.037
Root MJ, Kay MS, Kim PS (2001) Protein design of an HIV-1 entry inhibitor. Science 291(5505):884–888. https://doi.org/10.1126/science.1057453
Qiu Z, Chong H, Yao X, Su Y, Cui S, He Y (2015) Identification and characterization of a subpocket on the N-trimer of HIV-1 Gp41: implication for viral entry and drug target. AIDS 29(9):1015–1024. https://doi.org/10.1097/QAD.0000000000000683
Eckert DM, Kim PS (2001) Design of potent inhibitors of HIV-1 entry from the gp41 N-peptide region. Proc Natl Acad Sci U S A 98(20):11187–11192. https://doi.org/10.1073/pnas.201392898
Izumi K, Nakamura S, Nakano H, Shimura K, Sakagami Y, Oishi S et al (2010) Characterization of HIV-1 resistance to a fusion inhibitor, N36, derived from the gp41 amino-terminal heptad repeat. Antiviral Res 87(2):179–186. https://doi.org/10.1016/j.antiviral.2010.04.011
Zhu Y, Lu L, Xu L, Yang H, Jiang S, Chen Y-H (2010) Identification of a gp41 core-binding molecule with homologous sequence of human TNNI3K-like protein as a novel human immunodeficiency virus type 1 entry inhibitor. J Virol 84(18):9359–9368. https://doi.org/10.1128/jvi.00644-10
Di RA, Ebralidze AK, Benoukraf T, Goff LA, Terragni J, Figueroa ME et al (2012) Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491(7424):406–412
Huang Y, Yu J, Lanzi A, Yao X, Andrews CD, Tsai L et al (2016) Engineered bispecific antibodies with exquisite HIV-1-neutralizing activity. Cell 165(7):1621–1631. https://doi.org/10.1016/j.cell.2016.05.024
Padte NN, Yu J, Huang Y, Ho DD (2018) Engineering multi-specific antibodies against HIV-1. Retrovirology 15(1). https://doi.org/10.1186/s12977-018-0439-9
West AP, Scharf L, Horwitz J, Klein F, Nussenzweig MC, Bjorkman PJ (2013) Computational analysis of anti-HIV-1 antibody neutralization panel data to identify potential functional epitope residues. Proc Natl Acad Sci U S A 110(26):10598–10603. https://doi.org/10.1073/pnas.1309215110
Huang J, Kang BH, Pancera M, Lee JH, Tong T, Feng Y et al (2014) Broad and potent HIV-1 neutralization by a human antibody that binds the gp41-gp120 interface. Nature 515(7525):138–142. https://doi.org/10.1038/nature13601
Kong R, Xu K, Zhou T, Acharya P, Lemmin T, Liu K et al (2016) Fusion peptide of HIV-1 as a site of vulnerability to neutralizing antibody. Science 352(6287):828–833. https://doi.org/10.1126/science.aae0474
Lazzarin A, Clotet B, Cooper D, Reynes J, Arastéh K, Nelson M et al (2003) Efficacy of enfuvirtide in patients infected with drug-resistant HIV-1 in Europe and Australia. N Engl J Med 348(22):2186–2195. https://doi.org/10.1056/nejmoa035211
Walmsley S, Henry K, Katlama C, Nelson M, Castagna A, Reynes J et al (2003) Enfuvirtide (T-20) cross-reactive glycoprotein 41 antibody does not impair the efficacy or safety of enfuvirtide. J Infect Dis 188(12):1827–1833. https://doi.org/10.1086/379810
Kilby JM, Lalezari JP, Eron JJ, Carlson M, Cohen C, Arduino RC et al (2002) The safety, plasma pharmacokinetics, and antiviral activity of subcutaneous enfuvirtide (T-20), a peptide inhibitor of gp41-mediated virus fusion. HIV-infected adults AIDS Res Hum Retroviruses 18(10):685–693. https://doi.org/10.1089/088922202760072294
Pu J, Wang Q, Xu W, Lu L, Jiang S (2019) Development of protein-and peptide-based hiv entry inhibitors targeting gp120 or gp41. Viruses 11(8) https://doi.org/10.3390/v11080705
Potapov V, Cohen M, Schreiber G (2009) Assessing computational methods for predicting protein stability upon mutation: good on average but not in the details. Protein Eng Des Sel 22(9):553–560. https://doi.org/10.1093/protein/gzp030
Su X, Wang Q, Wen Y, Jiang S, Lu L (2020) Protein- and peptide-based virus inactivators: inactivating viruses before their entry into cells. Front Microbiol 11 https://doi.org/10.3389/fmicb.2020.01063
Laskowski RA, Jabłońska J, Pravda L, Vařeková RS, Thornton JM (2018) PDBsum: structural summaries of PDB entries. Protein Sci 27(1):129–134. https://doi.org/10.1002/pro.3289
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2014) The I-TASSER suite: protein structure and function prediction. Nat Methods 12(1):7–8. https://doi.org/10.1038/nmeth.3213
Dominguez C, Boelens R, Bonvin AMJJ (2003) HADDOCK: A protein-protein docking approach based on biochemical or biophysical information. J Am Chem Soc 125(7):1731–1737. https://doi.org/10.1021/ja026939x
Schrodinger LLC (2015) The PyMOL molecular graphics system. Version 1:8
Xue LC, Rodrigues JP, Kastritis PL, Bonvin AM, Vangone A (2026) PRODIGY: a web server for predicting the binding affinity of protein-protein complexes. Bioinformatics 32(23):3676–3678. https://doi.org/10.1093/bioinformatics/btw514
Abraham MJ, Roland Schulz TM, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25
Sabzian-Molaei F, NasiriKhalili MA, Sabzian-Molaei M, Shahsavarani H, Fattah Pour A, Molaei Rad A et al (2022) Urtica dioica Agglutinin: a plant protein candidate for inhibition of SARS-COV-2 receptor-binding domain for control of Covid19 Infection. PLoS One 17(7):e0268156. https://doi.org/10.1371/journal.pone.0268156
Sabzian-Molaei F, Hosseini S, Alipour A, Ghaderi H, Fotouhi-Chahouki F, Hadi A, et al (2023) Urtica dioica agglutinin (UDA) as a potential candidate for inhibition of SARS-CoV-2 Omicron variants: in silico prediction and experimental validation. Phytomedicine 111. https://doi.org/10.1016/j.phymed.2023.154648
Kumari R, Kumar R, Lynn A (2014) G-mmpbsa -A GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54(7):1951–1962. https://doi.org/10.1021/ci500020m
Laimer J, Hofer H, Fritz M, Wegenkittl S, Lackner P (2015) MAESTRO - multi agent stability prediction upon point mutations. BMC Bioinformatics 16(1)
Hallen MA, Martin JW, Ojewole A, Jou JD, Lowegard AU, Frenkel MS et al (2018) OSPREY 3.0: open-source protein redesign for you, with powerful new features. J Comput Chem 39(30):2494–507. https://doi.org/10.1002/jcc.25522
Lowegard AU, Frenkel MS, Holt GT, Jou JD, Ojewole AA, Donald BR (2020) Novel, provable algorithms for efficient ensemble-based computational protein design and their application to the redesign of the c-Raf-RBD:KRas protein-protein interface. PLoS Comput Biol 16(6). https://doi.org/10.1371/journal.pcbi.1007447
MdMehedi H, Nalini S, Shaherin B, Gwang L, Watshara S, Balachandran M (2020) HLPpred-Fuse: improved and robust prediction of hemolytic peptide and its activity by fusing multiple feature representation. Bioinformatics 36(11):3350–3356
Dimitrov I, Bangov I, Flower DR, Doytchinova I (2014) AllerTOP v.2 - a server for in silico prediction of allergens. J Mol Model 20(6). https://doi.org/10.1007/s00894-014-2278-5
Wei L, Ye X, Sakurai T, Mu Z, Wei L (2022) ToxIBTL: prediction of peptide toxicity based on information bottleneck and transfer learning. Bioinformatics 38(6):1514–1524. https://doi.org/10.1093/bioinformatics/btac006
Urry DW (2004) The change in Gibbs free energy for hydrophobic association: derivation and evaluation by means of inverse temperature transitions. Chem Phys Lett 399(1–3):177–183. https://doi.org/10.1016/j.cplett.2004.09.137
Weinstein B, Fenselau AH (1964) Amino acids and peptides. J Chromatogr A 15:149–152. https://doi.org/10.1016/s0021-9673(01)82761-8
Acknowledgements
Some parts of the graphical abstract were obtained from Vecteezy graphics: Free vector art via https://www.vecteezy.com; Vector illustration credit: https://www.vecteezy.com.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sabzian-Molaei, F., Ahmadi, M.A., Nikfarjam, Z. et al. Inactivation of cell-free HIV-1 by designing potent peptides based on mutations in the CD4 binding site. Med Biol Eng Comput 62, 423–436 (2024). https://doi.org/10.1007/s11517-023-02950-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02950-8