Abstract
Due to high computational requirements, deep-learning decoders for motor imaginary (MI) electroencephalography (EEG) signals are usually implemented on bulky and heavy computing devices that are inconvenient for physical actions. To date, the application of deep-learning techniques in independent portable brain-computer-interface (BCI) devices has not been extensively explored. In this study, we proposed a high-accuracy MI EEG decoder by incorporating spatial-attention mechanism into convolution neural network (CNN), and deployed it on fully integrated single-chip microcontroller unit (MCU). After the CNN model was trained on workstation computer using GigaDB MI datasets (52 subjects), its parameters were then extracted and converted to build deep-learning architecture interpreter on MCU. For comparison, EEG-Inception model was also trained using the same dataset, and was deployed on MCU. The results indicate that our deep-learning model can independently decode imaginary left-/right-hand motions. The mean accuracy of the proposed compact CNN reaches 96.75 ± 2.41% (8 channels: Frontocentral3 (FC3), FC4, Central1 (C1), C2, Central-Parietal1 (CP1), CP2, C3, and C4), versus 76.96 ± 19.08% of EEG-Inception (6 channels: FC3, FC4, C1, C2, CP1, and CP2). To the best of our knowledge, this is the first portable deep-learning decoder for MI EEG signals. The findings demonstrate high-accuracy deep-learning decoding of MI EEG in a portable mode, which has great implications for hand-disabled patients. Our portable system can be used for developing artificial-intelligent wearable BCI devices, as it is less computationally expensive and convenient for real-life application.
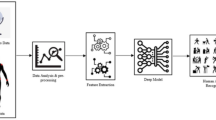
Graphical Abstract










Similar content being viewed by others
References
Xygonakis I, Athanasiou A, Pandria N et al (2018) Decoding motor imagery through common spatial pattern filters at the EEG source space. Comput Intell Neurosci 2018:7957408. https://doi.org/10.1155/2018/7957408
Qi FF, Li YQ, Wu W (2015) RSTFC: a novel algorithm for spatio-temporal filtering and classification of single-trial EEG. IEEE Trans Neural Netw Learn Syst 26(12):3070–3082. https://doi.org/10.1109/TNNLS.2015.2402694
Lesenfants D, Vanthornhout J, Verschueren E, Francart T (2019) Data-driven spatial filtering for improved measurement of cortical tracking of multiple representations of speech. J Neural Eng 16(6):066017. https://doi.org/10.1088/1741-2552/ab3c92
Wang L, Huang WJ, Yang Z, Zhang C (2020) Temporal-spatial-frequency depth extraction of brain-computer interface based on mental tasks. Biomed Signal Process Control 58:101845. https://doi.org/10.1016/j.bspc.2020.101845
Tautan AM, Rossi AC, de Francisco R, Ionescu B (2021) Dimensionality reduction for EEG-based sleep stage detection: comparison of autoencoders, principal component analysis and factor analysis. Biomed Eng-Biomed Tech 66(2):125–136. https://doi.org/10.1515/bmt-2020-0139
Garcia-Laencina PJ, Rodriguez-Bermudez G, Roca-Dorda J (2014) Exploring dimensionality reduction of EEG features in motor imagery task classification. Expert Syst Appl 41(11):5285–5295. https://doi.org/10.1016/j.eswa.2014.02.043
Haufe S, Dahne S, Nikulin VV (2014) Dimensionality reduction for the analysis of brain oscillations. Neuroimage 101:583–597. https://doi.org/10.1016/j.neuroimage.2014.06.073
Artoni F, Delorme A, Makeig S (2018) Applying dimension reduction to EEG data by principal component analysis reduces the quality of its subsequent independent component decomposition. Neuroimage 175:176–187. https://doi.org/10.1016/j.neuroimage.2018.03.016
Demuru M, La Cava SM, Pani SM, Fraschini M (2020) A comparison between power spectral density and network metrics: an EEG study. Biomed Signal Process Control 57:101760. https://doi.org/10.1016/j.bspc.2019.101760
Wang RF, Wang J, Yu HT, Wei XL, Yang C, Deng B (2015) Power spectral density and coherence analysis of Alzheimer’s EEG. Cogn Neurodyn 9(3):291–304. https://doi.org/10.1007/s11571-014-9325-x
Petroff OA, Spencer DD, Goncharova II, Zaveri HP (2016) A comparison of the power spectral density of scalp EEG and subjacent electrocorticograms. Clin Neurophysiol 127(2):1108–1112. https://doi.org/10.1016/j.clinph.2015.08.004
Jiang AM, Shang J, Liu XF, Tang YB, Kwan HK, Zhu YP (2020) Efficient CSP algorithm with spatio-temporal filtering for motor imagery classification. IEEE Trans Neural Syst Rehabil Eng 28(4):1006–1016. https://doi.org/10.1109/TNSRE.2020.2979464
Mishuhina V, Jiang XD (2021) Complex common spatial patterns on time-frequency decomposed EEG for brain-computer interface. Pattern Recog 115:107918. https://doi.org/10.1016/j.patcog.2021.107918
Guo Y, Zhang Y, Chen ZQ, Liu Y, Chen W (2020) EEG classification by filter band component regularized common spatial pattern for motor imagery. Biomed Signal Process Control 59:101917. https://doi.org/10.1016/j.bspc.2020.101917
Cote-Allard U, Fall CL, Drouin A, Campeau-Lecours A, Gosselin C, Glette K, Laviolette F, Gosselin B (2019) Deep learning for electromyographic hand gesture signal classification using transfer learning. IEEE Trans Neural Syst Rehabil Eng 27(4):760–771. https://doi.org/10.1109/TNSRE.2019.2896269
Kannathal N, Choo ML, Acharya UR, Sadasivan PK (2005) Entropies for detection of epilepsy in EEG. Comput Methods Programs Biomed 80(3):187–194. https://doi.org/10.1016/j.cmpb.2005.06.012
Liang ZH, Wang YH, Sun X, Li D, Voss LJ, Sleigh JW, Hagihira S, Li X (2015) EEG entropy measures in anesthesia. Front Comput Neurosci 9:16. https://doi.org/10.3389/fncom.2015.00016
Molina V, Lubeiro A, de Luis GR, Gomez-Pilar J, Martin-Santiago O, Igleaias-Tejedor M, Holgado-Madera P, Segarra-Echeverria R, Recio-Barbero M, Nunez P, Haidar MK, Fernandez-Sevillano J, Sanz-Fuentenebro J (2020) Deficits of entropy modulation of the EEG: a biomarker for altered function in schizophrenia and bipolar disorder? J Psychiatry Neurosci 45(5):322–333. https://doi.org/10.1503/jpn.190032
Xu Q, Zhou H, Wang YJ, Huang J (2009) Fuzzy support vector machine for classification of EEG signals using wavelet-based features. Med Eng Phys 31(7):858–865. https://doi.org/10.1016/j.medengphy.2009.04.005
Jrad N, Congedo M, Phlypo R, Rousseau S, Flamary R, Yger F, Rakotomamonjy A (2011) sw-SVM: sensor weighting support vector machines for EEG-based brain-computer interfaces. J Neural Eng 8(5):056004. https://doi.org/10.1088/1741-2560/8/5/056004
Kumar B, Gupta D (2021) Universum based Lagrangian twin bounded support vector machine to classify EEG signals. Comput Methods Prog Biomed 208:106244. https://doi.org/10.1016/j.cmpb.2021.106244
Wang JR, Bi LZ, Fei WJ, Guan CT (2021) Decoding single-hand and both-hand movement directions from noninvasive neural signals. IEEE Trans Biomed Eng 68(6):1932–1940. https://doi.org/10.1109/TBME.2020.3034112
Erla S, Faes L, Tranquillini E, Orrico D, Nollo G (2011) k-Nearest neighbour local linear prediction of scalp EEG activity during intermittent photic stimulation. Med Eng Phys 33(4):504–512. https://doi.org/10.1016/j.medengphy.2010.12.003
Li M, Xu HP, Liu XW, Lu SF (2018) Emotion recognition from multichannel EEG signals using K-nearest neighbor classification. Technol Health Care 26:509–519. https://doi.org/10.3233/THC-174836
Bablani A, Edla DR, Dodia S (2018) Classification of EEG data using k-nearest neighbor approach for concealed information Test. 8th Int Conf Adv Comput Commun (Icacc-2018) 143:242–249
Edla DR, Mangalorekar K, Dhavalikar G, Dodia S (2018) Classification of EEG data for human mental state analysis using random forest classifier. Procedia Comput Sci 132:1523–1532. https://doi.org/10.1016/j.procs.2018.10.392
Fraiwan L, Lweesy K, Khasawneh N, Wenz H, Dickhaus H (2012) Automated sleep stage identification system based on time-frequency analysis of a single EEG channel and random forest classifier. Comput Methods Programs Biomed 108(1):10–19. https://doi.org/10.1016/j.cmpb.2011.11.005
Al-Saegh A, Dawwd SA, Abdul-Jabbar JM (2021) Deep learning for motor imagery EEG-based classification: a review. Biomed Signal Process Control 63:102172. https://doi.org/10.1016/j.bspc.2020.102172
Lawhern VJ, Solon AJ, Waytowich NR, Gordon SM, Hung CP, Lance BJ (2018) EEGNet: a compact convolutional neural network for EEG-based brain-computer interfaces. J Neural Eng 15(5):056013. https://doi.org/10.1088/1741-2552/aace8c
Izzuddin TA, Safri NM, Othman MA (2021) Compact convolutional neural network (CNN) based on SincNet for end-to-end motor imagery decoding and analysis. Biocybernetics Biomed Eng 41(4):1629–1645. https://doi.org/10.1016/j.bbe.2021.10.001
Mumtaz W, Rasheed S, Irfan A (2021) Review of challenges associated with the EEG artifact removal methods. Biomed Signal Process Control 68:102741. https://doi.org/10.1016/j.bspc.2021.102741
Craik A, He YT, Contreras-Vidal JL (2019) Deep learning for electroencephalogram (EEG) classification tasks: a review. J Neural Eng 16(3):031001. https://doi.org/10.1088/1741-2552/ab0ab5
Gao Z, Dang WD, Wang XM, Hong X, Hou LH, Ma K, Perc M (2021) Complex networks and deep learning for EEG signal analysis. Cogn Neurodyn 15(3):369–388. https://doi.org/10.1007/s11571-020-09626-1
Chiarelli AM, Croce P, Merla A, Zappasodi F (2018) Deep learning for hybrid EEG-fNIRS brain-computer interface: application to motor imagery classification. J Neural Eng 15(3):036028. https://doi.org/10.1088/1741-2552/aaaf82
Wang P, Jiang AM, Liu XF, Shang J, Zhang L (2018) LSTM-based EEG classification in motor imagery tasks. IEEE Trans Neural Syst Rehabil Eng 26(11):2086–2095. https://doi.org/10.1109/TNSRE.2018.2876129
Kant P, Laskar SH, Hazarika J, Mahamune R (2020) CWT based transfer learning for motor imagery classification for brain computer interfaces. J Neurosci Methods 345:108886. https://doi.org/10.1016/j.jneumeth.2020.108886
Han Y, Wang B, Luo J, Li L, Li X (2022) A classification method for EEG motor imagery signals based on parallel convolutional neural network. Biomed Signal Process Control 71:103190. https://doi.org/10.1016/j.bspc.2021.103190
Xiao X, Fang Y (2021) Motor imagery EEG signal recognition using deep convolution neural network. Front Neurosci 15:655599. https://doi.org/10.3389/fnins.2021.655599
Santamaría-Vázquez E, Martínez-Cagigal V, Vaquerizo-Villar F, Hornero R (2020) EEG-Inception: a novel deep convolutional neural network for assistive ERP-based brain-computer interfaces. IEEE Trans Neural Syst Rehabil Eng 28(12):2773–2782. https://doi.org/10.1109/TNSRE.2020.3048106
Tabar YR, Halici U (2017) A novel deep learning approach for classification of EEG motor imagery signals. J Neural Eng 14(1):016003. https://doi.org/10.1088/1741-2560/14/1/016003
Dose H, Moller JS, Iversen HK, Puthusserypady S (2018) An end-to-end deep learning approach to MI-EEG signal classification for BCIs. Expert Syst Appl 114:532–542. https://doi.org/10.1016/j.eswa.2018.08.031
Amin SU, Alsulaiman M, Muhammad G, Mekhtiche MA, Hossain MS (2019) Deep learning for EEG motor imagery classification based on multi-layer CNNs feature fusion. Futur Gener Comput Syst- Int J Escience 101:542–554. https://doi.org/10.1016/j.future.2019.06.027
Zhang RL, Zong Q, Dou LQ, Zhao XY, Tang YF, Li ZY (2021) Hybrid deep neural network using transfer learning for EEG motor imagery decoding. Biomed Signal Process Control 63:102144. https://doi.org/10.1016/j.bspc.2020.102144
Karacsony T, Hansen JP, Iversen HK, Puthusserypady S (2019) Brain computer interface for neuro-rehabilitation with deep learning classification and virtual reality feedback. Proc 10th Augmented Human Int Conf 2019 (Ah2019). https://doi.org/10.1145/3311823.3311864
Lun XM, Yu ZL, Chen T, Wang F, Hou YM (2020) A simplified CNN classification method for MI-EEG via the electrode pairs signals. Front Human Neurosci 14:338. https://doi.org/10.3389/fnhum.2020.00338
Zhang H, Zhao X, Wu ZX, Sun B, Li T (2021) Motor imagery recognition with automatic EEG channel selection and deep learning. J Neural Eng 18(1):016004. https://doi.org/10.1088/1741-2552/abca16
Cho H, Ahn M, Ahn S, Kwon M, Jun SC (2017) EEG datasets for motor imagery brain-computer interface. Gigascience 6(7):1–8. https://doi.org/10.1088/1741-2552/abca16
Tsuchimoto S, Shibusawa S, Lwama S, Hayashi M, Okuyama K, Mizuguchi N, Kato K, Ushiba J (2021) Use of common average reference and large-Laplacian spatial-filters enhances EEG signal-to-noise ratios in intrinsic sensorimotor activity. J Neurosci Methods 353:109089. https://doi.org/10.1016/j.jneumeth.2021.109089
Belwafi K, Gannouni S, Aboalsamh H, Mathkour H, Belghith A (2019) A dynamic and self-adaptive classification algorithm for motor imagery EEG signals. J Neurosci Methods 327:108346. https://doi.org/10.1016/j.jneumeth.2019.108346
Ramos-Arguelles F, Morales G, Egozeue S, Pabon RM, Alonso MT (2009) Basic techniques of electroencephalography: principles and clinical applications. Anales Del Sistema Sanitario De Navarra 32:69–82. https://doi.org/10.23938/ASSN.0148
Kaya I (2020) A brief summary of EEG artifact handling. arXiv, 2001:00693. https://doi.org/10.5772/intechopen.99127
Wen JH, Thibeau-Sutre E, Diaz-Melo M, Samper-Gonzalez J, Routier A, Bottani S, Dormont D, Durrleman S, Burgos N, Colliot O (2020) Convolutional neural networks for classification of Alzheimer’s disease: overview and reproducible evaluation. Med Image Anal 63:101694. https://doi.org/10.1016/j.media.2020.101694
Gu BZ, Ge RJ, Chen Y, Luo LM, Coatrieux G (2021) Automatic and robust object detection in X-Ray baggage inspection using deep convolutional neural networks. IEEE Trans Industr Electron 68(10):10248–10257. https://doi.org/10.1109/TIE.2020.3026285
Wang Y, Luo XB, Ding L, Fu S, Wei X (2019) Detection based visual tracking with convolutional neural network. Knowl-Based Syst 175:62–71. https://doi.org/10.1016/j.knosys.2019.03.012
Schirrmeister RT, Springenberg JT, Fiederer LDJ, Glasstetter M, Eggensperger K, Tangermann M, Hutter F, Burgard W, Ball T (2017) Deep learning with convolutional neural networks for EEG decoding and visualization. Hum Brain Mapp 38(11):5391–5420. https://doi.org/10.1002/hbm.23730
Suchetha M, Madhumitha R, SornaMeena M, Sruthi R (2021) Sequential convolutional neural networks for classification of cognitive tasks from EEG signals. Appl Soft Comput 111:107664. https://doi.org/10.1016/j.asoc.2021.107664
Amin SU, Alsulaiman M, Muhammad G, Mekhtiche MA, Hossain MS (2019) Deep Learning for EEG motor imagery classification based on multi-layer CNNs feature fusion. Futur Gener Comput Syst 101:542–554. https://doi.org/10.1016/j.future.2019.06.027
Zhang R, Zong Q, Dou L, Zhao X, Tang Y, Li Z (2021) Hybrid deep neural network using transfer learning for EEG motor imagery decoding. Biomed Signal Process Control 63:10144. https://doi.org/10.1016/j.bspc.2020.102144
David R, Duke J, Jain A, Reddi VJ, Jeffries N, Li J, Kreeger N, Nappier I, Natraj, M, Regev S, Rhodes R, Wang T, Warden P (2021) TensorFlow lite micro: embedded machine learning on TinyML systems. ArXiv, 2010:08678. https://doi.org/10.48550/arXiv.2010.08678
Ren HG, Bello S, Alam MN, Manivannan N, Shagam T, D’Souza N (2021) Optimizing inference performance of a fundus image quality neural network model for edge computing using TensorFlow Lite. Investig Ophthalmol Visual Sci 62:113
Tariq M, Trivailo PM, Simic M (2018) EEG-based BCI control schemes for lower-limb assistive-robots. Front Human Neurosci 12:312. https://doi.org/10.3389/fnhum.2018.00312
Luu T, Nakagome S, He Y, Contreras-Vidal J (2017) Real-time EEG-based brain-computer interface to a virtual avatar enhances cortical involvement in human treadmill walking. Sci Rep 7:8895. https://doi.org/10.1038/s41598-017-09187-0
Wang J, Wang M (2021) Review of the emotional feature extraction and classification using EEG signals. Cognitive Robotics 1:29–40. https://doi.org/10.1016/j.cogr.2021.04.001
Al-Qazzaz NK, Alyasseri ZAA, Abdulkareem KH, Ali NS, Al-Mhiqani MN, Guger C (2021) EEG feature fusion for motor imagery: a new robust framework towards stroke patients rehabilitation. Comput Biol Med 137:104799. https://doi.org/10.1016/j.compbiomed.2021.104799
Roy Y, Banville H, Albuquerque I, Gramfort A, Falk TH, Faubert J (2019) Deep learning-based electroencephalography analysis: a systematic review. J Neural Eng 16(5):051001. https://doi.org/10.1088/1741-2552/ab260c
Mahmood M, Mzurikwao D, Kim YS, Lee Y, Mishra S, Herbert R, Duarte A, Ang CS, Yeo WH (2019) Fully portable and wireless universal brain-machine interfaces enabled by flexible scalp electronics and deep learning algorithm. Nature Mach Intell 1(9):412–422. https://doi.org/10.1038/s42256-019-0091-7
Kranczioch C, Zich C, Schierholz I, Sterr A (2014) Mobile EEG and its potential to promote the theory and application of imagery-based motor rehabilitation. Int J Psychophysiol 91(1):10–15. https://doi.org/10.1016/j.ijpsycho.2013.10.004
Netzer E, Frid A, Feldman D (2020) Real-time EEG classification via coresets for BCI applications. Eng Appl Artif Intell 89:103455. https://doi.org/10.1016/j.engappai.2019.103455
Hou YM, Zhou L, Jia SY, Lun XM (2020) A novel approach of decoding EEG four-class motor imagery tasks via scout ESI and CNN. J Neural Eng 17(1):016048. https://doi.org/10.1088/1741-2552/ab4af6
Funding
This study is supported by the Natural Science Foundation of Zhejiang Province (LY20E070005, LY17E070007), “Pioneer” and “Leading Goose” R&D Program of Zhejiang (2022C01119), and National Natural Science Foundation of China (51207038, 62171169).
Author information
Authors and Affiliations
Contributions
Zhanxiong Wu: methodology, result analysis, writing, and supervision. Xudong Tang: EEG data preprocess, investigation. Jinhui Wu: CNN model construction and training. Jiye Huang: hardware, programming, and debugging. Jian Shen: EEG data analyzing. Hui Hong: review and language editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Z., Tang, X., Wu, J. et al. Portable deep-learning decoder for motor imaginary EEG signals based on a novel compact convolutional neural network incorporating spatial-attention mechanism. Med Biol Eng Comput 61, 2391–2404 (2023). https://doi.org/10.1007/s11517-023-02840-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02840-z