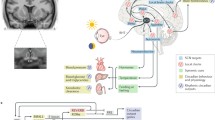
Abstract
The circadian clock coordinates rhythms in numerous physiological processes to maintain organismal homeostasis. Since the suprachiasmatic nucleus (SCN) is widely accepted as the circadian pacemaker, it is critical to understand the neural mechanisms by which rhythmic information is transferred from the SCN to peripheral clocks. Here, we present the first comprehensive map of SCN efferent connections and suggest a molecular logic underlying these projections. The SCN projects broadly to most major regions of the brain, rather than solely to the hypothalamus and thalamus. The efferent projections from different subtypes of SCN neurons vary in distance and intensity, and blocking synaptic transmission of these circuits affects circadian rhythms in locomotion and feeding to different extents. We also developed a barcoding system to integrate retrograde tracing with in-situ sequencing, allowing us to link circuit anatomy and spatial patterns of gene expression. Analyses using this system revealed that brain regions functioning downstream of the SCN receive input from multiple neuropeptidergic cell types within the SCN, and that individual SCN neurons generally project to a single downstream brain region. This map of SCN efferent connections provides a critical foundation for future investigations into the neural circuits underlying SCN-mediated rhythms in physiology. Further, our new barcoded tracing method provides a tool for revealing the molecular logic of neuronal circuits within heterogeneous brain regions.
Similar content being viewed by others
References
Abrahamson, E.E., and Moore, R.Y. (2001). Suprachiasmatic nucleus in the mouse: retinal innervation, intrinsic organization and efferent projections. Brain Res 916, 172–191.
Aton, S.J., Colwell, C.S., Harmar, A.J., Waschek, J., and Herzog, E.D. (2005). Vasoactive intestinal polypeptide mediates circadian rhythmicity and synchrony in mammalian clock neurons. Nat Neurosci 8, 476–483.
Bankhead, P., Loughrey, M.B., Fernández, J.A., Dombrowski, Y., McArt, D.G., Dunne, P.D., McQuaid, S., Gray, R.T., Murray, L.J., Coleman, H.G., et al. (2017). QuPath: open source software for digital pathology image analysis. Sci Rep 7, 16878.
Campos, L.M.G., Cruz-Rizzolo, R.J., Watanabe, I.S., Pinato, L., and Nogueira, M.I. (2014). Efferent projections of the suprachiasmatic nucleus based on the distribution of vasoactive intestinal peptide (VIP) and arginine vasopressin (AVP) immunoreactive fibers in the hypothalamus of Sapajus apella. J Chem Neuroanat 57–58, 42–53.
Chen, J., Wang, Z., Li, Z., Peng, D., and Fang, Y. (2021). Disturbances of affective cognition in mood disorders. Sci China Life Sci 64, 938–941.
Chen, X., Sun, Y.C., Zhan, H., Kebschull, J.M., Fischer, S., Matho, K., Huang, Z.J., Gillis, J., and Zador, A.M. (2019). High-throughput mapping of long-range neuronal projection using in situ sequencing. Cell 179, 772–786.e19.
Evans, J.A., Leise, T.L., Castanon-Cervantes, O., and Davidson, A.J. (2011). Intrinsic regulation of spatiotemporal organization within the suprachiasmatic nucleus. PLoS ONE 6, e15869.
Evans, J.A., Suen, T.C., Callif, B.L., Mitchell, A.S., Castanon-Cervantes, O., Baker, K. M., Kloehn, I., Baba, K., Teubner, B.J.W., Ehlen, J.C., et al. (2015). Shell neurons of the master circadian clock coordinate the phase of tissue clocks throughout the brain and body. BMC Biol 13, 43.
Fang, F., and Hu, H. (2021). Recent progress on mechanisms of human cognition and brain disorders. Sci China Life Sci 64, 843–846.
Freeman, J.G.M., Krock, R.M., Aton, S.J., Thaben, P., and Herzog, E.D. (2013). GABA networks destabilize genetic oscillations in the circadian pacemaker. Neuron 78, 799–806.
Gizowski, C., Zaelzer, C., and Bourque, C.W. (2016). Clock-driven vasopressin neurotransmission mediates anticipatory thirst prior to sleep. Nature 537, 685–688.
Harmar, A.J., Marston, H.M., Shen, S., Spratt, C., West, K.M., Sheward, W.J., Morrison, C.F., Dorin, J.R., Piggins, H.D., Reubi, J.C., et al. (2002). The VPAC2 receptor is essential for circadian function in the mouse suprachiasmatic nuclei. Cell 109, 497–508.
Huang, L., Kebschull, J.M., Fürth, D., Musall, S., Kaufman, M.T., Churchland, A.K., and Zador, A.M. (2020). BRICseq bridges brain-wide interregional connectivity to neural activity and gene expression in single animals. Cell 183, 2040.
Inouye, S.I.T., and Kawamura, H. (1979). Persistence of circadian rhythmicity in a mammalian hypothalamic “island” containing the suprachiasmatic nucleus.. Proc Natl Acad Sci USA 76, 5962–5966.
Ju, D., Zhang, W., Yan, J., Zhao, H., Li, W., Wang, J., Liao, M., Xu, Z., Wang, Z., Zhou, G., et al. (2020). Chemical perturbations reveal that RUVBL2 regulates the circadian phase in mammals. Sci Transl Med 12, eaba0769.
Ke, R., Mignardi, M., Pacureanu, A., Svedlund, J., Botling, J., Wählby, C., and Nilsson, M. (2013). In situ sequencing for RNA analysis in preserved tissue and cells. Nat Methods 10, 857–860.
Kebschull, J.M., Garcia da Silva, P., Reid, A.P., Peikon, I.D., Albeanu, D.F., and Zador, A.M. (2016). High-throughput mapping of single-neuron projections by sequencing of barcoded RNA. Neuron 91, 975–987.
Lee, I.T., Chang, A.S., Manandhar, M., Shan, Y., Fan, J., Izumo, M., Ikeda, Y., Motoike, T., Dixon, S., Seinfeld, J.E., et al. (2015). Neuromedin s-producing neurons act as essential pacemakers in the suprachiasmatic nucleus to couple clock neurons and dictate circadian rhythms. Neuron 85, 1086–1102.
LeGates, T.A., Fernandez, D.C., and Hattar, S. (2014). Light as a central modulator of circadian rhythms, sleep and affect. Nat Rev Neurosci 15, 443–454.
Liu, C., Wang, Y., Sun, X., Wang, Y., and Fang, F. (2023). Decoding six basic emotions from brain functional connectivity patterns. Sci China Life Sci 66, 835–847.
Maywood, E.S., Reddy, A.B., Wong, G.K.Y., O’Neill, J.S., O’Brien, J.A., McMahon, D. G., Harmar, A.J., Okamura, H., and Hastings, M.H. (2006). Synchronization and maintenance of timekeeping in suprachiasmatic circadian clock cells by neuropeptidergic signaling. Curr Biol 16, 599–605.
Mazuski, C., Chen, S.P., and Herzog, E.D. (2020). Different roles for VIP neurons in the neonatal and adult suprachiasmatic nucleus. J Biol Rhythms 35, 465–475.
Mieda, M., Ono, D., Hasegawa, E., Okamoto, H., Honma, K., Honma, S., and Sakurai, T. (2015). Cellular clocks in AVP neurons of the SCN are critical for interneuronal coupling regulating circadian behavior rhythm. Neuron 85, 1103–1116.
Mohawk, J.A., Green, C.B., and Takahashi, J.S. (2012). Central and peripheral circadian clocks in mammals. Annu Rev Neurosci 35, 445–462.
Moore, R.Y. (2013). The suprachiasmatic nucleus and the circadian timing system. Prog Mol Biol Transl Sci 119, 1–28.
Moore, R.Y., and Eichler, V.B. (1972). Loss of a circadian adrenal corticosterone rhythm following suprachiasmatic lesions in the rat. Brain Res 42, 201–206.
Morris, E.L., Patton, A.P., Chesham, J.E., Crisp, A., Adamson, A., and Hastings, M.H. (2021). Single-cell transcriptomics of suprachiasmatic nuclei reveal a Prokineticindriven circadian network. EMBO J 40, e108614.
Nagoshi, E., Saini, C., Bauer, C., Laroche, T., Naef, F., and Schibler, U. (2004). Circadian gene expression in individual fibroblasts. Cell 119, 693–705.
Ni, Y., and Li, J. (2021). Neural mechanisms of social learning and decision-making. Sci China Life Sci 64, 897–910.
Ono, D., Honma, K., and Honma, S. (2021). Roles of neuropeptides, VIP and AVP, in the mammalian central circadian clock. Front Neurosci 15, 650154.
Ono, D., Mukai, Y., Hung, C.J., Chowdhury, S., Sugiyama, T., and Yamanaka, A. (2020). The mammalian circadian pacemaker regulates wakefulness via CRF neurons in the paraventricular nucleus of the hypothalamus. Sci Adv 6, eabd0384.
Paul, S., Hanna, L., Harding, C., Hayter, E.A., Walmsley, L., Bechtold, D.A., and Brown, T.M. (2020). Output from VIP cells of the mammalian central clock regulates daily physiological rhythms. Nat Commun 11, 1453.
Reppert, S.M., and Weaver, D.R. (2002). Coordination of circadian timing in mammals. Nature 418, 935–941.
Scammell, T.E., Arrigoni, E., and Lipton, J.O. (2017). Neural circuitry of wakefulness and sleep. Neuron 93, 747–765.
Silver, R., Lesauter, J., Tresco, P.A., and Lehman, M.N. (1996). A diffusible coupling signal from the transplanted suprachiasmatic nucleus controlling circadian locomotor rhythms. Nature 382, 810–813.
Stringer, C., Wang, T., Michaelos, M., and Pachitariu, M. (2021). Cellpose: a generalist algorithm for cellular segmentation. Nat Methods 18, 100–106.
Sun, Y.C., Chen, X., Fischer, S., Lu, S., Zhan, H., Gillis, J., and Zador, A.M. (2021). Integrating barcoded neuroanatomy with spatial transcriptional profiling enables identification of gene correlates of projections. Nat Neurosci 24, 873–885.
Tasic, B., Yao, Z., Graybuck, L.T., Smith, K.A., Nguyen, T.N., Bertagnolli, D., Goldy, J., Garren, E., Economo, M.N., Viswanathan, S., et al. (2018). Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78.
Todd, W.D., Fenselau, H., Wang, J.L., Zhang, R., Machado, N.L., Venner, A., Broadhurst, R.Y., Kaur, S., Lynagh, T., Olson, D.P., et al. (2018). A hypothalamic circuit for the circadian control of aggression. Nat Neurosci 21, 717–724.
Todd, W.D., Venner, A., Anaclet, C., Broadhurst, R.Y., De Luca, R., Bandaru, S.S., Issokson, L., Hablitz, L.M., Cravetchi, O., Arrigoni, E., et al. (2020). Suprachiasmatic VIP neurons are required for normal circadian rhythmicity and comprised of molecularly distinct subpopulations. Nat Commun 11, 4410.
Vadnie, C.A., and McClung, C.A. (2017). Circadian rhythm disturbances in mood disorders: insights into the role of the suprachiasmatic nucleus. Neural Plast 2017, 1–28.
Varela, L., and Horvath, T.L. (2019). Parallel paths in PVH control of feeding. Neuron 102, 514–516.
Welsh, D.K., Takahashi, J.S., and Kay, S.A. (2010). Suprachiasmatic nucleus: cell autonomy and network properties. Annu Rev Physiol 72, 551–577.
Welsh, D.K., Yoo, S.H., Liu, A.C., Takahashi, J.S., and Kay, S.A. (2004). Bioluminescence imaging of individual fibroblasts reveals persistent, independently phased circadian rhythms of clock gene expression. Curr Biol 14, 2289–2295.
Wen, S., Ma, D., Zhao, M., Xie, L., Wu, Q., Gou, L., Zhu, C., Fan, Y., Wang, H., and Yan, J. (2020). Spatiotemporal single-cell analysis of gene expression in the mouse suprachiasmatic nucleus. Nat Neurosci 23, 456–467.
Xu, P., Berto, S., Kulkarni, A., Jeong, B., Joseph, C., Cox, K.H., Greenberg, M.E., Kim, T.K., Konopka, G., and Takahashi, J.S. (2021). NPAS4 regulates the transcriptional response of the suprachiasmatic nucleus to light and circadian behavior. Neuron 109, 3268–3282.e6.
Xu, X., Holmes, T.C., Luo, M.H., Beier, K.T., Horwitz, G.D., Zhao, F., Zeng, W., Hui, M., Semler, B.L., and Sandri-Goldin, R.M. (2020). Viral vectors for neural circuit mapping and recent advances in trans-synaptic anterograde tracers. Neuron 107, 1029–1047.
Yamaguchi, S., Isejima, H., Matsuo, T., Okura, R., Yagita, K., Kobayashi, M., and Okamura, H. (2003). Synchronization of cellular clocks in the suprachiasmatic nucleus. Science 302, 1408–1412.
Yoo, S.H., Yamazaki, S., Lowrey, P.L., Shimomura, K., Ko, C.H., Buhr, E.D., Siepka, S. M., Hong, H.K., Oh, W.J., Yoo, O.J., et al. (2004). PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc Natl Acad Sci USA 101, 5339–5346.
Yu, Y., Zeng, Z., Xie, D., Chen, R., Sha, Y., Huang, S., Cai, W., Chen, W., Li, W., Ke, R., et al. (2021). Interneuron origin and molecular diversity in the human fetal brain. Nat Neurosci 24, 1745–1756.
Acknowledgement
This work was supported by the National Natural Science Foundation of China (32171157, 31971090), Ministry of Science and Technology of the People’s Republic of China (2021ZD0203400) and Kuanren Talents’ Project of The Second Affiliated Hospital of Chongqing Medical University. We would like to thank Dr. David O’Keefe for careful reading and English language editing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The author(s) declare that they have no conflict of Interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Liao, M., Gao, X., Chen, C. et al. Integrated neural tracing and in-situ barcoded sequencing reveals the logic of SCN efferent circuits in regulating circadian behaviors. Sci. China Life Sci. 67, 518–528 (2024). https://doi.org/10.1007/s11427-023-2420-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-023-2420-7