Abstract
Microplastics (MPs) are ubiquitous pollutants persisting almost everywhere in the environment. With the increase in anthropogenic activities, MP accumulation is increasing enormously in aquatic, marine, and terrestrial ecosystems. Owing to the slow degradation of plastics, MPs show an increased biomagnification probability of persistent, bioaccumulative, and toxic substances thereby creating a threat to environmental biota. Thus, remediation of MP-pollutants requires efficient strategies to circumvent the mobilization of contaminants leaching into the water, soil, and ultimately to human beings. Over the years, several microorganisms have been characterized by the potential to degrade different plastic polymers through enzymatic actions. Metagenomics (MGs) is an effective way to discover novel microbial communities and access their functional genetics for the exploration and characterization of plastic-degrading microbial consortia and enzymes. MGs in combination with metatranscriptomics and metabolomics approaches are a powerful tool to identify and select remediation-efficient microbes in situ. Advancement in bioinformatics and sequencing tools allows rapid screening, mining, and prediction of genes that are capable of polymer degradation. This review comprehensively summarizes the growing threat of microplastics around the world and highlights the role of MGs and computational biology in building effective response strategies for MP remediation.
Similar content being viewed by others
Introduction
The use of plastic has escalated tremendously over the last 50 years due to industrialization. Plastic rise from 1.5 million metric tons (MMTs) in 1950 to 367 MMTs in 2020 is a testament to the global plastic surge (Peng et al. 2021). Even though there has been a decrease of 0.3% in plastic production, the shoot-up in the usage of masks, gloves, sanitizer bottles, and medical equipment during the ongoing COVID-19 pandemic has contributed to unforeseen environmental crisis (Patrício Silva et al. 2021). Microplastics (MPs), the plastic fragments with less than 5 mm in size, are insoluble, biodegradable, and non-biodegradable waste particles having a half-life of about 100–1000 years. Based on the occurrence, MPs are classified into primary and secondary types. Primary MPs exist in nature in standard MP-size such as microbeads and plastic pellets, whereas secondary MPs arise from the breakdown of larger plastic materials like fishing nets, soda bottles, microwave containers, and other plastic products. Chemically, MPs are synthetic or semi-synthetic polymers composed of carbon, nitrogen, oxygen, hydrogen, chloride, silicon, etc. Depending on the nature of side chains, polymer backbone, physical properties, tensile strength, density, and thermal resistant plastics are classified into seven types each numbered according to their recycling codes as (1) polyethylene terephthalate/PET (beverage bottles, polyester clothing, rope), (2) high-density polyethylene/HDPE (detergent bottles, toys, buckets, rigid pipes), (3) polyvinyl chloride/PVC (credit cards, medical tubing, rain gutters), (4) low-density polyethylene/LDPE (grocery bags, beverage cups, bread bags), (5) polypropylene/PP (straws, packaging tape, disposable diapers), (6) polystyrene/styrofoam/PS (insulations, takeout food containers, cutlery), and (7) others/O (bisphenol A, polyamimide, polycarbonate) (Verla et al. 2019; Henderson and Green 2020; Veerasingam et al. 2020; Frias et al. 2021).
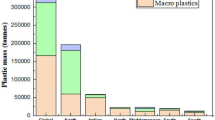
The top countries in the generation plastic waste per year in million tons in 2020 include the USA (58.02) (Law et al. 2020), India (55.06) (Shams et al. 2021), the UK (39.7) (Burgess et al. 2021), South Korea (38.1) (Shin et al. 2020), Germany (36) (Nelles et al. 2016), Thailand (32.4) (Parashar and Hait 2021), Malaysia (29.8) (Fauziah et al. 2021), Argentina (29.7) (Ronda et al. 2021), Russia (28) (Filiciotto and Rothenberg 2021), Italy (24.5) (Geyer et al. 2017), and Brazil (23.2) (Almeida et al. 2021). Most ecosystems are under threat of plastic pollution because of the properties like non-biodegradability, limited recovery, toxicity, higher ingestion, accumulation, and incorporation associated with MPs (Campanale et al. 2020; Issac and Kandasubramanian 2021). Since MP particles bear a resemblance with the food of marine biota, fishes, mammals, and plankton easily engulf it and accumulate in the body leading to blockage of the digestive system (Walkinshaw et al. 2020). Wang et al. (2019a) studied the effect of ingested PS-MPs on Artemia parthenogenetica (microcrustacean) and reported the occurrence of several abnormal epithelial cells in the digestive tract. Exposure of zooplankton crustacean Daphnia magna to PET textile microfibers resulted in increased mortality of daphnids (Jemec et al. 2016). MPs not only affect the ecosystem directly, but they also act as carriers for other environmental contaminants like heavy metals such as zinc and copper (Brennecke et al. 2016), polychlorinated biphenyl (Gerdes et al. 2019), and polyaromatic hydrocarbons (Sørensen et al. 2020) (Ye et al. 2020a). Humans may suffer chronic effects by ingestion, inhalation, and dermal contact of MPs leading to cell damage, inflammation, and hypersensitive reactions (Visalli et al. 2021; Domenech and Marcos 2021; Blackburn and Green 2021). A 2016–2017 UN report documented about 800 animal species contaminated with plastic via entanglement and ingestion, which is almost 70% greater than that of 1977 UN report. This makes humans prone to harmful effects of plastic in the upcoming decades (Smith et al. 2018).
Hwang et al. (2019) assessed the PP toxicity in human-derived cells and found that PP-MPs induce pro-inflammatory cytokines in a size-dependent manner. Likewise, Wu et al. (2019) studied the size-dependent effects of PS-MPs on cytotoxicity and efflux pump inhibition in human colon adenocarcinoma Caco-2 cells. They reported higher mitochondrial depolarization through 5 μm PS-MPs while 0.1 μm PS-MPs induced higher inhibition of adenosine triphosphate-binding cassette transporter. The traditional disposal methods like recycling, incineration, and landfill have been reported to show negative effects by generating secondary pollutants that cause disastrous effects on the environment (Rhodes 2018). Therefore, microbial degradation has emerged as a method of choice for expunging plastic and other pollutants. Several studies have been carried out in studying the biodegradation of MPs such as PE (Restrepo-Flórez et al. 2014), PS (Kim et al. 2021), PP (Jeon et al. 2021), and PET (Farzi et al. 2019). Kim et al. (2020) reported that the Pseudomonas aeruginosa DSM 50,071 strain, isolated from the gut of Zophobas atratus larvae, mediates the degradation of PS-MPs through enzymatic action. Zalerion maritimum (Paço et al. 2017), Aspergillus versicolor (Akhtar and Mannan 2020), Vibrio parahemolyticus (Kesy et al. 2020), and Psychrobacter sp. (Chattopadhyay 2022) have been also reported to exhibit the MP-remediation potential. A challenge in using microbial degradation on large scale is the slow rate of plastic degradation. Moreover, most of the reports published on the biodegradation of MPs have been performed in the laboratory set-ups.
Many microbes cannot be cultured in the laboratory conditions; hence, culture-based approaches have proved to be insufficient for the exploration and characterization of microorganisms. Besides, plastic biodegradation is also an outcome of the microbial consortia acting synergistically, which is difficult to study through culture-based approach. Metagenomics offers a gateway to overcome this problem (Handelsman 2004; Wani et al. 2022a). MGs in association with other meta-omics approaches are proving to be the standout approach for the identification of novel uncultivable microorganisms capable of MP-remediation (Bharagava et al. 2018; Wani et al. 2022b). This review offers a comprehensive outlook of the MP threat around the globe besides highlighting the fundamental MP remediation studies mediated by microorganisms isolated through culture-dependent and culture-independent approaches.
Microplastics (MPs): generation and escalation
Millions of tons of plastics are released into the environment every year. As a result, the quantity and distribution of MPs have escalated in the atmosphere, aquatic, and terrestrial ecosystems (Fig. 1). It is estimated that by 2060 plastic accumulation can reach up to 155–265 million tons, and it is believed that about 13.2% of this weight could be MPs (Eriksen et al. 2014; Geyer et al. 2017; Bergmann et al. 2019). The presence of MPs in different environments was revealed during the early 1970s (Buchanan 1971; Carpenter and Smith 1972). However, in recent times, scientists have started to investigate MP spread, accumulation, and ecological implications (Huang et al. 2021; Chen et al. 2021a; Vaid et al. 2021; Kallenbach et al. 2022). MP pollution in terrestrial and freshwater ecosystems has been less extensively studied in comparison to marine ecosystems (Chen et al. 2021a). Afrin et al. (2020) investigated MP presence in landfill sites of Dhaka, Bangladesh, and reported the presence of LDPE, HDPE, and cellulose acetate. Liu et al. (2018) also reported PP (50.51%) and PE (43.43%) in the suburbs of Shanghai, China. In total, 10% of the plastic ends up in the ocean and about 7–8 million plastic pieces escape into the oceans from land terrestrial sources. At present, most of the world’s seas and oceans are MP-contaminated. The Mediterranean Sea, with a 1500 m average depth, is recognized as a plastic contamination hotspot because its MP-concentration is fourfold greater than the North Pacific Ocean. This is attributed to the distinguishing semi-enclosed morphology of the Mediterranean Sea, and surrounding plastic waste-generating countries (Sharma et al. 2021). Table 1 gives insight about the growing MP contamination in different parts of the world. Lacerda et al. (2019) evaluated and characterized plastics in sea surface waters of the Antarctic Peninsula and did not find any statistical difference between the amount of MPs (54%) and mesoplastics (46%). They found smaller fragments composed of polyamide, PET, and polyurethane (PU).
Bioaccumulation and Ecotoxicological repercussions of MPs
The resistance (Sharma and Chatterjee 2017), high durability (Lim 2021), high consumption (Chen et al. 2021b), and low recycling (Muncke et al. 2020) of plastic polymers contribute to the escalation of plastic in the environment. Oceans are the largest known sinks for MPs (Kvale et al. 2020). The plastic debris from sewage treatment plants, transport and cosmetic industries, manufacturing, fishing, packaging, and shipping industries reaches the marine environment and is estimated to be 5–12 million metric tons per annum (Thushari and Senevirathna 2020; Vriend et al. 2021; Lim 2021; Peng et al. 2021). MP accumulation in terrestrial and aquatic biota through absorption, ingestion, or respiration has been widely recognized (Duis and Coors 2016; Souza Machado et al. 2018; Amobonye et al. 2021). Arenicola marina, an annelid species, has been reported to have MPs embedded in its gastrointestinal tracts (Besseling et al. 2013). Some crustaceans like Carcinus maenas have also been reported with the presence of MPs in the digestive and respiratory tracts (McGoran et al. 2020). These plastic particles are mistaken for food, leading to the blockade of essential body tracts which results in the generation of incorrect signals (Smith et al. 2018; Ugwu et al. 2021). Several studies have shown that MP accumulation or continuous exposure in aquatic organisms leads to the deterioration of inflammatory and oxidative intestinal balance, and permeability disruption of gut epithelial cells besides promoting the growth of pathogens on cell surfaces (Viršek et al. 2017; Limonta et al. 2019; Yang et al. 2020). Red tilapia when exposed to 0.3, 5, and 70 μm PS fragments for 14 days induced oxidative stress, neurotoxicity, and inhibition of cytochrome P450 enzyme activity (Ding et al. 2020). The accumulation of PS in Oryzias melastigmas (Ye et al. 2021) and PE in Dicentrachus labrax (Barboza et al. 2020) has been reported to cause negative effects on histology, immunity, and metabolism. Barboza et al. (2020) reported that PE and polyester in wild fish cause oxidative damage in muscles and gills besides increasing acetylcholinesterase activity in the brain. Bisphenol A and petroleum hydrocarbon aggravate immunotoxicity in blood clams and increase the toxicity of cadmium in fishes (Prüst et al. 2020). Benthic sea cucumbers, a non-selective bottom feeder, feed on the ocean floor debris and engulf a large amount of sediment (Sfriso et al. 2020). A study reported that Holothuria floridana, Thyonella gemmate, and Cucumaria frondose ingested 2–20 times more filter feeders have been reported to ingest MPs which decreases their filtration ability leading to effects like neurotoxicity and immunotoxicity (Mohsen et al. 2019; Bulleri et al. 2021). In 2019, marine biologists reported that seagrass beds in Makassar Strait, Indonesia, contain MP contaminants in the form of beads, pellets, fragments, and fibers (Tahir et al. 2019). Zooplankton also ingests MP beads which upon excretion can stick to the exoskeleton and appendages (Hasegawa and Nakaoka 2021).
The bioaccumulation of MPs in humans largely remains obscure, yet the MP consumption by crustaceans and fishes which are subsequently eaten by humans is still a matter of concern. There has been no study that evaluates the direct effect of plastic polymers on humans. A major concern in determining the negative effects of MPs on humans is the lack of information on human exposure. Thus, a better understanding of the MP-ability to cross epithelial barriers, skin, and gastrointestinal tract is needed to alleviate the uncertainty in human risk assessment of MPs (Prata et al. 2020; Vethaak and Legler 2021). However, several laboratory studies involving human cells and tissues and model organisms like rats and mice have shown negative implications of MPs. Researchers have started to investigate the presence of MPs in human tissues to extrapolate the effects of MPs that are directly human-oriented rather than in vitro. Ragusa et al. gave the first evidence of PPMP presence in the human placenta (Ragusa et al. 2021). Even though the presence and implications of MP in human tissues are obscure, there is a need to track and monitor MP pollution continuously. Exposure of mice to PE showed inflammation (Li et al. 2020) and smaller pups (Park et al. 2020), and exposure to PS reduced sperm count in mice (Jin et al. 2021). In mice gut, MPs increased intestinal permeability, altered gut microbiota composition, and enhanced intestinal inflammation (Deng et al. 2020a, b). One of the sub-chronic studies reported the accumulation of methacrylate polymer beads only in the gastrointestinal tract of mice (Groborz et al. 2020). Rodriguez-Seijo et al. (2017) reported the accumulation of PE-MPs in the earthworm gut causing damage to the epithelium of the gut wall. Seabirds also feed on marine debris and several studies have reported the presence of MPs in samples targeted for dietary studies, regurgitated cadavers, and feces. After engulfing, seabirds likely get rid of MPs through excretion or regurgitation (Blight and Burger 1997; Gil-Delgado et al. 2017; Hamilton et al. 2021). However, there is a risk of exposing offspring to the MPs at the time of feeding. Kühn and van Franeker (2012) found plastic in the intestine of juveniles rather than in adult birds.
Table 2 gives insight into the effect of different MPs on aquatic and terrestrial living systems of the earth. Figure 2 illustrates the potential threat of MPs on the biotic components of the earth.
Microplastic remediation mediated by microorganisms
MPs degrade mechanically (Schyns and Shaver 2021), chemically (Zhou et al. 2021), and biologically (Arpia et al. 2021) in the environment. Degradation rates mainly depend on structure, chemical composition, temperature, humidity, and deposition environment (Soil, water, sand). Mechanical degradation of MPs occurs through particle contact with anthropogenic (littered trash, boats, vehicles, groynes) and natural items (sediment, woody debris, shells) (Strayer and Findlay 2010; Qiao et al. 2019). Mechanical abrasion of MPs produces items that are similar in morphology to sediment grains. Song et al. (2017) examined the effect of UV exposure on MPs in the replicated beach environment and reported that the degradation rate varies with plastic type. PE and PP showed low degradation possibility through mechanical abrasion, but PS was found to fragment into more pieces. The exposure of floating plastic to UV light leads to the polymer degradation and the generation of chain scission products (Gewert et al. 2018). Enfrin et al. (2020) investigated weathering of MPs when exposed to stress using pumping, ultrasonic irradiation, and stirring. They reported that MP break down into nanoplastics (NPs) under low stress thus introducing more plastic debris to the environment. The weathering process of MPs is initiated or sometimes enhanced by chemical degradation through thermal oxidation, hydrolysis, and photooxidation. Plastics upon degradation produce different hydrocarbon gases such as methane, ethane, propylene, and ethylene when exposed to the solar radiation. Thus, climate-relevant trace gases are expected to increase with the accumulation of more plastic in the environment (Royer et al. 2018).
Besides, chemical degradation in seawater or replicated seawater has been reported to advance at a higher rate as compared to freshwater because of the variations in pH, and biotic community (Weinstein et al. 2016; Da Costa et al. 2018). Multiple chemical processes that are involved in the chemical degradation of MPs have been extensively reported and reviewed in great detail by different authors (Min et al. 2020; Ye et al. 2020b; Miao et al. 2020; Venkataramana et al. 2021; Zhou et al. 2021; Akhtar et al. 2022). Both natural and synthetic plastics are degraded by microbial action (Zeenat et al. 2021). Microorganisms degrade MPs using oxygen as an electron acceptor in the case of aerobic biodegradation (Yoshikawa et al. 2016). MPs are not transported directly into microorganisms because of their large size and water-insoluble nature (Cavicchioli et al. 2019). The degradation of MPs occurs through a series of events including, microbial attachment forming biofilms (Oberbeckmann and Labrenz 2020), and utilization of MPs as a carbon source (Lear et al. 2021). The microbial attachment to the MPs leads to the secretion of enzymes changing large MPs into monomers and oligomers having a low molecular weight (Lin et al. 2022). The microorganisms can change the surface properties of MPs followed by their bio-fragmentation through enzymatic action (Pathak and Navneet 2017). Hou and Majumder (2021) identified cytochrome 4500 s, monooxygenases, and hydrolases from microbial sources with PS-degrading potential. Several other microorganisms have been reported to have MP-degradation potential with varying biodegradation efficiency. Pseudomonas fluorescens, Bacillus sp., and Paenibacillus sp. degrade PE (Kathiresan 2003; Park and Kim 2019), B. vallismortis, Aspergillus oryzae, B. cereus, Trichoderma viride, A. nomius, and B. siamensis degrade LDPE (Skariyachan et al. 2017; Montazer et al. 2018; Nourollahi et al. 2019), and Klebsiella pneumoniae, and A. flavus degrade HDPE (Awasthi et al. 2017; Taghavi et al. 2021). The bio-fragmented MPs enter microorganisms through the cell membrane. The large monomers stay outside the microbial cells whereas small monomers move inside. Within the microbial cells, the monomers undergo oxidation which leads to energy generation used for biomass production (Lucas et al. 2008; Ru et al. 2020). MP biodegradability is largely affected by the factors like structural complexity, functional groups, morphology, polymer toughness, and bond strength (Klein et al. 2018). Biodegradability of MPs can be enhanced by combining MPs with several additives like nitric acid or pre-treating MPs with heat or UV (Montazer et al. 2018; Falkenstein et al. 2020). B. amyloliquefaciens degrades LDPE upon preliminary heat treatment by depolymerization reaction (Das and Kumar 2015). Similarly, B. safensis and B. mycoides degrades LDPE and HDPE upon pretreatment with 0.1% mercuric acid and sunlight respectively (Ibiene et al. 2013; Das and Kumar 2015). Microorganisms are known to adapt to varying environmental conditions including the pollution sites through a cascade of cellular and genetic pathways (Wani et al. 2022c). Microorganisms colonize surface of MPs which causes changes in mechanical properties like roughness, strength, and reduction in molecular weight (McGivney et al. 2020). The attachment changes hydrophobic MP surfaces into hydrophilic which makes them prone to degradation through the action of enzymes like tyrosinase, laccase, lipase, and peroxidase. For example, K. pneumoniae releases certain surfactants that mediate hydrophobic and hydrophilic phase exchange assisting in easy microbial penetration into PE for its degradation (Awasthi et al. 2017). Table 3 highlights the MP-degrading potential of microorganisms.
Enzymatic degradation of MPs
Owing to the presence of the homoatomic and heteroatomic backbone in plastics, MP degradation by microorganisms is an arduous process (Edmondson and Gilbert 2017). There is considerable weight loss in the plastic polymer with the action microorganisms but the process is significantly slower than chemically mediated biodegradation processes (Jaiswal et al. 2020). The polymer chains of MPs are broken by enzymes secreted by microbes (Mohanan et al. 2020; Lv et al. 2022; Kaur et al. 2022; Gaur et al. 2022). ATP-binding cassette transporters couple hydrolysis process to mediate the uptake and efflux of small fragments across the cell membrane in prokaryotic and eukaryotic cells. These transporters also play role in the secretion of toxins (Giuliani et al. 2011). Enzymatic actions like oxidation, hydrolysis, and hydroxylation cleave the MPs into monomers (Rana et al. 2022). The high molecular weight MPs are degraded first by extracellular enzymes and then incorporate into microbial cells (Urbanek et al. 2018). Within the microorganisms, the degraded MPs are catabolically channeled to yield energy for intracellular polymerization and integration into cellular structures (Müller et al. 2019; Rogers et al. 2020). Cutinase, an esterase sub-class, isolated from F. solani, Thermobifida fusca, T. alba, and T. cellulosilytica is effective in hydrolyzing polyester MPs (Ribitsch et al. 2012; Dong et al. 2020). Several studies have reported that PET degradation is mediated by PET hydrolases belonging to cutinases (Kawai et al. 2019; Furukawa et al. 2019; Carr et al. 2020). The enzymatic degradation of PET occurs either by surface modification of polyester fibers or polymer hydrolysis (Bååth et al. 2020). Several hydrolases have been reported to cause PET surface hydrophilization, such as lipases from Thermomyces sp., Candida antartica (Carniel et al. 2017), cutinases from Penicillium citrinum, Humicola insolens, and Saccharomonospora viridis (Liebminger et al. 2007), and carboxylesterases from T. halotolerans (Samak et al. 2020). PU degradation by membrane-associated (PudA) and extracellular (PueA, PueB) esterases isolated from Comamonas acidovorans, P.fluorescens, and P. chlororaphis has been characterized (Stern and Howard 2000). The blending of certain natural polymers like starch with synthetic MPs has been shown to increase the rate of MP-biodegradation (Vroman and Tighzert 2009). This is attributed to the rapid hydrolysis of starch making the MPs susceptible to microbial degradations. Karimi and Biria (2019) have reported LDPE degradation by the action of amylase when blended with starch. Currently, the least information on the enzymes acting on MPs with high molecular weight like PVC, PP, PS and polyamide is available. Even though mixed microbial communities have been reported to cause the weight loss of these MPs, the effectiveness of gene products is yet to be ascertained completely. Extreme environments are rich reservoirs of hydrolytic enzymes stable at fluctuating environmental conditions like temperature, pH, salinity, and pressure. The search for MP-degrading microorganisms and enzymes is already gaining research attention through metagenomic strategies. Table 4 gives an overview of the enzymes isolated and characterized from microbial sources with MP-degrading potential.
Metagenomics (MGs): gateway to microbial and enzyme mining
Even though microorganisms are present everywhere in the environment, limitations in traditional culture techniques have crippled the exploration of vast microbial flora (Lewis et al. 2021). Microbiologists estimate that only 1–2% of the total microbial flora is culturable, which leaves the majority of the microorganisms unexplored. MGs offer an efficient lens to reveal the hidden microbial diversity in a culture-independent manner (Handelsman 2004; Wani et al. 2022d). Figure 3 highlights the fundamental methodology of the sequence- and function-based metagenomic approach for the exploration of microorganisms and gene products. The taxonomic profiling and functional gene annotation of microbial communities of river Ganga (sediment) using whole-genome MGs have also been done (Rout et al. 2022). Several other research groups have identified novel bacteria from different sites including extreme environments like hot springs, deserts, and deep-sea sediments for bioprospecting using a MG approach (Tang et al. 2018; Najar et al. 2020; Alotaibi et al. 2020; Zhu et al. 2022; Wani et al. 2022b). Global ocean sampling revealed about 40 million non-redundant novel genes from more than 30,000 species, whereas over 97% of the 150 million genes reported in topsoil globally cannot be found in the existing gene catalogue. This is a strong indicator that microbiomes carry huge functional potential, with unculturable microorganisms as acting enzyme reservoir (Sunagawa et al. 2015; Bahram et al. 2018). In a study, hidden Markov models were constructed from experimentally verified enzymes and mined soil and ocean metagenomes to assess the ability of microorganisms in degrading plastics. They compiled almost 30,000 non-redundant enzymes that were homologues with known enzymes having plastic degrading potential (Zrimec et al. 2021). Chow et al. (2023) present a sequence-based in silico strategy for screening and characterization of PETases from MG datasets. The MG screening of a novel PET esterase through in vitro expression system has also been developed using next-generation sequencing (Han et al. 2023). In a recent study, distinct microbial communities have been unveiled through MGs that degrade hydrocarbon chains, which are units of plastic polymers (Hauptfeld et al. 2022). Using 16S rRNA datasets obtained through MGs, the taxonomic and functional characteristics of PE-degrading microorganisms have been analyzed from one of the waste recycling sites in Tehran, Iran (Hesami Zokaei et al. 2021).
Metagenomic (MG)-driven search operation for MP-degrading microorganisms through function and sequence-based metagenomic approaches. The function-based approach is followed by random screening for different enzymes while the sequence-based approach ensures the prediction of several genes that are effective in producing MP-degrading enzymes
Integrated microbial genome (IMG) helps to identify candidate genes from different metagenomes (Zaidi et al. 2021). In a MG study, two heat-stable enzymes with application in plastic degradation were partially characterized (Danso et al. 2018). Shotgun MGs have revealed the microbial community response to plastic contamination in coastal environments (Pinnell and Turner 2019). Shotgun MGs generated 3,314,688 contigs (DNA sequences that overlap providing a contiguous representation of a genomic region) and 120 microbial genomes. This was followed by the functional gene annotation to identify microbiomes that harbor genes encoding esterases, lipases, and monooxygenases that are known to degrade different types of plastics (Radwan et al. 2020). Hu et al. (2021) reported hydrolysis of PET by a metalloprotease and a serine protease. The study provided intrinsic insight into PET degradation and opened a gateway for hunting more plastic-degrading enzymes. Bollinger et al. (2020), also characterized a novel polyester hydrolase from P. aestusingri for the degradation of synthetic PET. Table 5 highlights some of the abundant microbes and enzymes isolated and characterized from microorganisms through culture-based and sequence- and function-based MG approaches having MP-degrading potential. Even though the MP degradation by microorganisms and their gene products is effective, the rate of degradation has always been a matter of concern. MG investigation allows upscaling the degradation rate by modifying the microbial composition and genome engineering.
Microbial manipulation
The manipulation of human, animal, soil, plant, and water microbiome is the contemporary strategy followed for increasing the benefits offered by them (Huynh et al. 2016; Hussain et al. 2018; Jochum et al. 2019). It includes several cellular, molecular, and chemical methods for extensive manipulation with higher specificity and magnitude. The prebiotic (chemical) approach enables modification in microbial communities to increase their adaptability and functionality in a particular environment (Gianoulis et al. 2009; Raes et al. 2021). Polysaccharides and oligosaccharides affect microbiome composition and support the growth of MP-degrading microorganisms (Grondin et al. 2017). Chitin, starch, lipopeptides, glycolipids, etc. help in biofilm formation by acting as surfactants on MP-surfaces (Shilpa et al. 2022). Similarly, probiotic cultures are applied for the better performance of MP-degraders through bioaugmentation (Kamilya and Devi 2022). The microorganisms like Pseudomonas, Micrococcus, Moraxella, Streptomyces, Thermoactinomyces, Penicillium, and Aspergillus are preferred over the native microorganism (Spini et al. 2018). Microbiome transplantation and probiotic bioaugmentation remain unsuccessful owing to the slow microbial growth, low cell viability, limited distribution, and reduced functionality. These issues are likely to be solved by metagenome engineering followed by bioaugmentation.
Microorganisms are genetically modified to produce novel strains that express unique and well-defined genetic determinants or to introduce genetic variants that cause phenotypic changes. The process is used to investigate the biotechnological potential linked to environmentally useful microorganisms and to make use of functional genes when put into the right host (Zeaiter et al. 2018). There have also been attempts to chemically alter marine microbes. Besides natural competence, wild-type and DNase-negative Vibrio cholerae strains are effectively electroporated and transformed by the researchers for biotechnological applications (Marcus et al. 1990; Jaskólska et al. 2018). Although the outcome of the electroporation can also be influenced by other parameters, including growth conditions, the pulse used, and the type of exogenous DNA, the electroporation efficiency is strain-dependent. Several marine strains from various genera, including Roseobacter, Vibrio, Pseudoalteromonas, Caulobacter, Cyanobacteria, and Halomona, have been successfully modified for the expression of environment-useful genes (Kivelä et al. 2008; Borg et al. 2016; Laurenceau et al. 2020).
Genetic engineering
With the progress in molecular biology and genetic engineering, the development of genetically modified microorganisms as potent MP degraders has advanced significantly. The construction of metagenomic libraries makes it likely to create genetic circuits with novel and precise functionalities (Bacha et al. 2021). The synthetic microbial cells created through genome editing, protein engineering, or genetic engineering can be employed for metagenome engineering in the plastisphere (Austin et al. 2018; Jaiswal et al. 2019). Since the biodegradation of MPs involves a cascade of oxidation processes which is difficult and slow by the action of single species (Klein et al. 2018). Metagenome engineering can be applied for complementing multiple genes involved in MP-degrading metabolic pathways. This will ensure the production of multiple enzymes that regulate biofilm formation and quorum sensing. Genome modification of B. subtilis and E. coli for the expression of PETase enzyme for the degradation of PET is a common example. PETase and MHETase have been identified in Ideonella sakaiensis 201-F6 and cloned in a suitable PUCIDT vector for the creation of recombinants with higher PET-degrading potential (Janatunaim and Fibriani 2020). Puspitasari et al. (2021) showed that the rate of PETase hydrolysis increases significantly in the presence of hydrophobin. Since the core metagenome of any site is constant, therefore, rather than modifying a single genome, it is possible to engineer the entire metagenome. The direct in situ metagenome engineering of microbial population is achievable through horizontal gene transfer of plasmid construct through genetic augmentation. The applicability of bacteriophages as gene-delivery agents is advancing. The strategy can very well be applied to the gene delivery with having MP-degrading potential. However, there is a growing concern about the release of genetically engineered microorganisms into the environment owing to their adverse effects. There are chances that engineered microorganisms may affect the biodiversity by creating more infectious pathogens, harm non-target species, and disrupt ecological balance (Lenski 1993; Clark 2006).
Metagenome analysis through computational tools
The development in computational tools and advancement of computational power has enormously aided in metagenome refinement and analysis. The sequencing of metagenome samples with the potential to degrade contaminants is a method of choice for identifying novel microorganisms and predicting genes. Shotgun MGs give insight into the microbial community members and the possible metabolic pathways mediated by them. Since metagenome collection from environments is largely uncontrolled, the organisms present in abundance are highly represented in sequence data. To achieve equal coverage of all the microbial members, the random shotgun sequencing resolves genomes uniformly and ensures the identification of lesser-presented organisms. The metagenome data is often enormous containing fragmented and raw data (Wooley et al. 2010). The metagenome sequencing of cow rumen generated more than 250 gigabases, while the gut microbiome of human-generated more than 550 gigabases of sequence data (Qin et al. 2010; Hess et al. 2011). Thus, the identification, collection, and curation of useful data from huge metagenome datasets are challenging for many researchers. Almeida et al. employed in silico screening method for the identification of potential PETase-like enzymes. They identified the PETase-like gene SM14est in streptomyces after analyzing more than 50 genomes (Almeida et al. 2019). Figure 4 represents the basic methodology of metagenome data analysis useful for understanding microbial diversity and predicting useful genes. One of the standalone metagenome analyzing tools is meta genome analyzer (MEGAN). It was initially used for studying metagenomes obtained from mammoth bone (Poinar et al. 2006). The tool is used to perform functional and taxonomic binning using the lowest common ancestor algorithm. More efficient, accurate, and faster computational tools are being developed to keep up the face with high-throughput sequencing. Metagenomic rapid annotations using subsystems technology (MG-RAST) is one of the biggest metagenome repositories developed for automatic phylogenic and functional analysis of metagenomes. Wani et al. (2022e) have comprehensively reviewed the maximum number of computational tools used in the analysis of metagenome data sets.
Basic methodology of the metagenome analysis through computational tools. The generated metagenome sequences are prefiltered for the removal of low-quality and redundant sequences using Eu-detect and DeConseq. To increase the analytical efficiency of computational tools, the metagenome assemblies are developed using Phrap or Celera or MEGAHIT assembler. This is followed by the prediction of genes using the MEGAN or Gene Mark or Gene Locator and Interpolated Markov Modeler (GLIMMER) program. Function-based annotation and taxonomic profiling are carried out MetaPhlan or Automatic phylogenomic inference application (AMPHORA) or Metaphyler followed by integration into MG-RAST, Integrated Microbial Genomes and Metagenomes (IMG/M) and Genomic Encyclopedia of Bacteria and Archaea (GEBA) like tools
Limitations and way forward
MG-based studies allow the exploration of microbial diversity, genetic evolution, species composition, and bioprospecting. However, bottlenecks in MGs right from sample collection until the analysis have always been challenging (Scholz et al. 2012). Sample collection is one of the confounding factors that affect the sequencing outcomes owing to concerns like contamination, transportation, storage, and safety. The developments in sequencing technology have significantly advanced computational tools for functional annotations and analysis (Bharti and Grimm 2021). However, multiple challenges still exist owing to the complexity of metagenomic data. While analyzing the complex metagenome data sets, challenges like multiple genomes and inter- and intra-genomic repeats lead to uneven sequencing with a higher degree of sequencing errors. Although the gene prediction tools have an efficiency of about 90%, the small number of genes escaping detection can be novel and more useful (Coleman and Korem 2021). Downstream processing of MG data is also much crucial for understanding microbiome structures and metabolic pathways, but due to multivariate metagenomic data, the downstream analysis is difficult (Lindgreen et al. 2016). The discovery of enzymes is prevented by other limitations like limited thermostability, low stereoselectivity, and insufficient expression. Ribosome engineering can be useful in retrieving all possible candidate genes for synthesis and testing the activities (Uchiyama and Miyazaki 2009). Fungi despite their affinity for plastics have been largely neglected. MG findings provide evidence that the plastisphere is a suitable niche for various fungal organisms, including pathogenic species (Gkoutselis et al. 2021).
The technical glitches and problems in data evaluation and interpretation confronted during metagenome studies can be overcome by the combination of MGs and machine learning tools like artificial intelligence (Rhoads 2020; Wani et al. 2022f). This will help in the accurate and timely characterization of microorganisms and microbial products useful in remediation processes. Artificial intelligence can be utilized in developing new models to design effective bioremediation tools and evaluate the performance and functionality of microorganisms. The development of smart biomarkers as indicators of pollution is an efficient way to track environmental fluctuations (Krishna Kumar et al. 2011). Moreover, gene engineering within genomes and metagenomes using gene-editing tools like clustered regular interspaced short palindromic repeats-associated protein (CRISPR-Cas) system can revolutionize the microbe-mediated degradation processes owing to its specific nature (Jaiswal et al. 2019; Wani et al. 2022g; Mir et al. 2022). This will help to upregulate contaminant-degrading genes and pave way for understanding the molecular pathway involved in it. The applicability of artificial intelligence environmental and genome editing for microbial simulation will continue to be the method of choice in combatting plastic and other pollution.
Conclusion
The emergence of MP-contamination has become a serious concern for the biota owing to the small size and their ability to reach into the human body through secondary sources like food. Moreover, research investigations and evidence based on the ecological toxicity of microplastics to aquatic biota revealed numerous toxic effects on organisms, posing serious ecological risks. The hazardous effect of microplastic is outlined as single and combined toxicity of various pollutants, which has reportedly impacted mortality rates, development, food intake capacity, reproductive capability, and gene expression in aquatic organisms. Considering the degradation potential of microbes and enzymes, it is possible to detoxify and degrade MPs into non-toxic end products. Thus, it is necessary to explore microorganisms that can mediate the bioremediation process of these MPs. MGs is a powerful genome-centric culture-independent technique to identify novel microorganisms and their products for bioprospecting including the degradation of environmental contaminants. MGs with other meta-omics strategies can be useful in building a timely response strategy for combatting the growing plastic threat and its associated concerns. Overall, MGs have enabled scientific studies of complex microbiomes, which have assisted to explain certain metabolic processes of polymer degradation. As a result, extensive research in this area is required, which may significantly reduce global plastic pollution while also ensuring the health of future generations.
Data availability
Not applicable.
References
Afrin S, Uddin MdK, Rahman MdM (2020) Microplastics contamination in the soil from urban landfill site, Dhaka, Bangladesh. Heliyon 6:e05572. https://doi.org/10.1016/j.heliyon.2020.e05572
Akhtar N, Mannan MA (2020) Mycoremediation: an unexplored gold mine. In: New and Future Developments in Microbial Biotechnology and Bioengineering, 1st edn. Elsevier, pp 11–24. https://doi.org/10.1016/B978-0-12-821007-9.00002-4
Akhtar N, Wani AK, Dhanjal DS, Mukherjee S (2022) Insights into the beneficial roles of dark septate endophytes in plants under challenging environment: resilience to biotic and abiotic stresses. World J Microbiol Biotechnol 38:79. https://doi.org/10.1007/s11274-022-03264-x
Alfaro-Núñez A, Astorga D, Cáceres-Farías L et al (2021) Microplastic pollution in seawater and marine organisms across the Tropical Eastern Pacific and Galápagos. Sci Rep 11:6424. https://doi.org/10.1038/s41598-021-85939-3
Almeida EL, Carrillo Rincón AF, Jackson SA, Dobson ADW (2019) In silico screening and heterologous expression of a polyethylene terephthalate hydrolase (PETase)-like enzyme (SM14est) with polycaprolactone (PCL)-degrading activity, from the marine sponge-derived strain Streptomyces sp. SM14. Front Microbiol 10:2187. https://doi.org/10.3389/fmicb.2019.02187
Alotaibi MO, Sonbol HS, Alwakeel SS et al (2020) Microbial diversity of some Sabkha and desert sites in Saudi Arabia. Saudi J Biol Sci 27:2778–2789. https://doi.org/10.1016/j.sjbs.2020.06.038
Alves VEN, Figueiredo GM (2019) Microplastic in the sediments of a highly eutrophic tropical estuary. Mar Pollut Bull 146:326–335. https://doi.org/10.1016/j.marpolbul.2019.06.042
Amobonye A, Bhagwat P, Raveendran S et al (2021) Environmental impacts of microplastics and nanoplastics: a current overview. Front Microbiol. https://doi.org/10.3389/fmicb.2021.768297
Antunes J, Frias J, Sobral P (2018) Microplastics on the Portuguese coast. Mar Pollut Bull 131:294–302. https://doi.org/10.1016/j.marpolbul.2018.04.025
Arpia AA, Chen W-H, Ubando AT et al (2021) Microplastic degradation as a sustainable concurrent approach for producing biofuel and obliterating hazardous environmental effects: a state-of-the-art review. J Hazard Mater 418:126381. https://doi.org/10.1016/j.jhazmat.2021.126381
Austin HP, Allen MD, Donohoe BS et al (2018) Characterization and engineering of a plastic-degrading aromatic polyesterase. Proc Natl Acad Sci 115:E4350–E4357. https://doi.org/10.1073/pnas.1718804115
Auta HS, Emenike CU, Jayanthi B, Fauziah SH (2018) Growth kinetics and biodeterioration of polypropylene microplastics by Bacillus sp. and Rhodococcus sp. isolated from mangrove sediment. Mar Pollut Bull 127:15–21. https://doi.org/10.1016/j.marpolbul.2017.11.036
Awasthi S, Srivastava P, Singh P et al (2017) Biodegradation of thermally treated high-density polyethylene (HDPE) by Klebsiella pneumoniae CH001. 3 Biotech 7:332. https://doi.org/10.1007/s13205-017-0959-3
Azizi A, Setyowati WN, Fairus S et al (2021) Microplastic pollution in the sediment of Jakarta Bay, Indonesia. IOP Conf Ser: Earth Environ Sci 930:012010. https://doi.org/10.1088/1755-1315/930/1/012010
Bååth JA, Borch K, Jensen K et al (2020) Comparative biochemistry of four polyester (PET) hydrolases. Chem Bio Chem 22:1627–1637. https://doi.org/10.1002/cbic.202000793
Bacha A-U-R, Nabi I, Zhang L (2021) Mechanisms and the engineering approaches for the degradation of microplastics. ACS EST Eng 1:1481–1501. https://doi.org/10.1021/acsestengg.1c00216
Bahram M, Hildebrand F, Forslund SK et al (2018) Structure and function of the global topsoil microbiome. Nature 560:233–237
Barboza LGA, Lopes C, Oliveira P et al (2020) Microplastics in wild fish from North East Atlantic Ocean and its potential for causing neurotoxic effects, lipid oxidative damage, and human health risks associated with ingestion exposure. Sci Total Environ 717:134625. https://doi.org/10.1016/j.scitotenv.2019.134625
Bardají DKR, Furlan JPR, Stehling EG (2019) Isolation of a polyethylene degrading Paenibacillus sp. from a landfill in Brazil. Arch Microbiol 201:699–704. https://doi.org/10.1007/s00203-019-01637-9
Bergmann M, Mützel S, Primpke S et al (2019) White and wonderful? Microplastics prevail in snow from the Alps to the Arctic. Sci Adv 5:eaax1157. https://doi.org/10.1126/sciadv.aax1157
Besseling E, Wegner A, Foekema EM et al (2013) Effects of microplastic on fitness and PCB bioaccumulation by the lugworm Arenicola marina (L.). Environ Sci Technol 47:593–600. https://doi.org/10.1021/es302763x
Bharagava RN, Purchase D, Saxena G, Mulla SI (2018) Applications of metagenomics in microbial bioremediation of pollutants: from genomics to environmental cleanup. From Genomics to Environmental Cleanup. In: Microbial Diversity in the Genomic Era, 1st edn. Elsevier, pp 459–477. https://doi.org/10.1016/B978-0-12-814849-5.00026-5
Bharti R, Grimm DG (2021) Current challenges and best-practice protocols for microbiome analysis. Brief Bioinform 22:178–193
Blackburn K, Green D (2021) The potential effects of microplastics on human health: what is known and what is unknown. Ambio. https://doi.org/10.1007/s13280-021-01589-9
Blight LK, Burger AE (1997) Occurrence of plastic particles in seabirds from the eastern North Pacific. Mar Pollut Bull 34:323–325. https://doi.org/10.1016/S0025-326X(96)00095-1
Bollinger A, Thies S, Knieps-Grünhagen E et al (2020) A Novel polyester hydrolase from the marine bacterium Pseudomonas aestusnigri – structural and functional insights. Front Microbiol 11:114. https://doi.org/10.3389/fmicb.2020.00114
Borg Y, Grigonyte AM, Boeing P et al (2016) Open source approaches to establishing Roseobacter clade bacteria as synthetic biology chassis for biogeoengineering. Peer J 4:e2031. https://doi.org/10.7717/peerj.2031
Brennecke D, Duarte B, Paiva F et al (2016) Microplastics as vector for heavy metal contamination from the marine environment. Estuar Coast Shelf Sci 178:189–195. https://doi.org/10.1016/j.ecss.2015.12.003
Bryant JA, Clemente TM, Viviani DA, et al (2016) Diversity and activity of communities inhabiting plastic debris in the North Pacific Gyre. mSystems 1:e00024–16. https://doi.org/10.1128/mSystems.00024-16
Buchanan JB (1971) Pollution by synthetic fibres. Mar Pollut Bull 2:23. https://doi.org/10.1016/0025-326X(71)90136-6
Bulleri F, Ravaglioli C, Anselmi S, Renzi M (2021) The sea cucumber Holothuria tubulosa does not reduce the size of microplastics but enhances their resuspension in the water column. Sci Total Environ 781:146650. https://doi.org/10.1016/j.scitotenv.2021.146650
Burgess M, Holmes H, Sharmina M, Shaver MP (2021) The future of UK plastics recycling: one bin to rule them all. Resour Conserv Recycl 164:105191. https://doi.org/10.1016/j.resconrec.2020.105191
Buss N, Sander B, Hua J (2021) Effects of polyester microplastic fiber contamination on amphibian-trematode interactions. Environ Toxicol Chem. https://doi.org/10.1002/etc.5035
Campanale C, Massarelli C, Savino I et al (2020) A detailed review study on potential effects of microplastics and additives of concern on human health. Int J Environ Res Public Health 17:1212. https://doi.org/10.3390/ijerph17041212
Carniel A, Valoni É, Nicomedes J et al (2017) Lipase from Candida antarctica (CALB) and cutinase from Humicola insolens act synergistically for PET hydrolysis to terephthalic acid. Process Biochem 59:84–90. https://doi.org/10.1016/j.procbio.2016.07.023
Carpenter EJ, Smith KL (1972) Plastics on the Sargasso sea surface. Science 175:1240–1241. https://doi.org/10.1126/science.175.4027.1240
Carr CM, Clarke DJ, Dobson ADW (2020) Microbial polyethylene terephthalate hydrolases: current and future perspectives. Front Microbiol 11:571265. https://doi.org/10.3389/fmicb.2020.571265
Cavicchioli R, Ripple WJ, Timmis KN et al (2019) Scientists’ warning to humanity: microorganisms and climate change. Nat Rev Microbiol 17:569–586. https://doi.org/10.1038/s41579-019-0222-5
Chattopadhyay I (2022) Role of microbiome and biofilm in environmental plastic degradation. Biocatal Agric Biotechnol 39:102263. https://doi.org/10.1016/j.bcab.2021.102263
Chen HL, Selvam SB, Ting KN, Gibbins CN (2021a) Microplastic pollution in freshwater systems in Southeast Asia: contamination levels, sources, and ecological impacts. Environ Sci Pollut Res 28:54222–54237. https://doi.org/10.1007/s11356-021-15826-x
Chen Y, Awasthi AK, Wei F et al (2021) Single-use plastics: production, usage, disposal, and adverse impacts. Sci Total Environ 752:141772. https://doi.org/10.1016/j.scitotenv.2020.141772
Chow J, Pérez-García P, Dierkes RF et al (2023) The PET-degrading potential of global metagenomes: from in silico mining to active enzymes. In: Streit WR, Daniel R (eds) Metagenomics: Methods and Protocols. Springer, US, New York, NY, pp 139–151
Clark EA (2006) Environmental risks of genetic engineering. Euphytica 148:47–60. https://doi.org/10.1007/s10681-006-5940-x
Coleman I, Korem T (2021) Embracing metagenomic complexity with a genome-free approach. Msystems 6:e00816-e821
Colzi I, Renna L, Bianchi E et al (2022) Impact of microplastics on growth, photosynthesis and essential elements in Cucurbita pepo L. J Hazard Mater 423:127238. https://doi.org/10.1016/j.jhazmat.2021.127238
Crump A, Mullens C, Bethell EJ et al (2020) Microplastics disrupt hermit crab shell selection. Biol Lett 16:20200030. https://doi.org/10.1098/rsbl.2020.0030
Da Costa JP, Nunes AR, Santos PSM et al (2018) Degradation of polyethylene microplastics in seawater: Insights into the environmental degradation of polymers. J Environ Sci Health A 53:866–875. https://doi.org/10.1080/10934529.2018.1455381
Danso D, Chow J, Streit WR (2019) plastics: environmental and biotechnological perspectives on microbial degradation. Appl Environ Microbiol 85:e01095-19. https://doi.org/10.1128/AEM.01095-19
Danso D, Schmeisser C, Chow J et al (2018) New insights into the function and global distribution of polyethylene terephthalate (PET)-degrading bacteria and enzymes in marine and terrestrial metagenomes. Appl Environ Microbiol 84:e02773-e2817. https://doi.org/10.1128/AEM.02773-17
Das MP, Kumar S (2015) An approach to low-density polyethylene biodegradation by Bacillus amyloliquefaciens. 3 Biotech 5:81–86. https://doi.org/10.1007/s13205-014-0205-1
de Almeida R, de Souza RG, Campos JC (2021) Lessons and challenges for the recycling sector of Brazil from the pandemic outbreak of COVID-19. Waste Dispos Sustain Energy 1–10. https://doi.org/10.1007/s42768-021-00075-y
de Faria É, Girard P, Nardes CS et al (2021) Microplastics pollution in the South American Pantanal. Case Stud Chem Environ Eng 3:100088. https://doi.org/10.1016/j.cscee.2021.100088
de Souza Machado AA, Kloas W, Zarfl C et al (2018) Microplastics as an emerging threat to terrestrial ecosystems. Glob Change Biol 24:1405–1416. https://doi.org/10.1111/gcb.14020
Deguchi T, Kitaoka Y, Kakezawa M, Nishida T (1998) Purification and characterization of a nylon-degrading enzyme. Appl Environ Microbiol 64:1366–1371
Deng Y, Yan Z, Shen R et al (2020) Microplastics release phthalate esters and cause aggravated adverse effects in the mouse gut. Environ Int 143:105916. https://doi.org/10.1016/j.envint.2020.105916
Deng Y, Yan Z, Shen R et al (2020) Microplastics release phthalate esters and cause aggravated adverse effects in the mouse gut. Environ Int 143:105916. https://doi.org/10.1016/j.envint.2020.105916
Ding J, Huang Y, Liu S et al (2020) Toxicological effects of nano- and micro-polystyrene plastics on red tilapia: are larger plastic particles more harmless? J Hazard Mater 396:122693. https://doi.org/10.1016/j.jhazmat.2020.122693
Domenech J, Marcos R (2021) Pathways of human exposure to microplastics, and estimation of the total burden. Curr Opin Food Sci 39:144–151. https://doi.org/10.1016/j.cofs.2021.01.004
Dong Q, Yuan S, Wu L et al (2020) Structure-guided engineering of a Thermobifida fusca cutinase for enhanced hydrolysis on natural polyester substrate. Bioresour Bioprocess 7:37. https://doi.org/10.1186/s40643-020-00324-8
Duis K, Coors A (2016) Microplastics in the aquatic and terrestrial environment: sources (with a specific focus on personal care products), fate and effects. Environ Sci Eur 28:2. https://doi.org/10.1186/s12302-015-0069-y
Edmondson S, Gilbert M (2017) Chapter 2 - the chemical nature of plastics polymerization. In: Gilbert M (ed) Brydson’s Plastics Materials (Eighth Edition). Butterworth-Heinemann, pp 19–37
Enfrin M, Lee J, Gibert Y et al (2020) Release of hazardous nanoplastic contaminants due to microplastics fragmentation under shear stress forces. J Hazard Mater 384:121393. https://doi.org/10.1016/j.jhazmat.2019.121393
Eriksen M, Lebreton LCM, Carson HS et al (2014) Plastic pollution in the world’s oceans: more than 5 trillion plastic pieces weighing over 250,000 tons afloat at sea. PLoS One 9:e111913. https://doi.org/10.1371/journal.pone.0111913
Esmaeili A, Pourbabaee AA, Alikhani HA et al (2013) Biodegradation of low-density polyethylene (LDPE) by mixed culture of Lysinibacillus xylanilyticus and Aspergillus niger in soil. PLoS One 8:e71720. https://doi.org/10.1371/journal.pone.0071720
Falahudin D, Cordova MR, Sun X et al (2020) The first occurrence, spatial distribution and characteristics of microplastic particles in sediments from Banten Bay, Indonesia. Sci Total Environ 705:135304. https://doi.org/10.1016/j.scitotenv.2019.135304
Falkenstein P, Gräsing D, Bielytskyi P et al (2020) UV Pretreatment impairs the enzymatic degradation of polyethylene terephthalate. Front Microbiol 11:689. https://doi.org/10.3389/fmicb.2020.00689
Farzi A, Dehnad A, Fotouhi AF (2019) Biodegradation of polyethylene terephthalate waste using Streptomyces species and kinetic modeling of the process. Biocatal Agric Biotechnol 17:25–31. https://doi.org/10.1016/j.bcab.2018.11.002
Fauziah SH, Rizman-Idid M, Cheah W et al (2021) Marine debris in Malaysia: a review on the pollution intensity and mitigating measures. Mar Pollut Bull 167:112258. https://doi.org/10.1016/j.marpolbul.2021.112258
Filiciotto L, Rothenberg G (2021) Biodegradable plastics: standards, policies, and impacts. Chemsuschem 14:56–72. https://doi.org/10.1002/cssc.202002044
Frias JP, Ivar do Sul JA, Panti C, Lima ARA (2021) Microplastics in the marine environment: sources, distribution, biological effects and socio-economic impacts. Front Environ Sci 90. https://doi.org/10.3389/fenvs.2021.676011
Fujisawa M, Hirai H, Nishida T (2001) Degradation of polyethylene and nylon-66 by the laccase-mediator system. J Polym Environ 9:103–108. https://doi.org/10.1023/A:1020472426516
Furukawa M, Kawakami N, Tomizawa A, Miyamoto K (2019) Efficient degradation of poly(ethylene terephthalate) with thermobifida fusca cutinase exhibiting improved catalytic activity generated using mutagenesis and additive-based approaches. Sci Rep 9:16038. https://doi.org/10.1038/s41598-019-52379-z
Ganesh Kumar A, Hinduja M, Sujitha K et al (2021) Biodegradation of polystyrene by deep-sea Bacillus paralicheniformis G1 and genome analysis. Sci Total Environ 774:145002. https://doi.org/10.1016/j.scitotenv.2021.145002
Gardon T, El Rakwe M, Paul-Pont I et al (2021) Microplastics contamination in pearl-farming lagoons of French Polynesia. J Hazard Mater 419:126396. https://doi.org/10.1016/j.jhazmat.2021.126396
Gaur VK, Gupta S, Sharma P et al (2022) Metabolic cascade for remediation of plastic waste: a case study on microplastic degradation. Curr Pollution Rep 8:30–50. https://doi.org/10.1007/s40726-021-00210-7
Gautam R, Bassi AS, Yanful EK (2007) Candida rugosa lipase-catalyzed polyurethane degradation in aqueous medium. Biotechnol Lett 29:1081–1086. https://doi.org/10.1007/s10529-007-9354-1
Gaytán I, Sánchez-Reyes A, Burelo M et al (2019) Degradation of recalcitrant polyurethane and xenobiotic additives by a selected landfill microbial community and its biodegradative potential revealed by proximity ligation-based metagenomic analysis. Front Microbiol 10:2986. https://doi.org/10.3389/fmicb.2019.02986
Gerdes Z, Ogonowski M, Nybom I et al (2019) Microplastic-mediated transport of PCBs? A depuration study with Daphnia magna. PLoS One 14:e0205378. https://doi.org/10.1371/journal.pone.0205378
Gewert B, Plassmann M, Sandblom O, MacLeod M (2018) Identification of chain scission products released to water by plastic exposed to ultraviolet light. Environ Sci Technol Lett 5:272–276
Geyer R, Jambeck JR, Law KL (2017) Production, use, and fate of all plastics ever made. Sci Adv 3:e1700782. https://doi.org/10.1126/sciadv.1700782
Ghatge S, Yang Y, Ahn J-H, Hur H-G (2020) Biodegradation of polyethylene: a brief review. Appl Biol Chem 63:27. https://doi.org/10.1186/s13765-020-00511-3
Giacomucci L, Raddadi N, Soccio M et al (2020) Biodegradation of polyvinyl chloride plastic films by enriched anaerobic marine consortia. Mar Environ Res 158:104949. https://doi.org/10.1016/j.marenvres.2020.104949
Gianoulis TA, Raes J, Patel PV et al (2009) Quantifying environmental adaptation of metabolic pathways in metagenomics. Proc Natl Acad Sci 106:1374–1379. https://doi.org/10.1073/pnas.0808022106
Gil-Delgado JA, Guijarro D, Gosálvez RU et al (2017) Presence of plastic particles in waterbirds faeces collected in Spanish lakes. Environ Pollut 220:732–736. https://doi.org/10.1016/j.envpol.2016.09.054
Giuliani SE, Frank AM, Corgliano DM et al (2011) Environment sensing and response mediated by ABC transporters. BMC Genomics 12:S8. https://doi.org/10.1186/1471-2164-12-S1-S8
Gkoutselis G, Rohrbach S, Harjes J et al (2021) Microplastics accumulate fungal pathogens in terrestrial ecosystems. Sci Rep 11:13214. https://doi.org/10.1038/s41598-021-92405-7
Groborz O, Poláková L, Kolouchová K et al (2020) Chelating Polymers for hereditary hemochromatosis treatment. Macromol Biosci 20:2000254. https://doi.org/10.1002/mabi.202000254
Grondin JM, Tamura K, Déjean G et al (2017) Polysaccharide Utilization Loci: Fueling Microbial Communities. J Bacteriol 199:e00860-e916. https://doi.org/10.1128/JB.00860-16
Habib S, Iruthayam A, Abd Shukor MY et al (2020) Biodeterioration of untreated polypropylene microplastic particles by antarctic bacteria. Polymers (basel) 12:E2616. https://doi.org/10.3390/polym12112616
Hajighasemi M, Tchigvintsev A, Nocek B et al (2018) Screening and characterization of novel polyesterases from environmental metagenomes with high hydrolytic activity against synthetic polyesters. Environ Sci Technol 52:12388–12401. https://doi.org/10.1021/acs.est.8b04252
Hamilton BM, Bourdages MPT, Geoffroy C et al (2021) Microplastics around an arctic seabird colony: particle community composition varies across environmental matrices. Sci Total Environ 773:145536. https://doi.org/10.1016/j.scitotenv.2021.145536
Han Y, Dierkes RF, Streit WR (2023) Metagenomic Screening of a Novel PET Esterase via in vitro expression system. In: Streit WR, Daniel R (eds) Metagenomics: Methods and Protocols. Springer, US, New York, NY, pp 167–179
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68:669 LP – 685. https://doi.org/10.1128/MMBR.68.4.669-685.2004
Hasegawa T, Nakaoka M (2021) Trophic transfer of microplastics from mysids to fish greatly exceeds direct ingestion from the water column. Environ Pollut 273:116468. https://doi.org/10.1016/j.envpol.2021.116468
Hauptfeld E, Pelkmans J, Huisman TT et al (2022) A metagenomic portrait of the microbial community responsible for two decades of bioremediation of poly-contaminated groundwater. Water Res 221:118767. https://doi.org/10.1016/j.watres.2022.118767
Henderson L, Green C (2020) Making sense of microplastics? Public understandings of plastic pollution. Mar Pollut Bull 152:110908. https://doi.org/10.1016/j.marpolbul.2020.110908
Hesami Zokaei F, Gharavi S, Asgarani E et al (2021) A comparative taxonomic profile of microbial polyethylene and hydrocarbon-degrading communities in diverse environments. Iran J Biotechnol 19:e2955. https://doi.org/10.30498/IJB.2021.2955
Hess M, Sczyrba A, Egan R et al (2011) Metagenomic Discovery of biomass-degrading genes and genomes from cow rumen. Science. https://doi.org/10.1126/science.1200387
Hou J, Lei Z, Cui L et al (2021) Polystyrene microplastics lead to pyroptosis and apoptosis of ovarian granulosa cells via NLRP3/Caspase-1 signaling pathway in rats. Ecotoxicol Environ Saf 212:112012. https://doi.org/10.1016/j.ecoenv.2021.112012
Hou L, Majumder EL-W (2021) Potential for and distribution of enzymatic biodegradation of polystyrene by environmental microorganisms. Materials 14:503. https://doi.org/10.3390/ma14030503
Howard GT, Blake RC (1998) Growth of Pseudomonas fluorescens on a polyester-polyurethane and the purification and characterization of a polyurethanase-protease enzyme. Int Biodeter and Biodegra 42:213–220. https://doi.org/10.1016/S0964-8305(98)00051-1
Hu Q, Jayasinghe-Arachchige VM, Prabhakar R (2021) Degradation of a main plastic pollutant polyethylene terephthalate by two distinct proteases (neprilysin and cutinase-like enzyme). J Chem Inf Model 61:764–776. https://doi.org/10.1021/acs.jcim.0c00797
Huang B, Sun L, Liu M et al (2021) Abundance and distribution characteristics of microplastic in plateau cultivated land of Yunnan Province, China. Environ Sci Pollut Res 28:1675–1688. https://doi.org/10.1007/s11356-020-10527-3
Huang J-N, Wen B, Zhu J-G et al (2020) Exposure to microplastics impairs digestive performance, stimulates immune response and induces microbiota dysbiosis in the gut of juvenile guppy (Poecilia reticulata). Sci Total Environ 733:138929. https://doi.org/10.1016/j.scitotenv.2020.138929
Hussain SS, Mehnaz S, Siddique KHM (2018) Harnessing the plant microbiome for improved abiotic stress tolerance. In: Egamberdieva D, Ahmad P (eds) Plant Microbiome: Stress Response. Springer, Singapore, pp 21–43
Huynh K, Schneider M, Gareau M (2016) Altering the gut microbiome for cognitive benefit? In: The Gut-Brain Axis Dietary, Probiotic, and Prebiotic Interventions on the Microbiota, Ist edn. Elsevier, pp 319–337. https://doi.org/10.1016/B978-0-12-802304-4.00015-3
Hwang J, Choi D, Han S et al (2019) An assessment of the toxicity of polypropylene microplastics in human derived cells. Sci Total Environ 684:657–669. https://doi.org/10.1016/j.scitotenv.2019.05.071
Ibiene AA, Stanley HO, Immanuel OM (2013) Biodegradation of polyethylene by Bacillus sp. Indigenous to the Niger Delta Mangrove Swamp. Nigerian J Biotechnol 26:68–78. https://doi.org/10.4314/njb.v26i1
Issac MN, Kandasubramanian B (2021) Effect of microplastics in water and aquatic systems. Environ Sci Pollut Res Int 28:19544–19562. https://doi.org/10.1007/s11356-021-13184-2
Jacquin J, Cheng J, Odobel C et al (2019) Microbial ecotoxicology of marine plastic debris: a review on colonization and biodegradation by the “plastisphere.” Front Microbiol 10:865. https://doi.org/10.3389/fmicb.2019.00865
Jaiswal S, Sharma B, Shukla P (2020) Integrated approaches in microbial degradation of plastics. Environ Technol Innov 17:100567. https://doi.org/10.1016/j.eti.2019.100567
Jaiswal S, Singh DK, Shukla P (2019) Gene editing and systems biology tools for pesticide bioremediation: a review. Front Microbiol. https://doi.org/10.3389/fmicb.2019.00087
Janatunaim RZ, Fibriani A (2020) Construction and cloning of plastic-degrading recombinant enzymes (MHETase). Recent Pat Biotechnol 14:229–234. https://doi.org/10.2174/1872208314666200311104541
Jankowska E, Gorman MR, Frischmann CJ (2022) Transforming the plastic production system presents opportunities to tackle the climate crisis. Sustainability 14:6539
Jaskólska M, Stutzmann S, Stoudmann C, Blokesch M (2018) QstR-dependent regulation of natural competence and type VI secretion in Vibrio cholerae. Nucleic Acids Res 46:10619–10634. https://doi.org/10.1093/nar/gky717
Jemec A, Horvat P, Kunej U et al (2016) Uptake and effects of microplastic textile fibers on freshwater crustacean Daphnia magna. Environ Pollut 219:201–209. https://doi.org/10.1016/j.envpol.2016.10.037
Jemec Kokalj A, Dolar A, Drobne D et al (2022) Environmental hazard of polypropylene microplastics from disposable medical masks: acute toxicity towards Daphnia magna and current knowledge on other polypropylene microplastics. Microplast Nanoplast 2:1. https://doi.org/10.1186/s43591-021-00020-0
Jeon HJ, Kim MN (2015) Functional analysis of alkane hydroxylase system derived from Pseudomonas aeruginosa E7 for low molecular weight polyethylene biodegradation. Int Biodeterior Biodegradation 103:141–146. https://doi.org/10.1016/j.ibiod.2015.04.024
Jeon J-M, Park S-J, Choi T-R et al (2021) Biodegradation of polyethylene and polypropylene by Lysinibacillus species JJY0216 isolated from soil grove. Polym Degrad Stab 191:109662. https://doi.org/10.1016/j.polymdegradstab.2021.109662
Ji J, Zhang Y, Liu Y et al (2020) Biodegradation of plastic monomer 2,6-dimethylphenol by Mycobacterium neoaurum B5–4. Environ Pollut 258:113793. https://doi.org/10.1016/j.envpol.2019.113793
Jiang S, Su T, Zhao J, Wang Z (2021) Isolation, identification, and characterization of polystyrene-degrading bacteria from the gut of Galleria mellonella (Lepidoptera: Pyralidae) larvae. Front Bioeng Biotechnol 9:736062. https://doi.org/10.3389/fbioe.2021.736062
Jimenez-Guri E, Roberts KE, García FC et al (2021) Transgenerational effects on development following microplastic exposure in Drosophila melanogaster. PeerJ 9:e11369. https://doi.org/10.7717/peerj.11369
Jin H, Ma T, Sha X et al (2021) Polystyrene microplastics induced male reproductive toxicity in mice. J Hazard Mater 401:123430. https://doi.org/10.1016/j.jhazmat.2020.123430
Jochum MD, McWilliams KL, Pierson EA, Jo Y-K (2019) Host-mediated microbiome engineering (HMME) of drought tolerance in the wheat rhizosphere. PLOS ONE 14:e0225933. https://doi.org/10.1371/journal.pone.0225933
Kallenbach EMF, Friberg N, Lusher A et al (2022) Anthropogenically impacted lake catchments in Denmark reveal low microplastic pollution. Environ Sci Pollut Res. https://doi.org/10.1007/s11356-022-19001-8
Kamilya D, Devi WM (2022) Bacillus probiotics and bioremediation: an aquaculture perspective. In: Islam MT, Rahman M, Pandey P (eds) Bacilli in Agrobiotechnology: Plant Stress Tolerance, Bioremediation, and Bioprospecting. Springer International Publishing, Cham, pp 335–347
Kang C-H, Oh K-H, Lee M-H et al (2011) A novel family VII esterase with industrial potential from compost metagenomic library. Microb Cell Fact 10:1–8
Karimi M, Biria D (2019) The promiscuous activity of alpha-amylase in biodegradation of low-density polyethylene in a polymer-starch blend. Sci Rep 9:2612. https://doi.org/10.1038/s41598-019-39366-0
Kathiresan K (2003) Polythene and Plastics-degrading microbes from the mangrove soil. Rev Biol Trop 51:629–633
Kaur P, Singh K, Singh B (2022) Microplastics in soil: impacts and microbial diversity and degradation. Pedosphere 32:49–60. https://doi.org/10.1016/S1002-0160(21)60060-7
Kawai F, Kawabata T, Oda M (2019) Current knowledge on enzymatic PET degradation and its possible application to waste stream management and other fields. Appl Microbiol Biotechnol 103:4253–4268. https://doi.org/10.1007/s00253-019-09717-y
Kesy K, Labrenz M, Scales BS et al (2020) Vibrio colonization is highly dynamic in early microplastic-associated biofilms as well as on field-collected microplastics. Microorganisms 9:E76. https://doi.org/10.3390/microorganisms9010076
Kim HR, Lee HM, Yu HC et al (2020) Biodegradation of Polystyrene by Pseudomonas sp. Isolated from the Gut of Superworms (Larvae of Zophobas atratus). Environ Sci Technol 54:6987–6996. https://doi.org/10.1021/acs.est.0c01495
Kim H-W, Jo JH, Kim Y-B et al (2021) Biodegradation of polystyrene by bacteria from the soil in common environments. J Hazard Mater 416:126239. https://doi.org/10.1016/j.jhazmat.2021.126239
Kirstein IV, Wichels A, Gullans E et al (2019) The plastisphere – uncovering tightly attached plastic “specific” microorganisms. PLoS One 14:e0215859. https://doi.org/10.1371/journal.pone.0215859
Kivelä HM, Madonna S, Krupovìč M et al (2008) Genetics for Pseudoalteromonas provides tools to manipulate marine bacterial virus PM2. J Bacteriol 190:1298–1307. https://doi.org/10.1128/JB.01639-07
Klein S, Dimzon IK, Eubeler J, Knepper TP (2018) Analysis, occurrence, and degradation of microplastics in the aqueous environment. In: Wagner M, Lambert S (eds) Freshwater Microplastics : Emerging Environmental Contaminants? Springer International Publishing, Cham, pp 51–67
Kleinschmidt JM, Janosik AM (2021) Microplastics in Florida, United States: a case study of quantification and characterization with intertidal snails. Front Ecol Evol 9:645727. https://doi.org/10.3389/fevo.2021.645727
Krishna Kumar PT, Vinod PT, Phoha VV et al (2011) Design of a smart biomarker for bioremediation: a machine learning approach. Comput Biol Med 41:357–360. https://doi.org/10.1016/j.compbiomed.2011.03.013
Kühn S, van Franeker JA (2012) Plastic ingestion by the northern fulmar (Fulmarus glacialis) in Iceland. Mar Pollut Bull 64:1252–1254. https://doi.org/10.1016/j.marpolbul.2012.02.027
Kumar AS, Varghese GK (2021) Microplastic pollution of Calicut beach - contributing factors and possible impacts. Mar Pollut Bull 169:112492. https://doi.org/10.1016/j.marpolbul.2021.112492
Kumar R, Pandit P, Kumar D et al (2021) Landfill microbiome harbour plastic degrading genes: a metagenomic study of solid waste dumping site of Gujarat. India. Sci Total Environ 779:146184. https://doi.org/10.1016/j.scitotenv.2021.146184
Kvale K, Prowe AEF, Chien C-T et al (2020) The global biological microplastic particle sink. Sci Rep 10:16670. https://doi.org/10.1038/s41598-020-72898-4
Lacerda AL, d F, Rodrigues L dos S, van Sebille E, et al (2019) Plastics in sea surface waters around the Antarctic Peninsula. Sci Rep 9:3977. https://doi.org/10.1038/s41598-019-40311-4
Laurenceau R, Bliem C, Osburne MS, et al (2020) Toward a genetic system in the marine cyanobacterium Prochlorococcus. Access Microbiol 2:acmi000107. https://doi.org/10.1099/acmi.0.000107
Law KL, Starr N, Siegler TR et al (2020) The United States’ contribution of plastic waste to land and ocean. Sci Adv 6:eabd0288. https://doi.org/10.1126/sciadv.abd0288
Lear G, Kingsbury JM, Franchini S et al (2021) Plastics and the microbiome: impacts and solutions. Environ Microbiome 16:2. https://doi.org/10.1186/s40793-020-00371-w
Lenski RE (1993) Evaluating the fate of genetically modified microorganisms in the environment: are they inherently less fit? Experientia 49:201–209. https://doi.org/10.1007/BF01923527
Lewis WH, Tahon G, Geesink P et al (2021) Innovations to culturing the uncultured microbial majority. Nat Rev Microbiol 19:225–240. https://doi.org/10.1038/s41579-020-00458-8
Li B, Ding Y, Cheng X et al (2020) Polyethylene microplastics affect the distribution of gut microbiota and inflammation development in mice. Chemosphere 244:125492. https://doi.org/10.1016/j.chemosphere.2019.125492
Li L, Xu R, Jiang L et al (2021) Effects of microplastics on immune responses of the yellow catfish Pelteobagrus fulvidraco under hypoxia. Front Physiol 12:753999. https://doi.org/10.3389/fphys.2021.753999
Liao Y-C, Nazygul J, Li M, et al (2019) Effects of microplastics on the growth, physiology, and biochemical characteristics of wheat (Triticum aestivum). Huan Jing Ke Xue 40:4661–4667. https://doi.org/10.13227/j.hjkx.201903113
Liebminger S, Eberl A, Sousa F et al (2007) Hydrolysis of PET and bis-(benzoyloxyethyl) terephthalate with a new polyesterase from Penicillium citrinum. Biocatal Biotransform 25:171–177. https://doi.org/10.1080/10242420701379734
Lim H-A, Raku T, Tokiwa Y (2005) Hydrolysis of polyesters by serine proteases. Biotechnol Lett 27:459–464. https://doi.org/10.1007/s10529-005-2217-8
Lim X (2021) Microplastics are everywhere — but are they harmful? Nature 593:22–25. https://doi.org/10.1038/d41586-021-01143-3
Limonta G, Mancia A, Benkhalqui A et al (2019) Microplastics induce transcriptional changes, immune response and behavioral alterations in adult zebrafish. Sci Rep 9:15775. https://doi.org/10.1038/s41598-019-52292-5
Lin Z, Jin T, Zou T et al (2022) Current progress on plastic/microplastic degradation: fact influences and mechanism. Environ Pollut 304:119159. https://doi.org/10.1016/j.envpol.2022.119159
Lindgreen S, Adair KL, Gardner PP (2016) An evaluation of the accuracy and speed of metagenome analysis tools. Sci Rep 6:19233. https://doi.org/10.1038/srep19233
Liu M, Lu S, Song Y et al (2018) Microplastic and mesoplastic pollution in farmland soils in suburbs of Shanghai, China. Environ Pollut 242:855–862. https://doi.org/10.1016/j.envpol.2018.07.051
Luan X, Kou X, Zhang L et al (2022) Estimation and prediction of plastic losses to the environment in China from 1950 to 2050. Resour Conserv Recycl 184:106386
Lucas N, Bienaime C, Belloy C et al (2008) Polymer biodegradation: mechanisms and estimation techniques – a review. Chemosphere 73:429–442. https://doi.org/10.1016/j.chemosphere.2008.06.064
Lv M, Jiang B, Xing Y et al (2022) Recent advances in the breakdown of microplastics: strategies and future prospectives. Environ Sci Pollut Res 29:65887–65903. https://doi.org/10.1007/s11356-022-22004-0
Mao L, Fang S, Zhao M et al (2021) Effects of bisphenol A and bisphenol S exposure at low doses on the metabolome of adolescent male Sprague-Dawley rats. Chem Res Toxicol 34:1578–1587. https://doi.org/10.1021/acs.chemrestox.1c00018
Marcus H, Ketley JM, Kaper JB, Holmes RK (1990) Effects of DNase production, plasmid size, and restriction barriers on transformation of Vibrio cholerae by electroporation and osmotic shock. FEMS Microbiol Lett 68:149–154. https://doi.org/10.1016/0378-1097(90)90141-C
Maroof L, Khan I, Yoo HS, et al (2021) Identification and characterization of low density polyethylene-degrading bacteria isolated from soils of waste disposal sites. Environ Eng Res 26. https://doi.org/10.4491/eer.2020.167
Marrone A, La Russa MF, Randazzo L et al (2021) Microplastics in the center of Mediterranean: comparison of the two calabrian coasts and distribution from coastal areas to the open sea. Int J Environ Res Public Health 18:10712. https://doi.org/10.3390/ijerph182010712
Masaki K, Kamini NR, Ikeda H, Iefuji H (2005) Cutinase-Like Enzyme from the Yeast Cryptococcus sp. Strain S-2 Hydrolyzes Polylactic Acid and Other Biodegradable Plastics. Appl Environ Microbiol 71:7548–7550. https://doi.org/10.1128/AEM.71.11.7548-7550.2005
Matsumura Y, Hosokawa C, Sasaki-Mori M et al (2009) Isolation and characterization of novel bisphenol-A–degrading bacteria from soils. Biocontrol Sci 14:161–169. https://doi.org/10.4265/bio.14.161
Matthews S, Xu EG, Dumont ER et al (2021) Polystyrene micro- and nanoplastics affect locomotion and daily activity of Drosophila melanogaster. Environ Sci: Nano 8:110–121. https://doi.org/10.1039/D0EN00942C
McGivney E, Cederholm L, Barth A et al (2020) Rapid physicochemical changes in microplastic induced by biofilm formation. Front Bioeng Biotechnol 8:205. https://doi.org/10.3389/fbioe.2020.00205
McGoran AR, Clark PF, Smith BD, Morritt D (2020) High prevalence of plastic ingestion by Eriocheir sinensis and Carcinus maenas (Crustacea: Decapoda: Brachyura) in the Thames Estuary. Environ Pollut 265:114972. https://doi.org/10.1016/j.envpol.2020.114972
Miao F, Liu Y, Gao M et al (2020) Degradation of polyvinyl chloride microplastics via an electro-Fenton-like system with a TiO2/graphite cathode. J Hazard Mater 399:123023. https://doi.org/10.1016/j.jhazmat.2020.123023
Min K, Cuiffi JD, Mathers RT (2020) Ranking environmental degradation trends of plastic marine debris based on physical properties and molecular structure. Nat Commun 11:727. https://doi.org/10.1038/s41467-020-14538-z
Mir TUG, Wani AK, Akhtar N, Shukla S (2022) CRISPR/Cas9: Regulations and challenges for law enforcement to combat its dual-use. Forensic Sci Int 334:111274. https://doi.org/10.1016/j.forsciint.2022.111274
Mohanan N, Montazer Z, Sharma PK, Levin DB (2020) Microbial and enzymatic degradation of synthetic plastics. Front Microbiol 11:580709. https://doi.org/10.3389/fmicb.2020.580709
Mohsen M, Wang Q, Zhang L et al (2019) Microplastic ingestion by the farmed sea cucumber Apostichopus japonicus in China. Environ Pollut 245:1071–1078. https://doi.org/10.1016/j.envpol.2018.11.083
Montazer Z, Habibi-Najafi MB, Mohebbi M, Oromiehei A (2018) Microbial degradation of UV-pretreated low-density polyethylene films by novel polyethylene-degrading bacteria isolated from plastic-dump soil. J Polym Environ 26:3613–3625. https://doi.org/10.1007/s10924-018-1245-0
Montero D, Rimoldi S, Torrecillas S et al (2022) Impact of polypropylene microplastics and chemical pollutants on European sea bass (Dicentrarchus labrax) gut microbiota and health. Sci Total Environ 805:150402. https://doi.org/10.1016/j.scitotenv.2021.150402
Mor R, Sivan A (2008) Biofilm formation and partial biodegradation of polystyrene by the actinomycete Rhodococcus ruber: biodegradation of polystyrene. Biodegradation 19:851–858. https://doi.org/10.1007/s10532-008-9188-0
Mouafo Tamnou EB, Tamsa Arfao A, Nougang ME et al (2021) Biodegradation of polyethylene by the bacterium Pseudomonas aeruginosa in acidic aquatic microcosm and effect of the environmental temperature. Environ Challenges 3:100056. https://doi.org/10.1016/j.envc.2021.100056
Müller R-J, Schrader H, Profe J et al (2005) Enzymatic degradation of poly(ethylene terephthalate): rapid hydrolyse using a hydrolase from T. fusca. Macromol Rapid Commun 26:1400–1405. https://doi.org/10.1002/marc.200500410
Müller WEG, Schröder HC, Wang X (2019) Inorganic polyphosphates as storage for and generator of metabolic energy in the extracellular matrix. Chem Rev 119:12337–12374. https://doi.org/10.1021/acs.chemrev.9b00460
Muncke J, Andersson A-M, Backhaus T et al (2020) Impacts of food contact chemicals on human health: a consensus statement. Environ Health 19:25. https://doi.org/10.1186/s12940-020-0572-5
Murano C, Agnisola C, Caramiello D et al (2020) How sea urchins face microplastics: uptake, tissue distribution and immune system response. Environ Pollut 264:114685. https://doi.org/10.1016/j.envpol.2020.114685
Najar IN, Sherpa MT, Das S et al (2020) Diversity analysis and metagenomic insights into antibiotic and metal resistance among Himalayan hot spring bacteriobiome insinuating inherent environmental baseline levels of antibiotic and metal tolerance. J Glob Antimicrob Resist 21:342–352. https://doi.org/10.1016/j.jgar.2020.03.026
Napper IE, Davies BFR, Clifford H et al (2020) Reaching new heights in plastic pollution—preliminary findings of microplastics on Mount Everest. One Earth 3:621–630. https://doi.org/10.1016/j.oneear.2020.10.020
Nelles M, Grünes J, Morscheck G (2016) Waste Management in Germany – development to a sustainable circular economy? Procedia Environ Sci 35:6–14. https://doi.org/10.1016/j.proenv.2016.07.001
Nourollahi A, Sedighi-Khavidak S, Mokhtari M et al (2019) Isolation and identification of low-density polyethylene (LDPE) biodegrading bacteria from waste landfill in Yazd. Int J Environ Stud 76:236–250. https://doi.org/10.1080/00207233.2018.1551986
Oberbeckmann S, Labrenz M (2020) Marine microbial assemblages on microplastics: diversity, adaptation, and role in degradation. Ann Rev Mar Sci 12:209–232. https://doi.org/10.1146/annurev-marine-010419-010633
Oceguera-Cervantes A, Carrillo-García A, López N et al (2007) Characterization of the polyurethanolytic activity of two Alicycliphilus sp. strains able to degrade polyurethane and N-methylpyrrolidone. Appl Environ Microbiol 73:6214–6223. https://doi.org/10.1128/AEM.01230-07
Oda Y, Oida N, Urakami T, Tonomura K (1997) Polycaprolactone depolymerase produced by the bacterium Alcaligenes faecalis. FEMS Microbiol Lett 152:339–343. https://doi.org/10.1111/j.1574-6968.1997.tb10449.x
Ojha N, Pradhan N, Singh S et al (2017) Evaluation of HDPE and LDPE degradation by fungus, implemented by statistical optimization. Sci Rep 7:39515. https://doi.org/10.1038/srep39515
Okubo N, Takahashi S, Nakano Y (2018) Microplastics disturb the anthozoan-algae symbiotic relationship. Mar Pollut Bull 135:83–89. https://doi.org/10.1016/j.marpolbul.2018.07.016
O’Neill A, Araújo R, Casal M et al (2007) Effect of the agitation on the adsorption and hydrolytic efficiency of cutinases on polyethylene terephthalate fibres. Enzyme Microb Technol 40:1801–1805. https://doi.org/10.1016/j.enzmictec.2007.02.012
Pabortsava K, Lampitt RS (2020) High concentrations of plastic hidden beneath the surface of the Atlantic Ocean. Nat Commun 11:4073. https://doi.org/10.1038/s41467-020-17932-9
Paço A, Duarte K, da Costa JP et al (2017) Biodegradation of polyethylene microplastics by the marine fungus Zalerion maritimum. Sci Total Environ 586:10–15. https://doi.org/10.1016/j.scitotenv.2017.02.017
Pan Z, Liu Q, Sun X et al (2022) Widespread occurrence of microplastic pollution in open sea surface waters: Evidence from the mid-North Pacific Ocean. Gondwana Res 108:31–40. https://doi.org/10.1016/j.gr.2021.10.024
Parashar N, Hait S (2021) Plastics in the time of COVID-19 pandemic: protector or polluter? Sci Total Environ 759:144274. https://doi.org/10.1016/j.scitotenv.2020.144274
Park E-J, Han J-S, Park E-J et al (2020) Repeated-oral dose toxicity of polyethylene microplastics and the possible implications on reproduction and development of the next generation. Toxicol Lett 324:75–85. https://doi.org/10.1016/j.toxlet.2020.01.008
Park SY, Kim CG (2019) Biodegradation of micro-polyethylene particles by bacterial colonization of a mixed microbial consortium isolated from a landfill site. Chemosphere 222:527–533. https://doi.org/10.1016/j.chemosphere.2019.01.159
Parthasarathy A, Miranda RR, Eddingsaas NC et al (2022) Polystyrene degradation by Exiguobacterium sp. RIT 594: preliminary evidence for a pathway containing an atypical oxygenase. Microorganisms 10:1619. https://doi.org/10.3390/microorganisms10081619
Pathak VM, Navneet (2017) Review on the current status of polymer degradation: a microbial approach. Bioresour Bioprocess 4:15. https://doi.org/10.1186/s40643-017-0145-9
Patrício Silva AL, Prata JC, Walker TR et al (2021) Increased plastic pollution due to COVID-19 pandemic: challenges and recommendations. Chem Eng J 405:126683. https://doi.org/10.1016/j.cej.2020.126683
Peng Y, Wu P, Schartup AT, Zhang Y (2021) Plastic waste release caused by COVID-19 and its fate in the global ocean. PNAS 118. https://doi.org/10.1073/pnas.2111530118
Peter Guengerich F, Yoshimoto FK (2018) Formation and cleavage of C-C bonds by enzymatic oxidation-reduction reactions. Chem Rev 118:6573–6655. https://doi.org/10.1021/acs.chemrev.8b00031
Pinnell LJ, Turner JW (2019) Shotgun metagenomics reveals the benthic microbial community response to plastic and bioplastic in a coastal marine environment. Front Microbiol 10:1252. https://doi.org/10.3389/fmicb.2019.01252
Poinar H, Schwarz C, Qi J et al (2006) Metagenomics to Paleogenomics: large-scale sequencing of mammoth DNA. Sci 311:392–394. https://doi.org/10.1126/science.1123360
Prata JC, da Costa JP, Lopes I et al (2020) Environmental exposure to microplastics: an overview on possible human health effects. Sci Total Environ 702:134455
Prüst M, Meijer J, Westerink RHS (2020) The plastic brain: neurotoxicity of micro- and nanoplastics. Part Fibre Toxicol 17:24. https://doi.org/10.1186/s12989-020-00358-y
Puspitasari N, Tsai S-L, Lee C-K (2021) Fungal hydrophobin RolA enhanced PETase hydrolysis of polyethylene terephthalate. Appl Biochem Biotechnol 193:1284–1295. https://doi.org/10.1007/s12010-020-03358-y
Qiao R, Deng Y, Zhang S et al (2019) Accumulation of different shapes of microplastics initiates intestinal injury and gut microbiota dysbiosis in the gut of zebrafish. Chemosphere 236:124334. https://doi.org/10.1016/j.chemosphere.2019.07.065
Qin J, Li R, Raes J et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65. https://doi.org/10.1038/nature08821
Radwan O, Lee JS, Stote R et al (2020) Metagenomic characterization of microbial communities on plasticized fabric materials exposed to harsh tropical environments. Int Biodeterior Biodegrad 154:105061. https://doi.org/10.1016/j.ibiod.2020.105061
Raes EJ, Karsh K, Sow SLS et al (2021) Metabolic pathways inferred from a bacterial marker gene illuminate ecological changes across South Pacific frontal boundaries. Nat Commun 12:2213. https://doi.org/10.1038/s41467-021-22409-4
Ragusa A, Svelato A, Santacroce C et al (2021) Plasticenta: first evidence of microplastics in human placenta. Environ Int 146:106274. https://doi.org/10.1016/j.envint.2020.106274
Rajandas H, Parimannan S, Sathasivam K et al (2012) A novel FTIR-ATR spectroscopy based technique for the estimation of low-density polyethylene biodegradation. Polym Testing 31:1094–1099. https://doi.org/10.1016/j.polymertesting.2012.07.015
Rana AK, Thakur MK, Saini AK et al (2022) Recent developments in microbial degradation of polypropylene: Integrated approaches towards a sustainable environment. Sci Total Environ 826:154056. https://doi.org/10.1016/j.scitotenv.2022.154056
Reichert J, Arnold AL, Hoogenboom MO et al (2019) Impacts of microplastics on growth and health of hermatypic corals are species-specific. Environ Pollut 254:113074. https://doi.org/10.1016/j.envpol.2019.113074
Restrepo-Flórez J-M, Bassi A, Thompson MR (2014) Microbial degradation and deterioration of polyethylene–A review. Int Biodeterior Biodegradation 88:83–90
Rhoads DD (2020) Computer vision and artificial intelligence are emerging diagnostic tools for the clinical microbiologist. J Clin Microbiol 58:. https://doi.org/10.1128/JCM.00511-20
Rhodes CJ (2018) Plastic pollution and potential solutions. Sci Prog 101:207–260. https://doi.org/10.3184/003685018X15294876706211
Ribitsch D, Acero EH, Greimel K et al (2012) Characterization of a new cutinase from Thermobifida alba for PET-surface hydrolysis. Biocatal Biotransform 30:2–9. https://doi.org/10.3109/10242422.2012.644435
Ritchie H, Roser M (2018) Plastic pollution. Our World in Data. https://ourworldindata.org/plastic-pollution?utm_source=newsletter. Accessed 27 Aug 2022
Rodriguez-Seijo A, Lourenço J, Rocha-Santos TAP et al (2017) Histopathological and molecular effects of microplastics in Eisenia andrei Bouché. Environ Pollut 220:495–503. https://doi.org/10.1016/j.envpol.2016.09.092
Rogers KL, Carreres-Calabuig JA, Gorokhova E, Posth NR (2020) Micro-by-micro interactions: how microorganisms influence the fate of marine microplastics. Limnol Oceanogr Lett 5:18–36. https://doi.org/10.1002/lol2.10136
Romano N, Renukdas N, Fischer H et al (2020) Differential modulation of oxidative stress, antioxidant defense, histomorphology, ion-regulation and growth marker gene expression in goldfish (Carassius auratus) following exposure to different dose of virgin microplastics. Comp Biochem Physiol Part C: Toxicol Pharmacol 238:108862. https://doi.org/10.1016/j.cbpc.2020.108862
Ronda AC, Arias AH, Rimondino GN et al (2021) Plastic impacts in argentina: a critical research review contributing to the global knowledge. Curr Environ Health Rep 8:212–222. https://doi.org/10.1007/s40572-021-00323-7
Ronkvist ÅM, Xie W, Lu W, Gross RA (2009) Cutinase-catalyzed hydrolysis of poly(ethylene terephthalate). Macromolecules 42:5128–5138. https://doi.org/10.1021/ma9005318
Rout AK, Dehury B, Parida PK et al (2022) Taxonomic profiling and functional gene annotation of microbial communities in sediment of river Ganga at Kanpur, India: insights from whole-genome metagenomics study. Environ Sci Pollut Res Int. https://doi.org/10.1007/s11356-022-21644-6
Royer S-J, Ferrón S, Wilson ST, Karl DM (2018) Production of methane and ethylene from plastic in the environment. PLoS ONE 13:e0200574
Ru J, Huo Y, Yang Y (2020) Microbial degradation and valorization of plastic wastes. Front Microbiol 11:442. https://doi.org/10.3389/fmicb.2020.00442
Russell JR, Huang J, Anand P et al (2011) Biodegradation of polyester polyurethane by endophytic fungi. Appl Environ Microbiol 77:6076–6084. https://doi.org/10.1128/AEM.00521-11
Samak NA, Jia Y, Sharshar MM et al (2020) Recent advances in biocatalysts engineering for polyethylene terephthalate plastic waste green recycling. Environ Int 145:106144. https://doi.org/10.1016/j.envint.2020.106144
Sameshima-Yamashita Y, Ueda H, Koitabashi M, Kitamoto H (2019) Pretreatment with an esterase from the yeast Pseudozyma antarctica accelerates biodegradation of plastic mulch film in soil under laboratory conditions. J Biosci Bioeng 127:93–98. https://doi.org/10.1016/j.jbiosc.2018.06.011
Sangeetha Devi R, Rajesh Kannan V, Nivas D et al (2015) Biodegradation of HDPE by Aspergillus spp. from marine ecosystem of Gulf of Mannar, India. Mar Pollut Bull 96:32–40. https://doi.org/10.1016/j.marpolbul.2015.05.050
Santo M, Weitsman R, Sivan A (2013) The role of the copper-binding enzyme – laccase – in the biodegradation of polyethylene by the actinomycete Rhodococcus ruber. Int Biodeterior Biodegradation 84:204–210. https://doi.org/10.1016/j.ibiod.2012.03.001
Scholz MB, Lo C-C, Chain PS (2012) Next generation sequencing and bioinformatic bottlenecks: the current state of metagenomic data analysis. Curr Opin Biotechnol 23:9–15. https://doi.org/10.1016/j.copbio.2011.11.013
Schyns ZO, Shaver MP (2021) Mechanical recycling of packaging plastics: a review. Macromol Rapid Commun 42:2000415
Sekiguchi T, Ebisui A, Nomura K et al (2009) Biodegradation of several fibers submerged in deep sea water and isolation of biodegradable plastic degrading bacteria from deep ocean water. Nippon Suisan Gakkaishi (Japan) 75:1011–1018. https://doi.org/10.2331/SUISAN.75.1011
Sfriso AA, Tomio Y, Rosso B et al (2020) Microplastic accumulation in benthic invertebrates in Terra Nova Bay (Ross Sea, Antarctica). Environ Int 137:105587. https://doi.org/10.1016/j.envint.2020.105587
Shams M, Alam I, Mahbub MS (2021) Plastic pollution during COVID-19: plastic waste directives and its long-term impact on the environment. Environ Adv 5:100119. https://doi.org/10.1016/j.envadv.2021.100119
Shang Y, Wang X, Chang X et al (2021) The effect of microplastics on the bioenergetics of the mussel Mytilus coruscus assessed by cellular energy allocation approach. Front Mar Sci 8:754789. https://doi.org/10.3389/fmars.2021.754789
Sharma S, Chatterjee S (2017) Microplastic pollution, a threat to marine ecosystem and human health: a short review. Environ Sci Pollut Res Int 24:21530–21547. https://doi.org/10.1007/s11356-017-9910-8
Sharma S, Sharma V, Chatterjee S (2021) Microplastics in the Mediterranean Sea: sources, pollution intensity, Sea Health, and Regulatory Policies. Front Mar Sci 8:634934. https://doi.org/10.3389/fmars.2021.634934
Shilpa, Basak N, Meena SS (2022) Exploring the plastic degrading ability of microbial communities through metagenomic approach. Mater Today: Proc. https://doi.org/10.1016/j.matpr.2022.02.308
Shin S-K, Um N, Kim Y-J et al (2020) New policy framework with plastic waste control plan for effective plastic waste management. Sustainability 12:6049. https://doi.org/10.3390/su12156049
Silva PHS, de Sousa FDB (2021) Microplastic pollution of Patos Lagoon, south of Brazil. Environ Challenges 4:100076. https://doi.org/10.1016/j.envc.2021.100076
Skariyachan S, Setlur AS, Naik SY et al (2017) Enhanced biodegradation of low and high-density polyethylene by novel bacterial consortia formulated from plastic-contaminated cow dung under thermophilic conditions. Environ Sci Pollut Res 24:8443–8457. https://doi.org/10.1007/s11356-017-8537-0
Smith M, Love DC, Rochman CM, Neff RA (2018) Microplastics in seafood and the implications for human health. Curr Environ Health Rep 5:375–386. https://doi.org/10.1007/s40572-018-0206-z
Sobhani Z, Luo Y, Gibson CT et al (2021) Collecting microplastics in gardens: case study (i) of soil. Front Environ Sci 9:371. https://doi.org/10.3389/fenvs.2021.739775
Song Y, Cao C, Qiu R et al (2019) Uptake and adverse effects of polyethylene terephthalate microplastics fibers on terrestrial snails (Achatina fulica) after soil exposure. Environ Pollut 250:447–455. https://doi.org/10.1016/j.envpol.2019.04.066
Song YK, Hong SH, Jang M et al (2017) Combined effects of UV exposure duration and mechanical abrasion on microplastic fragmentation by polymer type. Environ Sci Technol 51:4368–4376. https://doi.org/10.1021/acs.est.6b06155
Sørensen L, Rogers E, Altin D et al (2020) Sorption of PAHs to microplastic and their bioavailability and toxicity to marine copepods under co-exposure conditions. Environ Pollut 258:113844. https://doi.org/10.1016/j.envpol.2019.113844
Spini G, Spina F, Poli A et al (2018) Molecular and microbiological insights on the enrichment procedures for the isolation of petroleum degrading bacteria and fungi. Front Microbiol 9:2543. https://doi.org/10.3389/fmicb.2018.02543
Stern RV, Howard GT (2000) The polyester polyurethanase gene (pueA) from Pseudomonas chlororaphis encodes a lipase. FEMS Microbiol Lett 185:163–168. https://doi.org/10.1111/j.1574-6968.2000.tb09056.x
Strayer DL, Findlay SEG (2010) Ecology of freshwater shore zones. Aquat Sci 72:127–163. https://doi.org/10.1007/s00027-010-0128-9
Sulaiman S, Yamato S, Kanaya E et al (2012) Isolation of a novel cutinase homolog with polyethylene terephthalate-degrading activity from leaf-branch compost by using a metagenomic approach. Appl Environ Microbiol 78:1556–1562. https://doi.org/10.1128/AEM.06725-11
Sun H, Chen N, Yang X et al (2021) Effects induced by polyethylene microplastics oral exposure on colon mucin release, inflammation, gut microflora composition and metabolism in mice. Ecotoxicol Environ Saf 220:112340. https://doi.org/10.1016/j.ecoenv.2021.112340
Sun J, Li P, Liu Z et al (2020) A novel thermostable serine protease from a metagenomic library derived from marine sediments in the East China Sea. Appl Microbiol Biotechnol 104:9229–9238. https://doi.org/10.1007/s00253-020-10879-3
Sunagawa S, Coelho LP, Chaffron S et al (2015) Structure and function of the global ocean microbiome. Science 348:1261359
Sunitha TG, Monisha V, Sivanesan S et al (2021) Micro-plastic pollution along the Bay of Bengal coastal stretch of Tamil Nadu, South India. Sci Total Environ 756:144073. https://doi.org/10.1016/j.scitotenv.2020.144073
Tachibana K, Hashimoto K, Yoshikawa M, Okawa H (2010) Isolation and characterization of microorganisms degrading nylon 4 in the composted soil. Polym Degrad Stab 95:912–917. https://doi.org/10.1016/j.polymdegradstab.2010.03.031
Taghavi N, Singhal N, Zhuang W-Q, Baroutian S (2021) Degradation of plastic waste using stimulated and naturally occurring microbial strains. Chemosphere 263:127975. https://doi.org/10.1016/j.chemosphere.2020.127975
Tahir A, Samawi MF, Sari K et al (2019) Studies on microplastic contamination in seagrass beds at Spermonde Archipelago of Makassar Strait, Indonesia. J Phys: Conf Ser 1341:022008. https://doi.org/10.1088/1742-6596/1341/2/022008
Tamargo A, Molinero N, Reinosa JJ et al (2022) PET microplastics affect human gut microbiota communities during simulated gastrointestinal digestion, first evidence of plausible polymer biodegradation during human digestion. Sci Rep 12:528. https://doi.org/10.1038/s41598-021-04489-w
Tang J, Liang Y, Jiang D et al (2018) Temperature-controlled thermophilic bacterial communities in hot springs of western Sichuan, China. BMC Microbiol 18:1–14. https://doi.org/10.1186/s12866-018-1271-z
Tchigvintsev A, Tran H, Popovic A et al (2015) The environment shapes microbial enzymes: five cold-active and salt-resistant carboxylesterases from marine metagenomes. Appl Microbiol Biotechnol 99:2165–2178. https://doi.org/10.1007/s00253-014-6038-3
Teng J, Zhao J, Zhang C et al (2020) A systems analysis of microplastic pollution in Laizhou Bay. China Sci Total Environ 745:140815. https://doi.org/10.1016/j.scitotenv.2020.140815
Thushari GGN, Senevirathna JDM (2020) Plastic pollution in the marine environment. Heliyon 6:e04709. https://doi.org/10.1016/j.heliyon.2020.e04709
Toda H, Imae R, Komio T, Itoh N (2012) Expression and characterization of styrene monooxygenases of Rhodococcus sp. ST-5 and ST-10 for synthesizing enantiopure (S)-epoxides. Appl Microbiol Biotechnol 96:407–418. https://doi.org/10.1007/s00253-011-3849-3
Tongo I, Erhunmwunse NO (2022) Effects of ingestion of polyethylene microplastics on survival rate, opercular respiration rate and swimming performance of African catfish (Clarias gariepinus). J Hazard Mater 423:127237. https://doi.org/10.1016/j.jhazmat.2021.127237
Tsering T, Sillanpää M, Sillanpää M et al (2021) Microplastics pollution in the Brahmaputra River and the Indus River of the Indian Himalaya. Sci Total Environ 789:147968. https://doi.org/10.1016/j.scitotenv.2021.147968
Uchiyama T, Miyazaki K (2009) Functional metagenomics for enzyme discovery: challenges to efficient screening. Curr Opin Biotechnol 20:616–622. https://doi.org/10.1016/j.copbio.2009.09.010
Ugwu K, Herrera A, Gómez M (2021) Microplastics in marine biota: a review. Mar Pollut Bull 169:112540. https://doi.org/10.1016/j.marpolbul.2021.112540
Urbanek AK, Rymowicz W, Mirończuk AM (2018) Degradation of plastics and plastic-degrading bacteria in cold marine habitats. Appl Microbiol Biotechnol 102:7669–7678. https://doi.org/10.1007/s00253-018-9195-y
Vaid M, Mehra K, Gupta A (2021) Microplastics as contaminants in Indian environment: a review. Environ Sci Pollut Res 28:68025–68052. https://doi.org/10.1007/s11356-021-16827-6
Varó I, Osorio K, Estensoro I et al (2021) Effect of virgin low density polyethylene microplastic ingestion on intestinal histopathology and microbiota of gilthead sea bream. Aquaculture 545:737245. https://doi.org/10.1016/j.aquaculture.2021.737245
Veerasingam S, Ranjani M, Venkatachalapathy R et al (2020) Microplastics in different environmental compartments in India: analytical methods, distribution, associated contaminants and research needs. TrAC Trends Anal Chem 133:116071. https://doi.org/10.1016/j.trac.2020.116071
Venkataramana C, Botsa SM, Shyamala P, Muralikrishna R (2021) Photocatalytic degradation of polyethylene plastics by NiAl2O4 spinels-synthesis and characterization. Chemosphere 265:129021. https://doi.org/10.1016/j.chemosphere.2020.129021
Verla AW, Enyoh CE, Verla EN, Nwarnorh KO (2019) Microplastic–toxic chemical interaction: a review study on quantified levels, mechanism and implication. SN Appl Sci 1:1400. https://doi.org/10.1007/s42452-019-1352-0
Vethaak AD, Legler J (2021) Microplastics and human health. Science 371:672–674
Viršek MK, Lovšin MN, Koren Š et al (2017) Microplastics as a vector for the transport of the bacterial fish pathogen species Aeromonas salmonicida. Mar Pollut Bull 125:301–309. https://doi.org/10.1016/j.marpolbul.2017.08.024
Visalli G, Facciolà A, Pruiti Ciarello M et al (2021) Acute and sub-chronic effects of microplastics (3 and 10 µm) on the human intestinal Cells HT-29. Int J Environ Res Public Health 18:5833. https://doi.org/10.3390/ijerph18115833
Vriend P, Hidayat H, van Leeuwen J et al (2021) Plastic pollution research in Indonesia: state of science and future research directions to reduce impacts. Front Environ Sci 9:187. https://doi.org/10.3389/fenvs.2021.692907
Vroman I, Tighzert L (2009) Biodegradable polymers. Materials (basel) 2:307–344. https://doi.org/10.3390/ma2020307
Walkinshaw C, Lindeque PK, Thompson R et al (2020) Microplastics and seafood: lower trophic organisms at highest risk of contamination. Ecotoxicol Environ Saf 190:110066. https://doi.org/10.1016/j.ecoenv.2019.110066
Wang J, Khokhar I, Ren C et al (2019) Characterization and 16S metagenomic analysis of organophosphorus flame retardants degrading consortia. J Hazard Mater 380:120881. https://doi.org/10.1016/j.jhazmat.2019a.120881
Wang Y, Zhang D, Zhang M et al (2019b) Effects of ingested polystyrene microplastics on brine shrimp, Artemia parthenogenetica. Environ Pollut 244:715–722. https://doi.org/10.1016/j.envpol.2018.10.024
Wani AK, Akhtar N, Datta B et al (2021) Cyanobacteria-derived small molecules: a new class of drugs. In: Volatiles and Metabolites of Microbes, 1st edn. Academic Press, pp 283–303. https://doi.org/10.1016/B978-0-12-824523-1.00003-1
Wani AK, Akhtar N, Naqash N, et al (2022a) Bioprospecting culturable and unculturable microbial consortia through metagenomics for bioremediation. Clean Chem Eng 100017. https://doi.org/10.1016/j.clce.2022.100017
Wani AK, Akhtar N, Sher F et al (2022b) Microbial adaptation to different environmental conditions: molecular perspective of evolved genetic and cellular systems. Arch Microbiol 204:144. https://doi.org/10.1007/s00203-022-02757-5
Wani AK, Akhtar N, Singh R et al (2022c) Prospects of advanced metagenomics and meta-omics in the investigation of phytomicrobiome to forecast beneficial and pathogenic response. Mol Biol Rep. https://doi.org/10.1007/s11033-022-07936-7
Wani AK, Akhtar N, Singh R et al (2022d) Genome centric engineering using ZFNs, TALENs and CRISPR-Cas9 systems for trait improvement and disease control in Animals. Vet Res Commun 1–16. https://doi.org/10.1007/s11259-022-09967-8
Wani AK, Hashem NM, Akhtar N et al (2022e) Understanding microbial networks of farm animals through genomics, metagenomics and other meta-omic approaches for livestock wellness and sustainability. Annals Anim Sci 839–853. https://doi.org/10.2478/aoas-2022-0002
Wani AK, Rahayu F, Kadarwati FT et al (2022) Metagenomic screening strategies for bioprospecting enzymes from environmental samples. IOP Conf Ser: Earth Environ Sci 974:012003. https://doi.org/10.1088/1755-1315/974/1/012003
Wani AK, Roy P, Kumar V, Mir T ul G (2022g) Metagenomics and artificial intelligence in the context of human health. Infect Genet Evol 105267. https://doi.org/10.1016/j.meegid.2022.105267
Webb HK, Arnott J, Crawford RJ, Ivanova EP (2013) Plastic degradation and its environmental implications with special reference to poly(ethylene terephthalate). Polymers 5:1–18. https://doi.org/10.3390/polym5010001
Weinstein JE, Crocker BK, Gray AD (2016) From macroplastic to microplastic: degradation of high-density polyethylene, polypropylene, and polystyrene in a salt marsh habitat. Environ Toxicol Chem 35:1632–1640. https://doi.org/10.1002/etc.3432
Wooley JC, Godzik A, Friedberg I (2010) A primer on metagenomics. PLOS Comput Biol 6:e1000667. https://doi.org/10.1371/journal.pcbi.1000667
Wright RJ, Bosch R, Langille MGI et al (2021) A multi-OMIC characterisation of biodegradation and microbial community succession within the PET plastisphere. Microbiome 9:141. https://doi.org/10.1186/s40168-021-01054-5
Wu B, Wu X, Liu S et al (2019) Size-dependent effects of polystyrene microplastics on cytotoxicity and efflux pump inhibition in human Caco-2 cells. Chemosphere 221:333–341. https://doi.org/10.1016/j.chemosphere.2019.01.056
Wu Q, Liu S, Chen P et al (2021) Microplastics in seawater and two sides of the Taiwan Strait: reflection of the social-economic development. Mar Pollut Bull 169:112588. https://doi.org/10.1016/j.marpolbul.2021.112588
Xia X, Sun M, Zhou M et al (2020) Polyvinyl chloride microplastics induce growth inhibition and oxidative stress in Cyprinus carpio var. larvae. Sci Total Environ 716:136479. https://doi.org/10.1016/j.scitotenv.2019.136479
Yang Y, Liu W, Zhang Z et al (2020) Microplastics provide new microbial niches in aquatic environments. Appl Microbiol Biotechnol 104:6501–6511. https://doi.org/10.1007/s00253-020-10704-x
Ye G, Zhang X, Liu X et al (2021) Polystyrene microplastics induce metabolic disturbances in marine medaka (Oryzias melastigmas) liver. Sci Total Environ 782:146885. https://doi.org/10.1016/j.scitotenv.2021.146885
Ye S, Cheng M, Zeng G et al (2020) Insights into catalytic removal and separation of attached metals from natural-aged microplastics by magnetic biochar activating oxidation process. Water Res 179:115876. https://doi.org/10.1016/j.watres.2020.115876
Ye X, Wang P, Wu Y et al (2020b) Microplastic acts as a vector for contaminants: the release behavior of dibutyl phthalate from polyvinyl chloride pipe fragments in water phase. Environ Sci Pollut Res 27:42082–42091. https://doi.org/10.1007/s11356-020-10136-0
Yoshida S, Hiraga K, Takehana T et al (2016) A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351:1196–1199. https://doi.org/10.1126/science.aad6359
Yoshikawa M, Zhang M, Toyota K (2016) Integrated anaerobic-aerobic biodegradation of multiple contaminants including chlorinated ethylenes, benzene, toluene, and dichloromethane. Water Air Soil Pollut 228:25. https://doi.org/10.1007/s11270-016-3216-1
Zaidi SSA, Kayani MUR, Zhang X et al (2021) Prediction and analysis of metagenomic operons via MetaRon: a pipeline for prediction of Metagenome and whole-genome opeRons. BMC Genomics 22:60. https://doi.org/10.1186/s12864-020-07357-5
Zaki MRM, Ying PX, Zainuddin AH et al (2021) Occurrence, abundance, and distribution of microplastics pollution: an evidence in surface tropical water of Klang River estuary, Malaysia. Environ Geochem Health 43:3733–3748. https://doi.org/10.1007/s10653-021-00872-8
Zeaiter Z, Mapelli F, Crotti E, Borin S (2018) Methods for the genetic manipulation of marine bacteria. Electron J Biotechnol 33:17–28. https://doi.org/10.1016/j.ejbt.2018.03.003
Zeenat, Elahi A, Bukhari DA et al (2021) Plastics degradation by microbes: a sustainable approach. J King Saud Univ-Sci 33:101538. https://doi.org/10.1016/j.jksus.2021.101538
Zerhouni K, Abbouni B, Kanoun K et al (2018) Isolation and identification of low density polythene-degrading bacteria from soil of North West of Algeria. South Asian J Exp Biol 8:76–82. https://doi.org/10.38150/sajeb.8(3).p76-82
Zhang D, Liu X, Huang W et al (2020) Microplastic pollution in deep-sea sediments and organisms of the Western Pacific Ocean. Environ Pollut 259:113948. https://doi.org/10.1016/j.envpol.2020.113948
Zhang G, Wang M, Xu H, Song Y (2021) Global and regional prediction and evaluation model of plastic pollution. IOP Publishing 692:032080. https://doi.org/10.1088/1755-1315/692/3/032080
Zheng Y, Li J, Cao W et al (2019) Distribution characteristics of microplastics in the seawater and sediment: a case study in Jiaozhou Bay, China. Sci Total Environ 674:27–35. https://doi.org/10.1016/j.scitotenv.2019.04.008
Zhou D, Chen J, Wu J et al (2021) Biodegradation and catalytic-chemical degradation strategies to mitigate microplastic pollution. Sustain Mater Technol 28:e00251. https://doi.org/10.1016/j.susmat.2021.e00251
Zhu D, Sethupathy S, Gao L et al (2022) Microbial diversity and community structure in deep-sea sediments of South Indian Ocean. Environ Sci Pollut Res. https://doi.org/10.1007/s11356-022-19157-3
Zhu J, Zhang Q, Li Y et al (2019) Microplastic pollution in the Maowei Sea, a typical mariculture bay of China. Sci Total Environ 658:62–68. https://doi.org/10.1016/j.scitotenv.2018.12.192
Zhu L, Bai H, Chen B et al (2018) Microplastic pollution in North Yellow Sea, China: observations on occurrence, distribution and identification. Sci Total Environ 636:20–29. https://doi.org/10.1016/j.scitotenv.2018.04.182
Zrimec J, Kokina M, Jonasson S et al (2021) Plastic-degrading potential across the global microbiome correlates with recent pollution trends. Mbio 12:e02155-e2221
Author information
Authors and Affiliations
Contributions
Atif Khurshid Wani: conceptualization, methodology, visualization, data curation, writing—original draft, writing—review & editing. Nahid Akhtar: methodology, visualization, data curation, writing—original draft, writing—review & editing. Nafiaah Naqash: visualization, data curation, writing—original draft, writing—review & editing. Farida Rahayu: data curation, writing—original draft, writing—review & editing. Djajadi Djajadi: data curation, writing—original draft, writing—review & editing. Chirag Chopra: data curation, writing—original draft, writing – review & editing. Reena Singh: visualization, data curation, writing—original draft, writing—review & editing. Sikandar I. Mulla: writing—original draft, writing—review & editing. Farooq Sher: writing—original draft, writing—review & editing. Juliana Heloisa Pinê Américo-Pinheiro: conceptualization, methodology, visualization, data curation, writing—original draft, visualization, writing—review & editing, supervision, project administration.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wani, A.K., Akhtar, N., Naqash, N. et al. Discovering untapped microbial communities through metagenomics for microplastic remediation: recent advances, challenges, and way forward. Environ Sci Pollut Res 30, 81450–81473 (2023). https://doi.org/10.1007/s11356-023-25192-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-25192-5