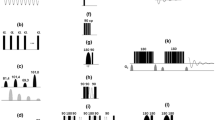
Abstract
Introduction
Nuclear Magnetic Resonance (NMR) spectroscopy stands as a preeminent analytical tool in the field of metabolomics. Nevertheless, when it comes to identifying metabolites present in scant amounts within various types of complex mixtures such as plants, honey, milk, and biological fluids and tissues, NMR-based metabolomics presents a formidable challenge. This predicament arises primarily from the fact that the signals emanating from metabolites existing in low concentrations tend to be overshadowed by the signals of highly concentrated metabolites within NMR spectra.
Objectives
The aim of this study is to tackle the issue of intense sugar signals overshadowing the desired metabolite signals, an optimal pulse sequence with band-selective excitation has been proposed for the suppression of sugar’s moiety signals (SSMS). This sequence serves the crucial purpose of suppressing unwanted signals, with a particular emphasis on mitigating the interference caused by sugar moieties' signals.
Methods
We have implemented this comprehensive approach to various NMR techniques, including 1D 1H presaturation (presat), 2D J-resolved (RES), 2D 1H-1H Total Correlation Spectroscopy (TOCSY), and 2D 1H-13C Heteronuclear Single Quantum Coherence (HSQC) for the samples of dates-flesh, honey, a standard stock solution of glucose, and nine amino acids, and commercial fetal bovine serum (FBS).
Results
The outcomes of this approach were significant. The suppression of the high-intensity sugar signals has considerably enhanced the visibility and sensitivity of the signals emanating from the desired metabolites.
Conclusion
This, in turn, enables the identification of a greater number of metabolites. Additionally, it streamlines the experimental process, reducing the time required for the comparative quantification of metabolites in statistical studies in the field of metabolomics.





Similar content being viewed by others
References
Alexandersson, E., Sandström, C., Lundqvist, L. C., & Nestor, G. (2020). Band-selective NMR experiments for suppression of unwanted signals in complex mixtures. RSC Advances, 10(54), 32511–32515. https://doi.org/10.1039/D0RA06828D
Chandra, K., Al-Harthi, S., Almulhim, F., Emwas, A.-H., Jaremko, Ł, & Jaremko, M. (2021a). The robust NMR toolbox for metabolomics. Molecular Omics, 17(5), 719–724. https://doi.org/10.1039/D1MO00118C
Chandra, K., Al-Harthi, S., Sukumaran, S., Almulhim, F., Emwas, A.-H., Atreya, H. S., Jaremko, Ł, & Jaremko, M. (2021b). NMR-based metabolomics with enhanced sensitivity. RSC Advances, 11(15), 8694–8700. https://doi.org/10.1039/D1RA01103K
Dasgupta, S., Ghosh, N., Choudhury, P., Joshi, M., Chowdhury, S. R., Bhattacharyya, P., & Chaudhury, K. (2022). NMR metabolomic and microarray-based transcriptomic data integration identifies unique molecular signatures of hypersensitivity pneumonitis. Molecular Omics, 18(2), 101–111. https://doi.org/10.1039/D1MO00209K
Emwas, A.-H., Szczepski, K., McKay, R. T., Asfour, H., Chang, C.-k., Lachowicz, J., & Jaremko, M. (2021). Pharmacometabolomics: A new horizon in personalized medicine. In Metabolomics-Methodology and Applications in Medical Sciences and Life Sciences. IntechOpen. https://doi.org/10.5772/intechopen.98911
Emwas, A.-H.M. (2015). The strengths and weaknesses of NMR spectroscopy and mass spectrometry with particular focus on metabolomics research. Metabonomics: Methods and Protocols. https://doi.org/10.1007/978-1-4939-2377-9_13
Emwas, A.-H., Alghrably, M., Al-Harthi, S., Poulson, B. G., Szczepski, K., Chandra, K., & Jaremko, M. (2019). New advances in fast methods of 2D NMR experiments. Nuclear Magnetic Resonance. https://doi.org/10.5772/intechopen.90263
Emwas, A.-H.M., Al-Rifai, N., Szczepski, K., Alsuhaymi, S., Rayyan, S., Almahasheer, H., Jaremko, M., Brennan, L., & Lachowicz, J. I. (2021b). You are what you eat: Application of metabolomics approaches to advance nutrition research. Foods, 10(6), 1249. https://doi.org/10.3390/foods10061249
Emwas, A.-H., Roy, R., McKay, R. T., Tenori, L., Saccenti, E., Gowda, G. N., Raftery, D., Alahmari, F., Jaremko, L., & Jaremko, M. (2019b). NMR Spectroscopy for Metabolomics Research. Metabolites, 9(7), 123. https://doi.org/10.3390/metabo9070123
Emwas, A.-H., Saunders, M., Ludwig, C., & Günther, U. (2008). Determinants for optimal enhancement in ex situ DNP experiments. Applied Magnetic Resonance, 34, 483–494. https://doi.org/10.1007/s00723-008-0120-x
Emwas, A.-H., Szczepski, K., Poulson, B. G., Chandra, K., McKay, R. T., Dhahri, M., Alahmari, F., Jaremko, L., Lachowicz, J. I., & Jaremko, M. (2020). NMR as a “gold standard” method in drug design and discovery. Molecules, 25(20), 4597. https://doi.org/10.3390/molecules25204597
Fan, T.W.-M., & Lane, A. N. (2016). Applications of NMR spectroscopy to systems biochemistry. Progress in Nuclear Magnetic Resonance Spectroscopy, 92, 18–53. https://doi.org/10.1016/j.pnmrs.2016.01.005
Foroozandeh, M., Adams, R. W., Meharry, N. J., Jeannerat, D., Nilsson, M., & Morris, G. A. (2014). Ultrahigh-resolution NMR spectroscopy. Angewandte Chemie International Edition, 53(27), 6990–6992. https://doi.org/10.1002/anie.201404111
Gathungu, R. M., Kautz, R., Kristal, B. S., Bird, S. S., & Vouros, P. (2020). The integration of LC-MS and NMR for the analysis of low molecular weight trace analytes in complex matrices. Mass Spectrometry Reviews, 39(1–2), 35–54. https://doi.org/10.1002/mas.21575
Geen, H., & Freeman, R. (1991). Band-selective radiofrequency pulses. Journal of Magnetic Resonance, 93(1), 93–141. https://doi.org/10.1016/0022-2364(91)90034-
Gowda, G. N., & Raftery, D. (2015). Can NMR solve some significant challenges in metabolomics? Journal of Magnetic Resonance, 260, 144–160. https://doi.org/10.1016/j.jmr.2015.07.014
Hansen, A. L., Kupče, E. R., Li, D.-W., Bruschweiler-Li, L., Wang, C., & Brüschweiler, R. (2021). 2D NMR-based metabolomics with HSQC/TOCSY NOAH supersequences. Analytical Chemistry, 93(15), 6112–6119. https://doi.org/10.1021/acs.analchem.0c05205
Hazrati, H., Kudsk, P., Ding, L., Uthe, H., & Fomsgaard, I. S. (2022). Integrated LC–MS and GC–MS-based metabolomics reveal the effects of plant competition on the rye metabolome. Journal of Agricultural and Food Chemistry, 70(9), 3056–3066. https://doi.org/10.1021/acs.jafc.1c06306
Hu, Y., Cheng, K., He, L., Zhang, X., Jiang, B., Jiang, L., Li, C., Wang, G., Yang, Y., & Liu, M. (2021). NMR-based methods for protein analysis. Analytical Chemistry, 93(4), 1866–1879. https://doi.org/10.1021/acs.analchem.0c03830
Hwang, T.-L., & Shaka, A. (1995). Water suppression that works. Excitation sculpting using arbitrary wave-forms and pulsed-field gradients. Journal of Magnetic Resonance, Series A, 112(2), 275–279. https://doi.org/10.1006/jmra.1995.1047
Kew, W., Bell, N. G., Goodall, I., & Uhrín, D. (2017). Advanced solvent signal suppression for the acquisition of 1D and 2D NMR spectra of Scotch Whisky. Magnetic Resonance in Chemistry, 55(9), 785–796. https://doi.org/10.1002/mrc.4621
Kim, H. K., Choi, Y. H., & Verpoorte, R. (2010). NMR-based metabolomic analysis of plants. Nature Protocols, 5(3), 536–549. https://doi.org/10.1038/nprot.2009.237
Lanzotti, V., Anzano, A., Grauso, L., Zotti, M., Sacco, A., Senatore, M., Moreno, M., Diano, M., Parente, M., & Esposito, S. (2022). NMR metabolomics and chemometrics of lettuce, Lactuca sativa L. under different foliar organic fertilization treatments. Plants, 11(16), 2164. https://doi.org/10.3390/plants11162164
Liu, D., Chen, Y.-Q., Xiao, X.-W., Zhong, R.-T., Yang, C.-F., Liu, B., & Zhao, C. (2019). Nutrient properties and nuclear magnetic resonance-based metabonomic analysis of macrofungi. Foods, 8(9), 397. https://doi.org/10.3390/foods8090397
Lohr, K. E., Khattri, R. B., Guingab-Cagmat, J., Camp, E. F., Merritt, M. E., Garrett, T. J., & Patterson, J. T. (2019). Metabolomic profiles differ among unique genotypes of a threatened Caribbean coral. Scientific Reports, 9(1), 6067. https://doi.org/10.1038/s41598-019-42434-0
Mariadoss, A. V., Vinyagam, R., Rajamanickam, V., Sankaran, V., Venkatesan, S., & David, E. (2019). Pharmacological aspects and potential use of phloretin: A systemic review. Mini Reviews in Medicinal Chemistry, 19(13), 1060–1067. https://doi.org/10.2174/1389557519666190311154425
Martineau, E., Dumez, J. N., & Giraudeau, P. (2020). Fast quantitative 2D NMR for metabolomics and lipidomics: A tutorial. Magnetic Resonance in Chemistry, 58(5), 390–403. https://doi.org/10.1002/mrc.4899
Morris, G. A., Aguilar, J. A., Evans, R., Haiber, S., & Nilsson, M. (2010). True chemical shift correlation maps: A TOCSY experiment with pure shifts in both dimensions. Journal of the American Chemical Society, 132(37), 12770–12772. https://doi.org/10.1021/ja1039715
Nagana Gowda, G., & Raftery, D. (2021). NMR-based metabolomics. Cancer Metabolomics: Methods and Applications. https://doi.org/10.1007/978-3-030-51652-9_2
Pan, Z., & Raftery, D. (2007). Comparing and combining NMR spectroscopy and mass spectrometry in metabolomics. Analytical and Bioanalytical Chemistry, 387, 525–527. https://doi.org/10.1007/s00216-006-0687-8
Perez de Souza, L., Alseekh, S., Scossa, F., & Fernie, A. R. (2021). Ultra-high-performance liquid chromatography high-resolution mass spectrometry variants for metabolomics research. Nature Methods, 18(7), 733–746. https://doi.org/10.1038/s41592-021-01116-4
Powers, R. (2007). Functional genomics and NMR spectroscopy. Combinatorial Chemistry & High Throughput Screening, 10(8), 676–697. https://doi.org/10.2174/138620707782507331
Rucker, S. P., & Shaka, A. (1989). Broadband homonuclear cross polarization in 2D NMR using DIPSI-2. Molecular Physics, 68(2), 509–517. https://doi.org/10.1080/00268978900102331
Sanchez, L. J., Zhu, D., Frew, R., & Kebede, B. (2021). Optimization of nuclear magnetic resonance and gas chromatography-mass spectrometry-based fingerprinting methods to characterize goat milk powder. Journal of Dairy Science, 104(1), 102–111. https://doi.org/10.3168/jds.2020-18467
Scarsini, M., Thurotte, A., Veidl, B., Amiard, F., Niepceron, F., Badawi, M., Lagarde, F., Schoefs, B., & Marchand, J. (2021). Metabolite quantification by Fourier Transform Infrared Spectroscopy in diatoms: Proof of concept on Phaeodactylum tricornutum. Frontiers in Plant Science, 12, 756421. https://doi.org/10.3389/fpls.2021.756421
Shah, Z., BADSHAH, S., Iqbal, A., Emwas, A.-H., & Jaremko, M. (2022). GC-MS based metabolomics and lipidiomics analyses of selected freshwater green macroalgae. Doi: https://doi.org/10.21203/rs.3.rs-1324666/v1
Simpson, A. J., Simpson, M. J., & Soong, R. (2018). Environmental nuclear magnetic resonance spectroscopy: An overview and a primer. Analytical Chemistry, 90(1), 628–639. https://doi.org/10.1021/acs.analchem.7b03241
Singh, U., Alsuhaymi, S., Al-Nemi, R., Emwas, A.-H., & Jaremko, M. (2023). Compound-specific 1D 1H NMR pulse sequence selection for metabolomics analyses. ACS Omega. https://doi.org/10.1021/acsomega.3c01688
Singh, U., Bhattacharya, S., & Baishya, B. (2020). Pure shift HMQC: Resolution and sensitivity enhancement by bilinear rotation decoupling in the indirect and direct dimensions. Journal of Magnetic Resonance, 311, 106684. https://doi.org/10.1016/j.jmr.2020.106684
Singh, U., Verma, A., & Baishya, B. (2017). Parallel acquisition of slice-selective 1H–1H soft COSY spectra. Journal of Magnetic Resonance, 284, 80–85. https://doi.org/10.1016/j.jmr.2017.09.012
Smallcombe, S. H., Patt, S. L., & Keifer, P. A. (1995). WET solvent suppression and its applications to LC NMR and high-resolution NMR spectroscopy. Journal of Magnetic Resonance, Series A, 117(2), 295–303. https://doi.org/10.1006/jmra.1995.0759
Sogin, E. M., Putnam, H. M., Nelson, C. E., Anderson, P., & Gates, R. D. (2017). Correspondence of coral holobiont metabolome with symbiotic bacteria, archaea and Symbiodinium communities. Environmental Microbiology Reports, 9(3), 310–315. https://doi.org/10.1111/1758-2229.12541
Son, H.-S., Hwang, G.-S., Kim, K. M., Ahn, H.-J., Park, W.-M., Van Den Berg, F., Hong, Y.-S., & Lee, C.-H. (2009). Metabolomic studies on geographical grapes and their wines using 1H NMR analysis coupled with multivariate statistics. Journal of Agricultural and Food Chemistry, 57(4), 1481–1490. https://doi.org/10.1021/jf803388w
Song, Z., Wang, H., Yin, X., Deng, P., & Jiang, W. (2019). Application of NMR metabolomics to search for human disease biomarkers in blood. Clinical Chemistry and Laboratory Medicine (CCLM), 57(4), 417–441. https://doi.org/10.1515/cclm-2018-0380
Spiteri, C., Lia, F., & Farrugia, C. (2020). Determination of the geographical origin of Maltese honey using 1H NMR fingerprinting. Foods, 9(10), 1455. https://doi.org/10.3390/foods9101455
Szczepski, K., Al-Younis, I., Dhahri, M., Lachowicz, J. I., Al-Talla, Z. A., Almahasheer, H., Alasmael, N., Rahman, M., Emwas, A.-H., & Jaremko, Ł. (2023). Metabolic biomarkers in cancer. In Metabolomics (pp. 173–198). Elsevier. https://doi.org/10.1016/B978-0-323-99924-3.00005-4
Thrippleton, M. J., & Keeler, J. (2003). Elimination of zero-quantum interference in two-dimensional NMR spectra. Angewandte Chemie International Edition, 42(33), 3938–3941. https://doi.org/10.1002/anie.200351947
Yuan, J., Zhang, B., Wang, C., & Brüschweiler, R. (2018). Carbohydrate background removal in metabolomics samples. Analytical Chemistry, 90(24), 14100–14104. https://doi.org/10.1021/acs.analchem.8b04482
Zangger, K., & Sterk, H. (1997). Homonuclear broadband-decoupled NMR spectra. Journal of Magnetic Resonance, 124(2), 486–489. https://doi.org/10.1006/jmre.1996.1063
Acknowledgements
The authors would like to thank King Abdullah University of Science and Technology (KAUST) for financial support. The Smart Health Initiative (SHI) is also acknowledged by Mariusz Jaremko for grants from the Baseline (BAS/1/1085-01-01) program for the period of 2021 to 2023.
Author information
Authors and Affiliations
Contributions
US was primarily responsible for the design of the methods, execution of these methods, studies, data collection, and data analysis. Additionally, US prepared the figures and tables and wrote the main manuscript text and RAl-N provided considerable assistance in the execution of these methods, studies, data collection, data analysis, figures, tables, and writing manuscript text. FA contributed significantly to providing honey samples, studies, data analysis, and editing the manuscript text. A-HE and MJ contributed significantly to assisting in designing methods, execution of these methods, studies, data collection, data analysis, writing, and editing manuscript text. All authors reviewed the manuscript. The NMR analysis was performed at the Core Lab of NMR, King Abdullah University of Science and Technology (KAUST). NMR core facility is funded by the Mariusz Jaremko-Smart-Health Initiative (SHI) and Red Sea Research Center (RSRC), Division of Biological and Environmental Sciences and Engineering (BESE), King Abdullah University of Science and Technology (KAUST), Thuwal, Makkah, 23955-6900, Saudi Arabia and Authority.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11306_2023_2069_MOESM1_ESM.docx
Experimental acquisition and processing parameters tables, in 1D and 2D NMR spectral for comparison of honey, dates-flesh, a mixture of glucose and nine metabolites as well as fetal bovine serum samples, Table of overall estimation of SNR, pulse sequences of current modified 1D and 2D NMR experiments, their pulse programming. (DOCX 7335 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, U., Al-Nemi, R., Alahmari, F. et al. Improving quality of analysis by suppression of unwanted signals through band-selective excitation in NMR spectroscopy for metabolomics studies. Metabolomics 20, 7 (2024). https://doi.org/10.1007/s11306-023-02069-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-023-02069-9