Abstract
Introduction
In the search for new potential therapies for pathologies of oxalate, such as kidney stone disease and primary hyperoxaluria, the intestinal microbiome has generated significant interest. Resident oxalate-degrading bacteria inhabit the gastrointestinal tract and reduce absorption of dietary oxalate, thereby potentially lowering the potency of oxalate as a risk factor for kidney stone formation. Although several species of bacteria have been shown to degrade oxalate, select strains of Oxalobacter formigenes (O. formigenes) have thus far demonstrated the unique ability among oxalotrophs to initiate a net intestinal oxalate secretion into the lumen from the bloodstream, allowing them to feed on both dietary and endogenous metabolic oxalate. There is significant interest in this function as a potential therapeutic application for circulating oxalate reduction, although its mechanism of action is still poorly understood. Since this species-exclusive, oxalate-regulating function is reported to be dependent on the use of a currently unidentified secreted bioactive compound, there is much interest in whether O. formigenes produces unique biochemicals that are not expressed by other oxalotrophs which lack the ability to transport oxalate. Hence, this study sought to analyze and compare the metabolomes of O. formigenes and another oxalate degrader, Bifidobacterium animalis subsp. lactis (B. animalis), to determine whether O. formigenes could produce features undetectable in another oxalotroph, thus supporting the theory of a species-exclusive secretagogue compound.
Methods
A comparative metabolomic analysis of O. formigenes strain HC1 (a human isolate) versus B. animalis, another oxalate-degrading human intestinal microbe, was performed by ultra-high-performance liquid chromatography-high-resolution mass spectrometry (UHPLC-HRMS). Bacteria were cultured independently in anaerobic conditions, harvested, lysed, and extracted by protein precipitation. Metabolite extracts were chromatographically separated and analyzed by UHPLC-HRMS using reverse phase gradient elution (ACE Excel 2 C18-Pentafluorophenyl column) paired with a Q Exactive™ mass spectrometer.
Objectives
The purpose of this study was to assess whether O. formigenes potentially produces unique biochemicals from other oxalate degraders to better understand its metabolic profile and provide support for the theoretical production of a species-exclusive secretagogue compound responsible for enhancing intestinal oxalate secretion.
Results
We report a panel of metabolites and lipids detected in the O. formigenes metabolome which were undetectable in B. animalis, several of which were identified either by mass-to-charge ratio and retention time matching to our method-specific metabolite library or MS/MS fragmentation. Furthermore, re-examination of data from our previous work showed most of these features were also undetected in the metabolomes of Lactobacillus acidophilus and Lactobacillus gasseri, two other intestinal oxalate degraders.
Conclusions
Our observation of O. formigenes metabolites and lipids which were undetectable in other oxalotrophs suggests that this bacterium likely holds the ability to produce biochemicals not expressed by at least a selection of other oxalate degraders. These findings provide support for the hypothesized biosynthesis of a species-exclusive secretagogue responsible for the stimulation of net intestinal oxalate secretion.
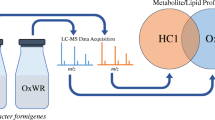
Graphic abstract




Similar content being viewed by others
References
Allison, M. J., Dawson, K. A., Mayberry, W. R., & Foss, J. G. (1985). Oxalobacter formigenes gen. nov., sp. nov.: Oxalate-degrading anaerobes that inhabit the gastrointestinal tract. Archives of Microbiology, 141, 1–7.
Allison, M. J., Macgregor, B. J., Sharp, R., & Stahl, D. A. (2005). Bergey’s Manual of Systematics of Archaea and Bacteria. Oxalobacter: Wiley.
Bertin, P. N., Heinrich-Salmeron, A., Pelletier, E., Goulhen-Chollet, F., Arsène-Ploetze, F., Gallien, S., et al. (2011). Metabolic diversity among main microorganisms inside an arsenic-rich ecosystem revealed by meta- and proteo-genomics. ISME Journal, 5, 1735–1747.
Chamberlain, C. A., Hatch, M., & Garrett, T. J. (2019a). Metabolomic and lipidomic characterization of Oxalobacter formigenes strains HC1 and OxWR by UHPLC-HRMS. Analytical and Bioanalytical Chemistry, 411(19), 4807–4818.
Chamberlain, C. A., Hatch, M., & Garrett, T. J. (2019b). Metabolomic profiling of oxalate-degrading probiotic Lactobacillus acidophilus and Lactobacillus gasseri. PLoS ONE, 14, e0222393.
Chamberlain, C. A., Hatch, M., & Garrett, T. J. (2020). Metabolomic alteration in the mouse distal colonic mucosa after oral gavage with oxalobacter formigenes. Metabolites, 10, 405.
Chamberlain, C. A., Rubio, V. Y., & Garrett, T. J. (2019). Impact of matrix effects and ionization efficiency in non-quantitative untargeted metabolomics. Metabolomics, 15, 135.
Chong, J., Soufan, O., Li, C., Caraus, I., Li, S., Bourque, G., et al. (2018). MetaboAnalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Research, 46, W486–W494.
Crits-Christoph, A., Diamond, S., Butterfield, C. N., Thomas, B. C., & Banfield, J. F. (2018). Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis. Nature, 558, 440–444.
Fahey, R. C., Brown, W. C., Adams, W. B., & Worsham, M. B. (1978). Occurrence of glutathione in bacteria. Journal of Bacteriology, 133, 1126–1129.
Gil De La Fuente, A., Godzien, J., Fernándezlópez, M., Rupérez, F. J., Barbas, C., & Otero, A. (2018). Knowledge-based metabolite annotation tool: CEU Mass Mediator. Journal of Pharmaceutical and Biomedical Analysis, 154, 138–149.
Green, R., Hanfrey, C. C., Elliott, K. A., McCloskey, D. E., Wang, X., Kanugula, S., et al. (2011). Independent evolutionary origins of functional polyamine biosynthetic enzyme fusions catalysing de novo diamine to triamine formation. Molecular Microbiology, 81, 1109–1124.
Hatch, M. (2017). Gut microbiota and oxalate homeostasis. Annals of Translation Medicine, 5, 36.
Hatch, M., Cornelius, J., Allison, M., Sidhu, H., Peck, A., & Freel, R. W. (2006). Oxalobacter sp. reduces urinary oxalate excretion by promoting enteric oxalate secretion. Kidney International, 69, 691–698.
Hatch, M., Gjymishka, A., Salido, E. C., Allison, M. J., & Freel, R. W. (2011). Enteric oxalate elimination is induced and oxalate is normalized in a mouse model of primary hyperoxaluria following intestinal colonization with Oxalobacter. American Journal of Physiology. Gastrointestinal and Liver Physiology, 300, G461–G469.
He, L., Diedrich, J., Chu, Y. Y., & Yates, J. R. (2015). Extracting accurate precursor information for tandem mass spectra by rawconverter. Analytical Chemistry, 87, 11361–11367.
Hobley, L., Li, B., Wood, J. L., Kim, S. H., Naidoo, J., Ferreira, A. S., et al. (2017). Spermidine promotes. Journal of Biological Chemistry, 292, 12041–12053.
Holm, S. (1979). A simple sequentially rejective multiple test procedure. Scandinavian Journal of Statistics, 6, 65–70.
Johnson, B. H., & Hecht, M. H. (1994). Recombinant proteins can be isolated from E. coli cells by repeated cycles of freezing and thawing. Biotechnology (N Y), 12, 1357–1360.
Klimesova, K., Whittamore, J. M., & Hatch, M. (2015). Bifidobacteriumanimalis subsp. lactis decreases urinary oxalate excretion in a mouse model of primary hyperoxaluria. Urolithiasis, 43, 107–117.
Li, Q., Chen, Q., Ruan, H., Zhu, D., & He, G. (2010). Isolation and characterisation of an oxygen, acid and bile resistant Bifidobacterium animalis subsp. lactis Qq08. Journal of the Science of Food and Agriculture, 90, 1340–1346.
Miller-Fleming, L., Olin-Sandoval, V., Campbell, K., & Ralser, M. (2015). Remaining mysteries of molecular biology: The role of polyamines in the cell. Journal of Molecular Biology, 427, 3389–3406.
O’Donnell, V. B., Dennis, E. A., Wakelam, M. J. O., & Subramaniam, S. (2019). LIPID MAPS: Serving the next generation of lipid researchers with tools, resources, data, and training. Science Signalling. https://doi.org/10.1126/scisignal.aaw2964.
Phale, P. S., Basu, A., Majhi, P. D., Deveryshetty, J., Vamsee-Krishna, C., & Shrivastava, R. (2007). Metabolic diversity in bacterial degradation of aromatic compounds. OMICS: A Journal of Integrative Biology, 11, 252–279.
Pluskal, T., Castillo, S., Villar-Briones, A., & Oresic, M. (2010). MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics, 11, 395.
Pophaly, S. D., Singh, R., Kaushik, J. K., & Tomar, S. K. (2012). Current status and emerging role of glutathione in food grade lactic acid bacteria. Microbial Cell Factories, 11, 114.
Sies, H. (1999). Glutathione and its role in cellular functions. Free Radical Biology and Medicine, 27, 916–921.
Sohlenkamp, C., & Geiger, O. (2016). Bacterial membrane lipids: Diversity in structures and pathways. FEMS Microbiology Reviews, 40, 133–159.
Sumner, L. W., Amberg, A., Barrett, D., Beale, M. H., Beger, R., Daykin, C. A., et al. (2007). Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics, 3, 211–221.
Tholl, D., Ober, D., Martin, W., Kellermann, J., & Hartmann, T. (1996). Purification, molecular cloning and expression in Escherichiacoli of homospermidine synthase from Rhodopseudomonas viridis. European Journal of Biochemistry, 240, 373–379.
van den Berg, R. A., Hoefsloot, H. C., Westerhuis, J. A., Smilde, A. K., & van der Werf, M. J. (2006). Centering, scaling, and transformations: Improving the biological information content of metabolomics data. BMC Genomics, 7, 142.
Wishart, D. S., Feunang, Y. D., Marcu, A., Guo, A. C., Liang, K., Vázquez-Fresno, R., et al. (2018). HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Research, 46, D608–D617.
Xu, B., Wang, Y., & Chen, Y. (1994). Isolation and identification of an oxygen-tolerant strain of Bifidobacterium from human feces. Wei Sheng Wu Xue Bao, 34, 9–13.
Yoshida, T., Sakamoto, A., Terui, Y., Takao, K., Sugita, Y., Yamamoto, K., et al. (2016). Effect of spermidine analogues on cell growth of Escherichiacoli polyamine requiring mutant MA261. PLoS ONE, 11, e0159494.
Acknowledgements
The authors would like to acknowledge Vanessa Y. Rubio (Department of Chemistry, University of Florida) for her contribution to the graphical abstract.
Funding
This work was funded by the National Institutes of Health grant 2R01DK088892-05A1.
Author information
Authors and Affiliations
Contributions
All authors have given approval to the final version of this manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chamberlain, C.A., Hatch, M. & Garrett, T.J. Oxalobacter formigenes produces metabolites and lipids undetectable in oxalotrophic Bifidobacterium animalis. Metabolomics 16, 122 (2020). https://doi.org/10.1007/s11306-020-01747-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01747-2