Abstract
The objective of this paper is to explore the function of the AOL-s00215g415 (Aog415) gene, which encodes for the synthesis of siderophore in the nematode trapping fungal model strain A. oligospora, in order to understand the relationship between siderophore biosynthesis and nematode trapping activity. After a through sequence analysis, it was determined that Aog415 is a siderophore-synthesizing NRPS. The product of this gene was then identified to be the hydroxamate siderophore desferriferrichrome, using mass spectrometry analysis. When compared to the WT strains, the Aog415 knockout strain exhibited a 60% decrease in siderophore content in fermentation broth. Additionally, the number of predatory rings of decreased by 23.21%, while the spore yield increased by 37.34%. The deletion of Aog415 did not affect the growth of A. oligospora in diverse nutrient medium. Lipid metabolism-related pathways were the primary targets of Aog415 disruption as revealed by the metabolomic analysis. In comparison to the WT, a significant reduction in the levels of glycerophospholipids, and glycolipids was observed in the mutation. The metabolic alteration in fatty acyls and amino acid-like molecules were significantly disrupted. The knockout of Aog415 impaired the biosynthesis of the hydroxamate siderophore desferriferrichrome, remodeled the flow of fatty acid in A. oligospora, and mainly reprogrammed the membrane lipid metabolism in cells. Desferriferrichrome, a hydroxamate siderophore affects the growth, metabolism and nematode trapping ability of A. oligospora by regulating iron intake and cell membrane homeostasis. Our study uncovered the significant contribution of siderophores to the growth and nematode trapping ability and constructed the relationship among siderophores biosynthesis, lipid metabolism and nematode trapping activity of A. oligospora, which provides a new insight for the development of nematode biocontrol agents based on nematode trapping fungi.
Graphical abstract







Similar content being viewed by others
Data availability
All supporting dataand protocols have been provided within the article or through supplementary data files. Supplementary material is available with the online version of this article.
References
Ahmed E, Holmström SJ (2014) Siderophores in environmental research: roles and applications. Microb Biotechnol 7(3):196–208. https://doi.org/10.1111/1751-7915.12117
Ambrosi C, Leoni L, Putignani L, Orsi N, Visca P (2000) Pseudobactin biogenesis in the plant growth-promoting rhizobacterium Pseudomonas strain B10: identification and functional analysis of the L-ornithine N(5)-oxygenase (psbA) gene. J Bacteriol 182(21):6233–6238. https://doi.org/10.1128/jb.182.21.6233-6238.2000
Amsaveni R, Sureshkumar M, Aravinth A, Reshma Mary J, Vivekanandhan G (2016) Production of Non-Ribosomal Peptide Synthetase (NRPS)-dependent siderophore by aeromonas isolates. Iran Biomed J 20(4):235–240. https://doi.org/10.7508/ibj.2016.04.008
Bai N, Zhang G, Wang W, Feng H, Yang X, Zheng Y et al (2022) Ric8 acts as a regulator of G-protein signalling required for nematode-trapping lifecycle of Arthrobotrys oligospora. Environ Microbiol 24(4):1714–1730. https://doi.org/10.1111/1462-2920.15735
Barry SM, Challis GL (2009) Recent advances in siderophore biosynthesis. Curr Opin Chem Biol 13(2):205–215. https://doi.org/10.1016/j.cbpa.2009.03.008
Bhat AA, Shakeel A, Waqar S, Handoo ZA, Khan AA (2023) Microbes vs. nematodes: insights into biocontrol through antagonistic organisms to control root-knot nematodes. Plants (basel). https://doi.org/10.3390/plants12030451
Chalut C (2016) MmpL transporter-mediated export of cell-wall associated lipids and siderophores in mycobacteria. Tuberculosis (edinb) 100:32–45. https://doi.org/10.1016/j.tube.2016.06.004
Dutta S, Yu SM, Jeong SC, Lee YH (2020) High-throughput analysis of genes involved in biocontrol performance of Pseudomonas fluorescens NBC275 against Gray mold. J Appl Microbiol 128(1):265–279. https://doi.org/10.1111/jam.14475
Esmaeel Q, Pupin M, Jacques P, Leclère V (2018) Nonribosomal peptides and polyketides of Burkholderia: new compounds potentially implicated in biocontrol and pharmaceuticals. Environ Sci Pollut Res Int 25(30):29794–29807. https://doi.org/10.1007/s11356-017-9166-3
Fan Y, Zhang W, Chen Y, Xiang M, Liu X (2021) DdaSTE12 is involved in trap formation, ring inflation, conidiation, and vegetative growth in the nematode-trapping fungus Drechslerella dactyloides. Appl Microbiol Biotechnol 105(19):7379–7393. https://doi.org/10.1007/s00253-021-11455-z
Haas H (2003) Molecular genetics of fungal siderophore biosynthesis and uptake: the role of siderophores in iron uptake and storage. Appl Microbiol Biotechnol 62(4):316–330. https://doi.org/10.1007/s00253-003-1335-2
Hider RC, Kong X (2010) Chemistry and biology of siderophores. Nat Prod Rep 27(5):637–657. https://doi.org/10.1039/b906679a
Himpsl SD, Mobley HLT (2019) Siderophore detection using chrome azurol s and cross-feeding assays. Methods Mol Biol 2021:97–108. https://doi.org/10.1007/978-1-4939-9601-8_10
Jiang X, Xiang M, Liu X (2017) Nematode-Trapping Fungi. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.FUNK-0022-2016
Jiang KX, Liu QQ, Bai N, Zhu MC, Zhang KQ, Yang JK (2022) AoSsk1, a response regulator required for mycelial growth and development, stress responses, trap formation, and the secondary metabolism in arthrobotrys oligospora. J Fungi (basel). https://doi.org/10.3390/jof8030260
Johnson LJ, Koulman A, Christensen M, Lane GA, Fraser K, Forester N et al (2013) An extracellular siderophore is required to maintain the mutualistic interaction of Epichloë festucae with Lolium perenne. PLoS Pathog 9(5):e1003332. https://doi.org/10.1371/journal.ppat.1003332
Kautsar SA, Blin K, Shaw S, Navarro-Muñoz JC, Terlouw BR, van der Hooft JJJ et al (2020) MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res 48(D1):D454-d458. https://doi.org/10.1093/nar/gkz882
Khasheii B, Mahmoodi P, Mohammadzadeh A (2021) Siderophores: Importance in bacterial pathogenesis and applications in medicine and industry. Microbiol Res 250:126790. https://doi.org/10.1016/j.micres.2021.126790
Kramer J, Özkaya Ö, Kümmerli R (2020) Bacterial siderophores in community and host interactions. Nat Rev Microbiol 18(3):152–163. https://doi.org/10.1038/s41579-019-0284-4
Kriegl L, Havlicek V, Dichtl K, Egger M, Hoenigl M (2022) Siderophores: a potential role as a diagnostic for invasive fungal disease. Curr Opin Infect Dis 35(6):485–492. https://doi.org/10.1097/qco.0000000000000862
Kuo TH, Yang CT, Chang HY, Hsueh YP, Hsu CC (2020) Nematode-trapping fungi produce diverse metabolites during predator-prey interaction. Metabolites. https://doi.org/10.3390/metabo10030117
Lee SR, Schalk F, Schwitalla JW, Guo H, Yu JS, Song M et al (2022) GNPS-guided discovery of madurastatin siderophores from the termite-associated Actinomadura sp. RB99. Chemistry. https://doi.org/10.1002/chem.202200612
Liu Q, Li D, Jiang K, Zhang KQ, Yang J (2022) AoPEX1 and AoPEX6 are required for mycelial growth, conidiation, stress response, fatty acid utilization, and trap formation in arthrobotrys oligospora. Microbiol Spectr 10(2):e0027522. https://doi.org/10.1128/spectrum.00275-22
Lu H, Wang S, Gu T, Sun L, Wang Y (2023) Roles of the fungal-specific lysine biosynthetic pathway in the nematode-trapping fungus Arthrobotrys oligospora identified through metabolomics analyses. J Fungi (basel). https://doi.org/10.3390/jof9020206
Medema MH, Blin K, Cimermancic P, de Jager V, Zakrzewski P, Fischbach MA et al (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. https://doi.org/10.1093/nar/gkr466
Montarry J, Mimee B, Danchin EGJ, Koutsovoulos GD, Ste-Croix DT, Grenier E (2021) Recent advances in population genomics of plant-parasitic nematodes. Phytopathology 111(1):40–48. https://doi.org/10.1094/phyto-09-20-0418-rvw
Peng H, Dong X, Lu H, Kong X, Zha X, Wang Y (2022) A putative F-box-domain-encoding gene AOL_s00076g207 regulates the development and pathogenicity of Arthrobotrys oligospora. J Basic Microbiol 62(1):74–81. https://doi.org/10.1002/jobm.202100388
Philpott CC (2006) Iron uptake in fungi: a system for every source. Biochim Biophys Acta 1763(7):636–645. https://doi.org/10.1016/j.bbamcr.2006.05.008
Pluháček T, Lemr K, Ghosh D, Milde D, Novák J, Havlíček V (2016) Characterization of microbial siderophores by mass spectrometry. Mass Spectrom Rev 35(1):35–47. https://doi.org/10.1002/mas.21461
Ronnebaum TA, Lamb AL (2018) Nonribosomal peptides for iron acquisition: pyochelin biosynthesis as a case study. Curr Opin Struct Biol 53:1–11. https://doi.org/10.1016/j.sbi.2018.01.015
Saha M, Sarkar S, Sarkar B, Sharma BK, Bhattacharjee S, Tribedi P (2016) Microbial siderophores and their potential applications: a review. Environ Sci Pollut Res Int 23(5):3984–3999. https://doi.org/10.1007/s11356-015-4294-0
Solanki MK, Singh RK, Srivastava S, Kumar S, Kashyap PL, Srivastava AK et al (2014) Isolation and characterization of siderophore producing antagonistic rhizobacteria against Rhizoctonia solani. J Basic Microbiol 54(6):585–597. https://doi.org/10.1002/jobm.201200564
Soliman MS, El-Deriny MM, Ibrahim DSS, Zakaria H, Ahmed Y (2021) Suppression of root-knot nematode Meloidogyne incognita on tomato plants using the nematode trapping fungus Arthrobotrys oligospora Fresenius. J Appl Microbiol 131(5):2402–2415. https://doi.org/10.1111/jam.15101
Song TY, Xu ZF, Chen YH, Ding QY, Sun YR, Miao Y et al (2017) Potent nematicidal activity and new hybrid metabolite production by disruption of a cytochrome P450 gene involved in the biosynthesis of morphological regulatory arthrosporols in nematode-trapping fungus Arthrobotrys oligospora. J Agric Food Chem 65(20):4111–4120. https://doi.org/10.1021/acs.jafc.7b01290
Swayambhu G, Bruno M, Gulick AM, Pfeifer BA (2021) Siderophore natural products as pharmaceutical agents. Curr Opin Biotechnol 69:242–251. https://doi.org/10.1016/j.copbio.2021.01.021
Teng LL, Song TY, Chen YH, Chen YG, Zhang KQ, Li SH et al (2020) Novel polyketide-terpenoid hybrid metabolites from a potent nematicidal Arthrobotrys oligospora Mutant ΔAOL_s00215g278. J Agric Food Chem 68(41):11449–11458. https://doi.org/10.1021/acs.jafc.0c04713
Wang Y, Zhang G, Huang Y, Guo M, Song J, Zhang T et al (2022) a potential biofertilizer-siderophilic bacteria isolated from the rhizosphere of Paris polyphylla var. yunnanensis. Front Microbiol. https://doi.org/10.3389/fmicb.2022.870413
Wang D, Ma N, Rao W, Zhang Y (2023) Recent advances in life history transition with nematode-trapping fungus Arthrobotrys oligospora and its application in sustainable agriculture. Pathogens. https://doi.org/10.3390/pathogens12030367
Wei LX, Zhang HX, Tan JL, Chu YS, Li N, Xue HX et al (2011) Arthrobotrisins A-C, oligosporons from the nematode-trapping fungus Arthrobotrys oligospora. J Nat Prod 74(6):1526–1530. https://doi.org/10.1021/np200187z
Xie M, Ma N, Bai N, Zhu M, Zhang KQ, Yang J (2022) Phospholipase C (AoPLC2) regulates mycelial development, trap morphogenesis, and pathogenicity of the nematode-trapping fungus Arthrobotrys oligospora. J Appl Microbiol 132(3):2144–2156. https://doi.org/10.1111/jam.15370
Xu G, Martinez JS, Groves JT, Butler A (2002) Membrane affinity of the amphiphilic marinobactin siderophores. J Am Chem Soc 124(45):13408–13415. https://doi.org/10.1021/ja026768w
Yang J, Wang L, Ji X, Feng Y, Li X, Zou C et al (2011) Genomic and proteomic analyses of the fungus Arthrobotrys oligospora provide insights into nematode-trap formation. PLoS Pathog 7(9):e1002179. https://doi.org/10.1371/journal.ppat.1002179
Zhang Y, Li S, Li H, Wang R, Zhang KQ, Xu J (2020) Fungi-nematode interactions: diversity, ecology, and biocontrol prospects in agriculture. J Fungi (basel). https://doi.org/10.3390/jof6040206
Zhang S, Chen Y, Zhu J, Lu Q, Cryle MJ, Zhang Y et al (2023) Structural diversity, biosynthesis, and biological functions of lipopeptides from Streptomyces. Nat Prod Rep 40(3):557–594. https://doi.org/10.1039/d2np00044j
Zhao X, Fan Y, Zhang W, Xiang M, Kang S, Wang S et al (2023) DhFIG_2, a gene of nematode-trapping fungus Dactylellina haptotyla that encodes a component of the low-affinity calcium uptake system, is required for conidiation and knob-trap formation. Fungal Genet Biol 166:103782. https://doi.org/10.1016/j.fgb.2023.103782
Zhou D, Zhu Y, Bai N, Xie M, Zhang KQ, Yang J (2021) Aolatg1 and Aolatg13 regulate autophagy and play different roles in conidiation, trap formation, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora. Front Cell Infect Microbiol 11:824407. https://doi.org/10.3389/fcimb.2021.824407
Zhou J, Wu QF, Li SH, Yan JX, Wu L, Cheng QY et al (2022) The Multifaceted Gene 275 Embedded in the PKS-PTS gene cluster was involved in the regulation of arthrobotrisin biosynthesis, TCA cycle, and septa formation in nematode-trapping fungus Arthrobotrys oligospora. J Fungi (basel). https://doi.org/10.3390/jof8121261
Zhu MC, Zhao N, Liu YK, Li XM, Zhen ZY, Zheng YQ et al (2022) The cAMP-PKA signalling pathway regulates hyphal growth, conidiation, trap morphogenesis, stress tolerance, and autophagy in Arthrobotrys oligospora. Environ Microbiol 24(12):6524–6538. https://doi.org/10.1111/1462-2920.16253
Acknowledgements
This research was funded by the National Natural Science Foundation of China (No. 32200042, 31770066), the Anhui Provincial Natural Science Foundation (No. 2208085QC67), the Natural Science Foundation of Anhui Higher Education Institutions of China (No. KJ2021A0058).
Funding
This article was funded by National Natural Science Foundation of China, no.31770066, no. 32200042, Anhui Provincial Natural Science Foundation,no. 2208085QC67, Natural Science Foundation of Anhui Higher Education Institutions of China, no. KJ2021A0058.
Author information
Authors and Affiliations
Contributions
Huiwen Liu, performed the experiments, supported metabolomics analysis, writing (review and editing), supported product identification. Liangyin Sun: performed the experiments, writing. Jintao Zhang supported product identification and manuscript revision, Yongzhong Wang: funding acquisition, supervision, and writing (review and editing). Hengqian Lu: designed the project, investigation, supported metabolomics analysis and data interpretation, writing (review and editing), funding acquisition. All authors discussed the results from the experiments and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11274_2023_3840_MOESM1_ESM.docx
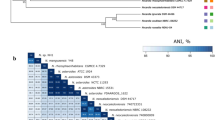
Supplementary material 1 (SM1) (WORD): Supplementary Table and FiguresTable S1. Primer sequences used in this study.Figure S1. The phylogenetic tree analysis of Aog415 in A. oligospora.Figure S2. Deletion of Aog415 in A. oligospora.Figure S3. The mirror match showed the matched MS/MS spectrum peak of desferriferrichrome.
11274_2023_3840_MOESM2_ESM.xlsx
Supplementary material 2 (SM2) (EXCEL): Data matrix of metabolomics analysis
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, H., Sun, L., Zhang, J. et al. Siderophore-synthesizing NRPS reprogram lipid metabolic profiles for phenotype and function changes of Arthrobotrys oligospora. World J Microbiol Biotechnol 40, 46 (2024). https://doi.org/10.1007/s11274-023-03840-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03840-9