Abstract
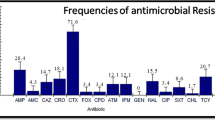
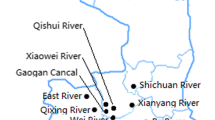
The present study aims to demonstrate the β-lactam resistance phenotypes and genotypes of Escherichia coli isolates from the Fu River in Chengdu, southwestern China. We obtained 108 E. coli isolates from nine sampling sites during May and December 2010. The total bacterial count varied from 79 colony-forming units (CFU)/ml to 14,550 CFU/ml, and coliform group number from 13 to 1,435 MPN/ml. Among the 108 isolates, 0–31.48% were resistant to β-lactams and β-lactam inhibitors, 1.85–7.40% to aminoglycoside, 1–20% to fluoroquinolone, and 50% to trimethoprim- sulfamethoxazole. The total bacterial count and antimicrobial resistance of different sites were significantly correlated (P < 0.05). Among the 34 ampicillin-resistant isolates, polymerase chain reaction (PCR) amplification and DNA sequencing showed that bla TEM, bla SHV, and bla CTX-M were detected in 85.29% (n = 29), 41.18% (n = 14), and 5.88% (n = 2) of the isolates, respectively, whereas bla KPC and bla GES were not observed in any of the isolates. Enterobacterial repetitive intergenic consensus-PCR patterns revealed that the 34 ampicillin-resistant E. coli isolates belonged to three distinct groups. Plasmid DNAs from the 14 SHV producer isolates yielded one to five bands of ca. 0.15–40 kb. To our knowledge, the current study is the first to describe the phenotypic and genetic characterizations of β-lactam resistance in E. coli isolates of river water origin from the Fu River, Chengdu, southwestern China. Results of the present study suggest that the river water may be considered as a reservoir for antibiotic resistance genes.


Similar content being viewed by others
References
American Public Health Association (APHA) The American Water Works Association (AWWA), and the Water Environment Federation (WEF) (1998) Standard methods for the examination of water and wastewater, 20th edn. Washington
Baquero F, Jose-Luis M, Canton R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotech 19:260–265
Biyela PT, Lin J, Bezuidenhou CC (2004) The role of aquatic ecosystems as reservoirs of antibiotic resistant bacteria and antibiotic resistance genes. Water Sci Technol 50:45–50
Castiglioni S, Pomati F, Miller K, Burns BP, Zuccato E, Calamari D, Neilan BA (2008) Novel homologs of the multiple resistance regulator marA in antibiotic-contaminated environments. Water Res 42:4271–4280
Chen H, Shu W, Chang X, Chen Ji-an, Guo Y, Tan Y (2010) The profile of antibiotics resistance and integrons of extended-spectrum beta-lactamase producing thermo tolerant coliforms isolated from the Yangtze River basin in Chongqing. Environ Pollut 158:2459–2464
Clinical and Laboratory Standards Institute (2009) Performance standards for antimicrobial susceptibility testing: 19th informational supplement M100–S19. CLSI, Wayne
D’Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Science 311:374–377
Davies JE (1997) Origins, acquisition and dissemination of antibiotic resistance determinants. Ciba Found Symp 207:15–27
Decré D, Burghoffer B, Gautier V, Petit JC, Arlet G (2004) Outbreak of multi-resistant Klebsiella oxytoca involving strains with extended-spectrum β-lactamases and strains with extended-spectrum activity of the chromosomal β-lactamase. J Antimicrob Chemoth 54:881–888
Edelstein M, Pimkin M, Palagin I, Edelstein I, Stratchounski L (2003) Prevalence and molecular epidemiology of CTX-M extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae in Russian hospitals. Antimicrob Agents Chemoth 47:3724–3732
GB/T 5750-2006, Standard examination methods for drinking water of China
Guo-Bao T, Hong-Ning W, Li-Kou Z, Jun-Ni T, Ying-Wang Z, Man-Yu Y, Jing-Yuan T, Yi Z, An-Yun Z, Xin Y, Chang-Wen X, Yue-Jun F (2009) Detection of CTX-M-15, CTX-M-22, and SHV-2 extended-spectrum β-lactamases (ESBLs) in Escherichia coli fecal-sample isolates from pig farms in China. Foodborne Pathog Dis 6:297–304
Harakeh S, Yassine H, El-Fadel M (2006) Antimicrobial-resistant patterns of Escherichia coli and Salmonella strains in the aquatic Lebanese environments. Environ Pollut 143:269–277
Henriques IS, Fonseca F, Alves A, Saavedra MJ, Correia A (2006) Occurrence and diversity of integrons and β-lactamase genes among ampicillin-resistant isolates from estuarine waters. Res Microbiol 157:938–947
Heritage J, MZali FH, Gascoyne-Binzi D, Hawke PM (1999) Evolution and spread of SHV extended-spectrum beta-lactamases in gram-negative. J Antimicrob Chemoth 44:309–318
Hoffmann H, Roggenkamp A (2003) Population genetics of the nomenspecies Enterobacter cloacae. Appl Environ Microb 69:5306–5318
Hossain A, Ferraro MJ, Pino RM, Dew RB III, Moland ES, Lockhart TJ, Thomson KS, Goering RV, Hanson ND (2004) Plasmid-mediated carbapenem-hydrolyzing enzyme KPC-2 in an Enterobacter sp. Antimicrob Agents Chemoth 48:4438–4440
Jeong SH, Bae W, Kim D (2005) First outbreak of Klebsiella pneumoniae clinical isolates producing GES-5 and SHV-12 extended-spectrum β-lactamases in Korea. Antimicrob Agents Chemoth 49:4809–4810
Kampfer P, Nienhuser A, Packroff G, Wernicke F, Mehlinge A, Nixdorf K, Fiedler S, Kolauch C, Esser M (2008) Molecular identification of coliform bacteria isolated from drinking water reservoirs with traditional methods and the Colilert-18 system. Int J Hy Environ Heal 211:374–384
Kemper N (2008) Veterinary antibiotics in the aquatic and terrestrial environment. Ecol Indic 8:1–13
Kiratisin P, Apisarnthanarak A, Laesripa C, Saifon P (2008) Molecular characterization and epidemiology of extended-spectrum-β-lactamase-producing Escherichia coli and Klebsiella pneumonia isolates causing health care-associated infection in Thailand, where the CTX-M family is endemic. Antimicrob Agents Chemoth 52:2818–2824
Le TX, Munekage Y, Kato S-I (2005) Antibiotic resistance in bacteria from shrimp farming in mangrove areas. Sci Total Environ 349:95–105
Li D, Yang M, Hu J, Zhang J, Liu R, Gu X, Zhang Y, Wang Z (2009) Antibiotic-resistance profile in environmental bacteria isolated from penicillin production wastewater treatment plant and the receiving river. Environ Microbiol 11:1506–1517
Li D, Yu T, Zhang Y, Yang M, Li A, Liu M, Qi R (2010) Antibiotic resistance characteristics of environmental bacteria from an oxytetracycline production wastewater treatment plant and the receiving river. Appl Environ Microbiol 76:3444–3451
Li-Kou Z, Hong-Ning W, Bo Z, Jin-Niang L, Xu-Ting L, Xu-Ting L, Guo-Bao T, Kun W, Ying-Shun Z, Chang-Wen Xu, Zhi-Rong Y (2011) Phenotypic and genotypic characterization of beta-lactam resistance in Klebsiella pneumoniae isolated from swine. Vet Microbiol 149:139–146
Livermore DM (1995) Beta-lactamases in laboratory and clinical resistance. Clin Microbiol Rev 8:557–584
Luo Y, Mao D, Michal RYSZ, Zhou Q, Zhang H, Zhang H, Xu L, Alvarez PJJ (2010) Trends in antibiotic resistance genes occurrence in the Haihe River, China. Environ Sci Technol 44:7220–7225
Maheux AF, Picard FJ, Boissinot M, Bissonnette L, Paradis S, Bergeron MG (2009) Analytical comparison of nine PCR primer sets designed to detect the presence of Escherichia coli/Shigella in water samples. Water Res 43:3019–3028
Martinez JL (2009) Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ Pollut 157:2893–2902
Meacham KJ, Zhang L, Foxman B, Bauer RJ, Marrs CF (2003) Evaluation of genotyping large numbers of Escherichia coli Isolates by enterobacterial repetitive intergenic consensus-PCR. J Clin Microbiol 41:5224–5226
Mendonça N, Ferreira E (2009) Genetic diversity of genes encoding OKP and LEN β-lactamases produced by clinical Klebsiella pneumoniae strains in Portugal. Diagn Microbiol Infec Dis 63:334–338
Moore JE, Moore PJA, Millar BC, Goldsmith CE, Loughrey A, Rooney PJ, Rao JR (2010) The presence of antibiotic resistant bacteria along the river Lagan. Agric Water Manage 98:217–221
Munoz-Aguayo J, Lang KS, LaPara TM, Gonzalez G, Singer RS (2007) Evaluating the effects of chlortetracycline on the proliferation of antibiotic-resistant bacteria in a simulated river water ecosystem. Appl Environ Microbiol 73:5421–5425
Pei R, Kim SC, Carlson KH, Pruden A (2006) Effect of river landscape on the sediment concentrations of antibiotics and corresponding antibiotic resistance genes (ARG). Water Res 40:2427–2435
Poglazova MN, Mitskevich IN, Kuzhinovsky VA (1996) A spectrofluorimetric method for the determination bacterial counts in environmental samples. J Microbiol Meth 24:211–218
Ram S, Vajpayee P, Shanker S (2007) Prevalence of multi-antimicrobial-agent resistant, shiga toxin and enterotoxin producing Escherichia coli in surface waters of river Ganga. Environ Sci Technol 41:7383–7388
Rompre A, Servais P, Baudart J, de-Roubin M-R, Laurent P (2002) Detection and enumeration of coliforms in drinking water: current methods and emerging approaches. J Microbiol Meth 49:31–54
Sarmah AJ, Meyer MT, Boxall ABA (2006) A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere 65:725–759
Schwartz T, Kohnen W, Jansen B, Obst U (2003) Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol Ecol 43:325–335
Servais P, Passerat J (2009) Antimicrobial resistance of fecal bacteria in waters of the Seine river watershed (France). Sci Total Environ 408:365–372
Sharma A, Singh SK, Kori L (2009) Molecular epidemiological characteristics of Shigella spp. isolated from river Narmada during 2005–2006. J Environ Heal 71:61–66
Tao R, Ying G–G, Su H-C, Zhou H-W, Sidhu JPS (2010) Detection of antibiotic resistance and tetracycline resistance genes in Enterobacteriaceae isolated from the Pearl Rivers in South China. Environ Pollut 158:2101–2109
Threlfall EJ (2002) Antimicrobial drug resistance in Salmonella: problems and perspectives in food-and water-borne infections. FEMS Microbiol Rev 26:141–148
Watkinson AJ, Micalizzi GB, Graham GM, Bates JB, Costanzo SD (2007) Antibiotic-resistant Escherichia coli in wastewaters, surface waters, and oysters from an urban riverine system. Antimicrob Agents Chemoth 73:5667–5670
Xian-Zhi L, Mehrotra M, Ghimire S, Adewoye L (2007) β-Lactam resistance and β-lactamases in bacteria of animal origin. Vet Microbiol 121:197–214
Yagi T, Kurokawa H, Shibata N, Shibayama K, Arakawa Y (2000) A preliminary survey of extended-spectrum β-lactamases (ESBLs) in clinical isolates of Klebsiella pneumoniae and Escherichia coli in Japan. FEMS Microbiol Lett 184:53–56
Acknowledgments
This work was supported by research projects 10ZA055 of the key program of the Education Bureau of Sichuan province.
Author information
Authors and Affiliations
Corresponding author
Additional information
Li-Wen Li, contribute equally to the article.
Rights and permissions
About this article
Cite this article
Zou, LK., Li, LW., Pan, X. et al. Molecular characterization of β-lactam-resistant Escherichia coli isolated from Fu River, China. World J Microbiol Biotechnol 28, 1891–1899 (2012). https://doi.org/10.1007/s11274-011-0987-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-011-0987-9