Abstract
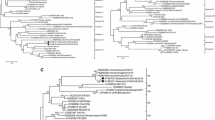
Fifty-seven Gallid alphaherpesvirus 2 (GaHV-2) isolates, collected during a 30-year period (1990–2019) from commercial poultry flocks affected by Marek’s disease (MD), were molecularly characterised. The GaHV-2 meq gene was amplified and sequenced to evaluate the virus virulence, based on the number of PPPPs within the proline-rich repeats (PRRs) of its transactivation domain. The present illustration of virus virulence evaluation on a large scale of field virus isolates by molecular analysis exemplifies the practical benefit and usefulness of the molecular marker in commercial GaVH-2 isolates. The alternative assay of GaVH-2 virulence pathotyping is the classical Gold Standard ADOL method, which is difficult and impossible to employ on a large scale using the Specific Pathogen Free (SPF) chicks of the ADOL strains kept in isolators for two months. The phylogenetic analysis performed in the present study showed that the meq gene amino acid sequences of the 57 Israeli strains divide into 16 phylogenetic branches. The virulence evaluation was performed in comparison with 36 GaHV-2 prototype strains, previously characterised by the in vivo Gold Standard ADOL assay. The results obtained revealed that the GaHV-2 strains circulating in Israel have evolved into a higher virulence potential during the years, as the four-proline stretches number in the meq gene decreased over the investigated period, typically of very virulent virus prototypes. The present study supports the meq gene molecular markers for the assessment of field GaVH-2 strains virulence.

Similar content being viewed by others
Data Availability
Not applicable.
References
Gimeno IM (2014) Marek’s disease and differential diagnosis with other tumor viral diseases of poultry. Encyclopedia of agriculture and food systems. Elsevier, Amsterdam, pp 156-171. https://doi.org/10.1016/B978-0-444-52512-3.00193-5
Gimeno IM, Schat KA (2018) Virus-induced immunosuppression in chickens. Avian Dis 62:272–285. https://doi.org/10.1637/11841-041318-Review
Schat KA, Nair V (2013) Neoplastic diseases: Marek’s disease. In: Swayne DE, Glisson JR, McDougald LR, Nolan LK, Suarez DL, Nair VL (eds) Diseases of poultry, 13th edn. Wiley-Blackwell Hoboken, Ames, IA, pp 515–552
Gatherer D, Depledge DP, Hartley CA, Szpara ML, Vaz PK, Benkő M, Brandt CR, Bryant NA, Dastjerdi A, Doszpoly A, Gompels UA, Inoue N, Jarosinski KW, Kaul R, Lacoste V, Norberg P, Origgi FC, Orton RJ, Pellett PE, Schmid DS, Spatz SJ, Stewart JP, Trimpert J, Waltzek TB, Davison AJ (2021) ICTV virus taxonomy profile: Herpesviridae 2021. J Gen Virol 102(10):1–2. https://doi.org/10.1099/JGV.0.001673
Nair V (2018) Spotlight on avian pathology: Marek’s disease. Avian Pathol 47:440–442. https://doi.org/10.1080/03079457.2018.1484073
Trimpert J, Groenke N, Jenckel M, He S, Kunec D, Szpara ML, Spatz SJ, Osterrieder N, McMahon DP (2017) A phylogenomic analysis of Marek’s disease virus reveals independent paths to virulence in Eurasia and North America. Evol Appl 10:1091–1101. https://doi.org/10.1111/eva.12515
Witter RL (1997) Increased virulence of Marek’s disease virus field isolates. Avian Dis 41:149. https://doi.org/10.2307/1592455
Witter RL, Calnek BW, Buscaglia C, Gimeno IM, Schat KA (2005) Classification of Marek’s disease viruses according to pathotype: philosophy and methodology. Avian Pathol 34:75–90. https://doi.org/10.1080/03079450500059255
Davidson I, Natour-Altory A, Raibstein I, Kin E, Dahan Y, Krispin H, Elkin N (2018) Monitoring the uptake of live avian vaccines by their detection in feathers. Vaccine 36:637–643. https://doi.org/10.1016/j.vaccine.2017.12.052
Davidson I, Natour-Altory N, Shimshon Y (2018) Evaluation of live vaccine application in commercial poultry flocks using feathers—in practice. Israel J Vet Med 73:8–13
Davidson I (2020) Out of sight but not out of mind aspects of the oncogenic herpesvirus, Marek’s Disease virus. Animals (Basel) 10(8):1319. https://doi.org/10.3390/ani10081319
Schat KA, van Santen VL (2013) Chicken infectious anemia. In: Swayne DE, Glisson JR, McDougald LR, Nolan LK, Suarez DL, Nair VL (eds) Diseases of poultry, 13th edn. Wiley-Blackwell Hoboken, Ames, IA, pp 248–275
Liu JL, Lin SF, Xia L, Brunovskis P, Li D, Davidson I, Lee LF, Kung HJ (1999) Meq and V-IL8: cellular genes in disguise? Acta Virol 43:94–101
Qian Z, Brunovskis P, Rauscher F 3rd, Lee L, Kung HJ (1995) Transactivation activity of Meq, a Marek’s disease herpesvirus bZIP protein persistently expressed in latently infected transformed T cells. J Virol 69:4037-4044.
Ross NLJ (1999) T-cell transformation by Marek’s disease virus. Trends Microbiol 7:22–29. https://doi.org/10.1016/S0966-842X(98)01427-9
Shamblin CE, Greene N, Arumugaswami V, Dienglewicz RL, Parcells MS (2004) Comparative analysis of Marek’s disease virus (MDV) glycoprotein-, lytic antigen pp38- and transformation antigen Meq-encoding genes: association of meq mutations with MDVs of high virulence. Vet Microbiol 102:147–167. https://doi.org/10.1016/j.vetmic.2004.06.007
Duffy S, Shackelton LA, Holmes EC (2008) Rates of evolutionary change in viruses: patterns and determinants. Nat Rev Genet 9:267–276. https://doi.org/10.1038/nrg2323
Firth C, Kitchen A, Shapiro B, Suchard MA, Holmes EC, Rambaut A (2010) Using time-structured data to estimate evolutionary rates of double-stranded DNA viruses. Mol Biol Evol 27:2038–2051. https://doi.org/10.1093/molbev/msq088
Padhi A, Parcells MS (2016) Positive selection drives rapid evolution of the meq oncogene of Marek’s disease virus. PLoS One 11(9):e0162180. https://doi.org/10.1371/journal.pone.0162180
Renz KG, Cooke J, Clarke N, Cheetham BF, Hussain Z, Fakhrul Islam AFM, Tannock GA, Walkden-Brown SW (2012) Pathotyping of Australian isolates of Marek’s disease virus and association of pathogenicity with meq gene polymorphism. Avian Pathol 41:161–176. https://doi.org/10.1080/03079457.2012.656077
Dudnikova E, Norkina S, Vlasov A, Slobodchuk A, Lee LF, Witter RL (2007) Evaluation of Marek’s disease field isolates by the “best fit” pathotyping assay. Avian Pathol 36:135–143. https://doi.org/10.1080/03079450701209857
Dunn JR, Black Pyrkosz A, Steep A, Cheng HH (2019) Identification of Marek’s disease virus genes associated with virulence of US strains. J Gen Virol 100:1132–1139. https://doi.org/10.1099/jgv.0.001288
Mescolini G, Lupini C, Felice V, Guerrini A, Silveira F, Cecchinato M, Catelli E (2019) Molecular characterization of the meq gene of Marek’s disease viruses detected in unvaccinated backyard chickens reveals the circulation of low- and high-virulence strains. Poult Sci 98:3130–3137. https://doi.org/10.3382/ps/pez095
Mescolini G, Lupini C, Davidson I, Massi P, Tosi G, Fiorentini L, Catelli E (2020) Molecular characterization of a Marek’s disease virus strain detected in tumour-bearing turkeys. Avian Pathol 2:202–207. https://doi.org/10.1080/03079457.2019.1691715
Mescolini G, Lupini C, Davidson I, Massi P, Tosi G, Catelli E (2020) Marek’s disease viruses circulating in commercial poultry in Italy in the years 2015–2018 are closely related by their meq gene phylogeny. Transbound Emerg Dis 67:98–107. https://doi.org/10.1111/tbed.13327
Davidson I, Weisman Y, Orgad U, Jacobson B, Perl S, Strenger C, Becker Y, Malkinson M (1988) Pathogenicity studies of Marek’s disease virus isolates in Israel. Israel J Vet Sci 44:223–232
Davidson I, Borenshtain R (1999) Multiple infection of chickens and turkeys with avian oncogenic viruses: prevalence and molecular analysis. Acta Virol 43:136–142
Davidson I (2007) Avian oncogenic viruses: the correlation between clinical signs and molecular virus identification, knowledge acquired from the examination of over 1000 flocks. Isr Vet Med J 62:42–47
Hassanin O, Abdallah F, El-Araby IE (2013) Molecular characterization and phylogenetic analysis of Marek’s disease virus from clinical cases of Marek’s disease in Egypt. Avian Dis 57:555–561. https://doi.org/10.1637/10337-082912-Reg.1
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Yu Z-H, Man T, Lou J, Wang X-W, Ding K, Yu L-L, Su J-W, Chi J-Q, Zhao P, Hu B, Zhang G-P, Liu J-X (2013) Molecular characteristics and evolutionary analysis of field Marek’s disease virus prevalent in vaccinated chicken flocks in recent years in China. Virus Genes 47:282–291. https://doi.org/10.1007/s11262-013-0942-y
Suresh P, Johnson Rajeswar J, Sukumar K, Harikrishnan TJ, Srinivasan P (2017) Complete nucleotide sequence analysis of the oncogene “Meq” from serotype 1 Marek’s disease virus isolates from India. Br Poult Sci 58:111–115. https://doi.org/10.1080/00071668.2016.1257780
Abdallah F, Hassanin O, Attar E, Ali H, Megahed M, Nair V (2018) Marek’s disease virus in Egypt: historical overview and current research based on the major MDV-encoded oncogene meq. Hosts Virus 5:35–43. https://doi.org/10.17582/journal.hv/2018/5.3.35.43
Woźniakowski G, Samorek-Salamonowicz E (2014) Molecular evolution of Marek’s Disease Virus (MDV) field strains in a 40-year time period. Avian Dis 58:550–557. https://doi.org/10.1637/10812-030614-Reg.1
Abd-Ellatieff H, Abou Rawash AA, Ellakany HF, Goda WM, Suzuki T, Yanai T (2018) Molecular characterization and phylogenetic analysis of a virulent Marek’s disease virus field strain in broiler chickens in Japan. Avian Pathol 47:47–57. https://doi.org/10.1080/03079457.2017.1362497
Ghalyanchilangeroudi A, Hossein H, Hadi HN, Aidin M, Omid D, Morshed R (2022) Molecular characterization and phylogenetic analysis of Marek’s disease virus in Iran. Avian Dis 66(3):1–5. https://doi.org/10.1637/aviandiseases-D-22-00018
Read AF, Baigent SJ, Powers C, Kgosana LB, Blackwell L, Smith LP, Kennedy DA, Walkden-Brown SW, Nair VK (2015) Imperfect vaccination can enhance the transmission of highly virulent pathogens. PLoS Biol 13:e1002198. https://doi.org/10.1371/journal.pbio.1002198
Spatz SJ, Petherbridge L, Zhao Y, Nair V (2007) Comparative full-length sequence analysis of oncogenic and vaccine (Rispens) strains of Marek’s disease virus. J Gen Virol 88:1080-1096. https://doi.org/10.1099/vir.0.82600-0
Spatz SJ, Silva RF (2007) Polymorphisms in the repeat long regions of oncogenic and attenuated pathotypes of Marek’s disease virus 1. Virus Genes 35:41–53. https://doi.org/10.1007/s11262-006-0024-5
Spatz SJ, Smith LP, Baigent SJ, Petherbridge L, Nair V (2011) Genotypic characterization of two artificial chromosome clones derived from a single DNA source of the very virulent gallid herpesvirus-2 strain C12/130. J Gen Viro 92:1500–1507. https://doi.org/10.1099/vir.0.027706-0
Tulman ER, Afonso CL, Lu Z, Zsak L, Rock DL, Kutish GF (2000) The genome of a very virulent Marek’s disease virus. J Virol 74:7980–7988. https://doi.org/10.1128/jvi.74.17.7980-7988.2000
Zhang F, Liu CJ, Zhaanalysisng YP, Li Z, Liu AL, Yan FH, Cheng Y (2012) Comparative full-length sequence analysis of Marek’s disease virus vaccine strain 814. Arch Virol 157:177–183. https://doi.org/10.1007/s00705-011-1131-8
Puro KU, Bhattacharjee U, Baruah S, Sen A, Das S, Ghatak S, Doley S, Sanjukta R, Shakuntala I (2018) Characterization of Marek's disease virus and phylogenetic analyses of meq gene from an outbreak in poultry in Meghalaya of Northeast India. Virus disease 29:167–172. https://doi.org/10.1007/s13337-018-0448-2
Author information
Authors and Affiliations
Contributions
ID collected and characterized the samples from commercial flocks, provided them for sequencing and wrote the manuscript. GM carried out the study conception and design, the experiment analysis, interpretation of results and manuscript preparation. EC and CL provided critical feedback and helped shape the research, analysis and manuscript. GC carried out the experiment and manuscript preparation. LM carried out the experiment.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical statement
The study reported in this paper was non-interventional and only used existing samples that were submitted for routine diagnostic purposes.
Additional information
Edited by Nicola Decaro.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Davidson, I., Lupini, C., Catelli, E. et al. Virulence evaluation of Israeli Marek’s disease virus isolates from commercial poultry using their meq gene sequence. Virus Genes 60, 32–43 (2024). https://doi.org/10.1007/s11262-023-02042-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-023-02042-7