Abstract
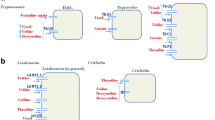
Trypanosomiasis is a major illness affecting camels in tropical and subtropical regions. Comparisons of camel and Trypanosoma evansi genomes can lead to the discovery of new drug targets for treating Trypanosoma infections. The synthesis pathways of cytosine, cytidine, cytidine monophosphate (CMP), cytidine diphosphate (CDP), cytidine triphosphate (CTP) deoxycytidine, deoxycytidine monophosphate (dCMP), deoxycytidine diphosphate (dCDP), and deoxycytidine triphosphate (dCTP) were compared in the dromedary camel (Camelus dromedarius) and T. evansi. None of the enzymes involved in cytosine pathway were detected in camels and T. evansi. Notably, cytidine kinase (CK) and 5′-nucleotidase, which interconverts cytidine to CMP, were not detected in T. evansi but were present in camels. UMP/CMP kinase was not predicted in T. evansi. Therefore, the presence of enzymes involved in the CTP synthesis cascade was not predicted in T. evansi. CMP synthesis might also be encoded by other enzymes, e.g., purine nucleotides kinases. Both camel and T. evansi share an efficient enzyme system for converting CDP to CTP. In conclusion, CTP synthase is important for homeostasis of cytosine nucleotides in T. evansi and could be a potential drug target against the parasite. In addition, the inhibition of UMP synthesis might contribute to parasite death as it is a shared source for CTP synthesis.





























Similar content being viewed by others
Availability of data and materials
All data are within the manuscript.
References
Abdi, R.D., Agga, G.E., Aregawi, W.G., Bekana, M., Van Leeuwen, T., Delespaux, V., Duchateau, L., 2017. A systematic review and meta-analysis of trypanosome prevalence in tsetse flies. BMC veterinary research,13, 100-100.
Artimo, P., Jonnalagedda, M., Arnold, K., Baratin, D., Csardi, G., de Castro, E., Duvaud, S., Flegel, V., Fortier, A., Gasteiger, E., Grosdidier, A., Hernandez, C., Ioannidis, V., Kuznetsov, D., Liechti, R., Moretti, S., Mostaguir, K., Redaschi, N., Rossier, G., Xenarios, I., Stockinger, H., 2012. ExPASy: SIB bioinformatics resource portal. Nucleic acids research,40, W597-W603.
Chávez-Fumagalli, M.A., Lage, D.P., Tavares, G.S., Mendonça, D.V., Dias, D.S., Ribeiro, P.A., Ludolf, F., Costa, L.E., Coelho, V.T., Coelho, E.A., 2019. In silico Leishmania proteome mining applied to identify drug target potential to be used to treat against visceral and tegumentary leishmaniasis. Journal of Molecular Graphics and Modelling,87, 89-97.
Cook, C.E., Bergman, M.T., Cochrane, G., Apweiler, R., Birney, E., 2018. The European Bioinformatics Institute in 2017: data coordination and integration. Nucleic acids research,46, D21-D29.
Cowman, A.F., Crabb, B.S., 2003. Functional genomics: identifying drug targets for parasitic diseases. TRENDS in Parasitology,19, 538-543.
Desquesnes, M., Dargantes, A., Lai, D.H., Lun, Z.R., Holzmuller, P., Jittapalapong, S., 2013. Trypanosoma evansi and surra: a review and perspectives on transmission, epidemiology and control, impact, and zoonotic aspects. BioMed research international,2013, 321237.
Ebhodaghe, F., Ohiolei, J.A., Isaac, C., 2018. A systematic review and meta-analysis of small ruminant and porcine trypanosomiasis prevalence in sub-Saharan Africa (1986 to 2018). Acta tropica,188, 118-131.
el Kouni, M.H., 2017. Pyrimidine metabolism in schistosomes: a comparison with other parasites and the search for potential chemotherapeutic targets. Comparative Biochemistry and Physiology Part B: Biochemistry and Molecular Biology,213, 55-80.
Fox, B., Belperron, A., Bzik, D., 1999. Stable transformation of Toxoplasma gondii based on a pyrimethamine resistant trifunctional dihydrofolate reductase-cytosine deaminase-thymidylate synthase gene that confers sensitivity to 5-fluorocytosine. Molecular and biochemical parasitology,98, 93-103.
Fox, B.A., Bzik, D.J., 2020. Biochemistry and metabolism of Toxoplasma gondii: purine and pyrimidine acquisition in Toxoplasma gondii and other Apicomplexa, Toxoplasma gondii. Elsevier, pp. 397-449.
Garavito, M.F., Narváez-Ortiz, H.Y., Zimmermann, B.H., 2015. Pyrimidine metabolism: dynamic and versatile pathways in pathogens and cellular development. Journal of genetics and genomics,42, 195-205.
Gasteiger, E., Gattiker, A., Hoogland, C., Ivanyi, I., Appel, R.D., Bairoch, A., 2003. ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic acids research,31, 3784-3788.
Ginger, M.L., Ngazoa, E.S., Pereira, C.A., Pullen, T.J., Kabiri, M., Becker, K., Gull, K., Steverding, D., 2005. Intracellular positioning of isoforms explains an unusually large adenylate kinase gene family in the parasite Trypanosoma brucei. Journal of Biological Chemistry,280, 11781-11789.
Habrian, C., Chandrasekhara, A., Shahrvini, B., Hua, B., Lee, J., Jesinghaus, R., Barry, R., Gitai, Z., Kollman, J., Baldwin, E.P., 2016. Inhibition of Escherichia coli CTP synthetase by NADH and Other Nicotinamides and Their Mutual Interactions with CTP and GTP. Biochemistry,55, 5554-5565.
Huson, D.H., Richter, D.C., Rausch, C., Dezulian, T., Franz, M., Rupp, R., 2007. Dendroscope: An interactive viewer for large phylogenetic trees. BMC bioinformatics,8, 460.
Jirimutu, Wang, Z., Ding, G., Chen, G., Sun, Y., Sun, Z., Zhang, H., Wang, L., Hasi, S., Zhang, Y., Li, J., Shi, Y., Xu, Z., He, C., Yu, S., Li, S., Zhang, W., Batmunkh, M., Ts, B., Narenbatu, Unierhu, Bat-Ireedui, S., Gao, H., Baysgalan, B., Li, Q., Jia, Z., Turigenbayila, Subudenggerile, Narenmanduhu, Wang, Z., Wang, J., Pan, L., Chen, Y., Ganerdene, Y., Dabxilt, Erdemt, Altansha, Altansukh, Liu, T., Cao, M., Aruuntsever, Bayart, Hosblig, He, F., Zha-ti, A., Zheng, G., Qiu, F., Sun, Z., Zhao, L., Zhao, W., Liu, B., Li, C., Chen, Y., Tang, X., Guo, C., Liu, W., Ming, L., Temuulen, Cui, A., Li, Y., Gao, J., Li, J., Wurentaodi, Niu, S., Sun, T., Zhai, Z., Zhang, M., Chen, C., Baldan, T., Bayaer, T., Li, Y., Meng, H., 2012. Genome sequences of wild and domestic bactrian camels. Nat Commun,3, 1202.
Kandeel, M., Ando, T., Kitamura, Y., Abdel-Aziz, M., Kitade, Y., 2009. Mutational, inhibitory and microcalorimetric analyses of Plasmodium falciparum TMP kinase. Implications for drug discovery. Parasitology,136, 11-25.
Kanehisa, M., Araki, M., Goto, S., Hattori, M., Hirakawa, M., Itoh, M., Katayama, T., Kawashima, S., Okuda, S., Tokimatsu, T., 2007. KEGG for linking genomes to life and the environment. Nucleic acids research,36, D480-D484.
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y., Morishima, K., 2016. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic acids research,45, D353-D361.
Labarga, A., Valentin, F., Anderson, M., Lopez, R., 2007. Web services at the European bioinformatics institute. Nucleic acids research,35, W6-W11.
Madden, T., 2013. The BLAST sequence analysis tool, The NCBI Handbook [Internet]. 2nd edition. National Center for Biotechnology Information (US).
Marchler-Bauer, A., Anderson, J.B., Cherukuri, P.F., DeWeese-Scott, C., Geer, L.Y., Gwadz, M., He, S., Hurwitz, D.I., Jackson, J.D., Ke, Z., 2005. CDD: a Conserved Domain Database for protein classification. Nucleic acids research,33, D192-D196.
Mori, G., Chiarelli, L.R., Esposito, M., Makarov, V., Bellinzoni, M., Hartkoorn, R.C., Degiacomi, G., Boldrin, F., Ekins, S., de Jesus Lopes Ribeiro, A.L., Marino, L.B., Centarova, I., Svetlikova, Z., Blasko, J., Kazakova, E., Lepioshkin, A., Barilone, N., Zanoni, G., Porta, A., Fondi, M., Fani, R., Baulard, A.R., Mikusova, K., Alzari, P.M., Manganelli, R., de Carvalho, L.P., Riccardi, G., Cole, S.T., Pasca, M.R., 2015. Thiophenecarboxamide Derivatives Activated by EthA Kill Mycobacterium tuberculosis by Inhibiting the CTP Synthetase PyrG. Chem Biol,22, 917-927.
Narvaez-Ortiz, H.Y., Lopez, A.J., Gupta, N., Zimmermann, B.H., 2018. A CTP synthase undergoing stage-specific spatial expression is essential for the survival of the intracellular parasite Toxoplasma gondii. Frontiers in Cellular and Infection Microbiology,8, 83.
Ogata, H., Goto, S., Fujibuchi, W., Kanehisa, M., 1998. Computation with the KEGG pathway database. Biosystems,47, 119-128.
Schimmel, K.J., Gelderblom, H., Guchelaar, H.J., 2007. Cyclopentenyl cytosine (CPEC): an overview of its in vitro and in vivo activity. Curr Cancer Drug Targets,7, 504-509.
Sequencing, H., 2011. CLC Genomics Workbench. Workbench.
Tamborini, L., Pinto, A., Smith, T.K., Major, L.L., Iannuzzi, M.C., Cosconati, S., Marinelli, L., Novellino, E., Lo Presti, L., Wong, P.E., Barrett, M.P., De Micheli, C., Conti, P., 2012. Synthesis and biological evaluation of CTP synthetase inhibitors as potential agents for the treatment of African trypanosomiasis. ChemMedChem,7, 1623-1634.
Yoshida, T., Nasu, H., Namba, E., Ubukata, O., Yamashita, M., 2012. Discovery of a compound that acts as a bacterial PyrG (CTP synthase) inhibitor. J Med Microbiol,61, 1280-1285.
Acknowledgements
The authors acknowledge the financial support of this project by King Abdul-Aziz City for Science and Technology (KACST), Basic Research Programs, National Transformation Program, under Research and Development Grants Program for National Research Institutions and Centers (GPURC), Kingdom of Saudi Arabia (Grant No. 2-17-04-004-0001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
Not apply to this work.
Informed consent
Not apply to this work.
Additional information
This article belongs to the Topical Collection: Camelids
Guest Editor: Bernard Faye
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kandeel, M., Al-Taher, A. Metabolic drug targets of the cytosine metabolism pathways in the dromedary camel (Camelus dromedarius) and blood parasite Trypanosoma evansi. Trop Anim Health Prod 52, 3337–3358 (2020). https://doi.org/10.1007/s11250-020-02366-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11250-020-02366-8