Abstract
Finite Gaussian mixture models are widely used for model-based clustering of continuous data. Nevertheless, since the number of model parameters scales quadratically with the number of variables, these models can be easily over-parameterized. For this reason, parsimonious models have been developed via covariance matrix decompositions or assuming local independence. However, these remedies do not allow for direct estimation of sparse covariance matrices nor do they take into account that the structure of association among the variables can vary from one cluster to the other. To this end, we introduce mixtures of Gaussian covariance graph models for model-based clustering with sparse covariance matrices. A penalized likelihood approach is employed for estimation and a general penalty term on the graph configurations can be used to induce different levels of sparsity and incorporate prior knowledge. Model estimation is carried out using a structural-EM algorithm for parameters and graph structure estimation, where two alternative strategies based on a genetic algorithm and an efficient stepwise search are proposed for inference. With this approach, sparse component covariance matrices are directly obtained. The framework results in a parsimonious model-based clustering of the data via a flexible model for the within-group joint distribution of the variables. Extensive simulated data experiments and application to illustrative datasets show that the method attains good classification performance and model quality. The general methodology for model-based clustering with sparse covariance matrices is implemented in the R package mixggm, available on CRAN.













Similar content being viewed by others
References
Amerine, M.A.: The composition of wines. Sci Mon 77(5), 250–254 (1953)
Azizyan, M., Singh, A., Wasserman, L.: Efficient sparse clustering of high-dimensional non-spherical Gaussian mixtures. In: Artificial Intelligence and Statistics, pp. 37–45 (2015)
Baladandayuthapani, V., Talluri, R., Ji, Y., Coombes, K.R., Lu, Y., Hennessy, B.T., Davies, M.A., Mallick, B.K.: Bayesian sparse graphical models for classification with application to protein expression data. Ann. Appl. Stat. 8(3), 1443–1468 (2014)
Banfield, J.D., Raftery, A.E.: Model-based Gaussian and non-Gaussian clustering. Biometrics 49(3), 803–821 (1993)
Barber, R.F., Drton, M.: High-dimensional Ising model selection with Bayesian information criteria. Electr. J. Stat. 9(1), 567–607 (2015)
Baudry, J.P., Celeux, G.: EM for mixtures Initialization requires special care. Stat. Comput. 25(4), 713–726 (2015)
Bellman, R.: Dynamic Programming. Princeton University Press, Princeton (1957)
Bien, J., Tibshirani, R.J.: Sparse estimation of a covariance matrix. Biometrika 98(4), 807–820 (2011)
Biernacki, C., Lourme, A.: Stable and visualizable Gaussian parsimonious clustering models. Stat. Comput. 24(6), 953–969 (2014)
Bollobas, B.: Random Graphs. Cambridge University Press, Cambridge (2001)
Bouveyron, C., Brunet, C.: Simultaneous model-based clustering and visualization in the fisher discriminative subspace. Stat. Comput. 22(1), 301–324 (2012)
Bouveyron, C., Brunet-Saumard, C.: Model-based clustering of high-dimensional data: a review. Comput. Stat. Data Anal. 71, 52–78 (2014)
Bozdogan, H.: Intelligent statistical data mining with information complexity and genetic algorithms. In: Statistical Data Mining and Knowledge Discovery, pp. 15–56 (2004)
Celeux, G., Govaert, G.: Gaussian parsimonious clustering models. Pattern Recogn. 28(5), 781–793 (1995)
Chalmond, B.: A macro-DAG structure based mixture model. Stat. Methodol. 25, 99–118 (2015)
Chatterjee, S., Laudato, M., Lynch, L.A.: Genetic algorithms and their statistical applications: an introduction. Comput. Stat. Data Anal. 22(6), 633–651 (1996)
Chaudhuri, S., Drton, M., Richardson, T.S.: Estimation of a covariance matrix with zeros. Biometrika 94(1), 199–216 (2007)
Chen, J., Chen, Z.: Extended Bayesian information criteria for model selection with large model spaces. Biometrika 95(3), 759–771 (2008)
Ciuperca, G., Ridolfi, A., Idier, J.: Penalized maximum likelihood estimator for normal mixtures. Scand. J. Stat. 30(1), 45–59 (2003)
Coomans, D., Broeckaert, M., Jonckheer, M., Massart, D.: Comparison of multivariate discriminant techniques for clinical data—application to the thyroid functional state. Methods Inf. Med. 22, 93–101 (1983)
Danaher, P., Wang, P., Witten, D.M.: The joint graphical lasso for inverse covariance estimation across multiple classes. J. R. Stat. Soc. Ser. B (Stat. Methodol.) 76(2), 373–397 (2014)
Dempster, A.: Covariance selection. Biometrics 28(1), 157–175 (1972)
Dempster, A.P., Laird, N.M., Rubin, D.B.: Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. Ser. B 39(1), 1–38 (1977)
Drton, M., Maathuis, M.H.: Structure learning in graphical modeling. Annu. Rev. Stat. Appl. 4(1), 365–393 (2017)
Edwards, D.: Introduction to Graphical Modelling. Springer, Berlin (2000)
Erdős, P., Rényi, A.: On random graphs I. Publ. Math. (Debrecen) 6, 290–297 (1959)
Erdős, P., Rényi, A.: On the evolution of random graphs. Publ. Math. Inst. Hung. Acad. Sci. 5(1), 17–60 (1960)
Fop, M., Murphy, T.B.: Variable selection methods for model-based clustering. Stat. Surv. 12, 18–65 (2018)
Forina, M., Armanino, C., Castino, M., Ubigli, M.: Multivariate data analysis as a discriminating method of the origin of wines. Vitis 25(3), 189–201 (1986)
Foygel, R., Drton, M.: Extended Bayesian information criteria for Gaussian graphical models. In: Advances in Neural Information Processing Systems, pp. 604–612 (2010)
Fraley, C., Raftery, A.E.: Model-based clustering, discriminant analysis and density estimation. J. Am. Stat. Assoc. 97, 611–631 (2002)
Fraley, C., Raftery, A.E.: Bayesian regularization for normal mixture estimation and model-based clustering. Technical Report 486, Department of Statistics, University of Washington (2005)
Fraley, C., Raftery, A.E.: Bayesian regularization for normal mixture estimation and model-based clustering. J. Classif. 24(2), 155–181 (2007)
Friedman, J., Hastie, T., Tibshirani, R.: Sparse inverse covariance estimation with the graphical lasso. Biostatistics 9(3), 432–441 (2008)
Friedman, N.: Learning belief networks in the presence of missing values and hidden variables. In: Fisher, D. (ed.) Proceedings of the Fourteenth International Conference on Machine Learning, pp. 125–133. Morgan Kaufmann (1997)
Friedman, N.: The Bayesian structural EM algorithm. In: Proceedings of the Fourteenth Conference on Uncertainty in Artificial Intelligence, pp. 129–138. Morgan Kaufmann (1998)
Frühwirth-Schnatter, S.: Finite Mixture and Markov Switching Models. Springer, Berlin (2006)
Galimberti, G., Soffritti, G.: Using conditional independence for parsimonious model-based Gaussian clustering. Stat. Comput. 23(5), 625–638 (2013)
Galimberti, G., Manisi, A., Soffritti, G.: Modelling the role of variables in model-based cluster analysis. Stat. Comput. 28, 1–25 (2017)
Gao, C., Zhu, Y., Shen, X., Pan, W.: Estimation of multiple networks in Gaussian mixture models. Electr. J. Stat. 10(1), 1133–1154 (2016)
Garber, J., Cobin, R., Gharib, H., Hennessey, J., Klein, I., Mechanick, J., Pessah-Pollack, R., Singer, P., Woeber, K.: Clinical practice guidelines for hypothyroidism in adults: cosponsored by the American Association of Clinical Endocrinologists and the American Thyroid Association. Endocr. Pract. 18(6), 988–1028 (2012)
Goldberg, D.: Genetic Algorithms in Search, Optimization, and Machine Learning. Addison-Wesley, Boston (1989)
Green, P.J.: On use of the EM for penalized likelihood estimation. J. R. Stat. Soc. Ser. B (Methodol.) 52, 443–452 (1990)
Greenhalgh, D., Marshall, S.: Convergence criteria for genetic algorithms. SIAM J. Comput. 30(1), 269–282 (2000)
Guo, J., Levina, E., Michailidis, G., Zhu, J.: Joint estimation of multiple graphical models. Biometrika 98(1), 1–15 (2011)
Harbertson, J.F., Spayd, S.: Measuring phenolics in the winery. Am. J. Enol. Vitic. 57(3), 280–288 (2006)
Hoeting, J.A., Madigan, D., Raftery, A.E., Volinsky, C.T.: Bayesian model averaging: a tutorial. Stat. Sci. 14(4), 382–417 (1999)
Holland, J.H.: Genetic algorithms. Sci. Am. 267(1), 66–72 (1992)
Huang, J.Z., Liu, N., Pourahmadi, M., Liu, L.: Covariance matrix selection and estimation via penalised normal likelihood. Biometrika 93(1), 85–98 (2006)
Hubert, L., Arabie, P.: Comparing partitions. J. Classif. 2, 193–218 (1985)
Kauermann, G.: On a dualization of graphical Gaussian models. Scand. J. Stat. 23(1), 105–116 (1996)
Koller, D., Friedman, N.: Probabilistic Graphical Models: Principles and Techniques. MIT Press, Cambridge (2009)
Kriegel, H.P., Schubert, E., Zimek, A.: The (black) art of runtime evaluation: are we comparing algorithms or implementations? Knowl. Inf. Syst. 52(2), 341–378 (2017)
Krishnamurthy, A.: High-dimensional clustering with sparse Gaussian mixture models. Unpublished paper (2011)
Kumar, M.S., Safa, A.M., Deodhar, S.D., SO, P.: The relationship of thyroid-stimulating hormone (TSH), thyroxine (T4), and triiodothyronine (T3) in primary thyroid failure. Am. J. Clin. Pathol. 68(6), 747–751 (1977)
Lee, KH., Xue, L.: Nonparametric finite mixture of Gaussian graphical models. Technometrics (2017)
Lotsi, A., Wit, E.: High dimensional sparse Gaussian graphical mixture model. arXiv preprint arXiv:1308.3381 (2013)
Ma, J., Michailidis, G.: Joint structural estimation of multiple graphical models. J. Mach. Learn. Res. 17(166), 1–48 (2016)
Madigan, D., Raftery, A.E.: Model selection and accounting for model uncertainty in graphical models using Occam’s window. J. Am. Stat. Assoc. 89(428), 1535–1546 (1994)
Malsiner-Walli, G., Frühwirth-Schnatter, S., Grün, B.: Model-based clustering based on sparse finite Gaussian mixtures. Stat. Comput. 26(1), 303–324 (2016)
MartÄśÌĄnez, A.M., Vitria, J.: Learning mixture models using a genetic version of the EM algorithm. Pattern Recogn. Lett. 21(8), 759–769 (2000)
Maugis, C., Celeux, G., Martin-Magniette, M.L.: Variable selection for clustering with Gaussian mixture models. Biometrics 65, 701–709 (2009)
McLachlan, G., Peel, D.: Finite Mixture Models. Wiley, New York (2000)
McLachlan, G.J., Rathnayake, S.: On the number of components in a Gaussian mixture model. Wiley Interdiscipl. Rev. Data Min. Knowl. Discov. 4(5), 341–355 (2014)
McNicholas, D.P., Murphy, T.B.: Parsimonious Gaussian mixture models. Stat. Comput. 18(3), 285–296 (2008)
McNicholas, P.D.: Model-based clustering. J. Classif. 33(3), 331–373 (2016)
Miller, A.: Subset Selection in Regression. Chapman & Hall/CRC, London (2002)
Mohan, K., Chung, M., Han, S., Witten, D., Lee, Si., Fazel, M.: Structured learning of Gaussian graphical models. In: Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q. (eds.) Advances in Neural Information Processing Systems, vol. 25, pp. 620–628 (2012)
Mohan, K., London, P., Fazel, M., Witten, D., Lee, S.I.: Node-based learning of multiple Gaussian graphical models. J. Mach. Learn. Res. 15(1), 445–488 (2014)
Pan, W., Shen, X.: Penalized model-based clustering with application to variable selection. J. Mach. Learn. Res. 8, 1145–1164 (2007)
Pan, W., Shen, X., Jiang, A., Hebbel, R.P.: Semi-supervised learning via penalized mixture model with application to microarray sample classification. Bioinformatics 22(19), 2388–2395 (2006)
Pernkopf, F., Bouchaffra, D.: Genetic-based EM algorithm for learning Gaussian mixture models. IEEE Trans. Pattern Anal. Mach. Intell. 27(8), 1344–1348 (2005)
Peterson, C., Stingo, F.C., Vannucci, M.: Bayesian inference of multiple Gaussian graphical models. J. Am. Stat. Assoc. 110(509), 159–174 (2015)
Poli, I., Roverato, A.: A genetic algorithm for graphical model selection. J. Ital. Stat. Soc. 7(2), 197–208 (1998)
Pourahmadi, M.: Covariance estimation: the GLM and regularization perspectives. Stat. Sci. 26(3), 369–387 (2011)
R Core Team: R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria (2017) https://www.R-project.org
Raftery, A.E., Dean, N.: Variable selection for model-based clustering. J. Am. Stat. Assoc. 101, 168–178 (2006)
Richardson, T., Spirtes, P.: Ancestral graph markov models. Ann. Stat. 30(4), 962–1030 (2002)
Rodríguez, A., Lenkoski, A., Dobra, A.: Sparse covariance estimation in heterogeneous samples. Electr. J. Stat. 5, 981–1014 (2011)
Rothman, A.J.: Positive definite estimators of large covariance matrices. Biometrika 99(3), 733–740 (2012)
Roverato, A.: Hyper inverse Wishart distribution for non-decomposable graphs and its application to Bayesian inference for Gaussian graphical models. Scand. J. Stat. 29(3), 391–411 (2002)
Roverato, A., Paterlini, S.: Technological modelling for graphical models: an approach based on genetic algorithms. Comput. Stat. Data Anal. 47(2), 323–337 (2004)
Ruan, L., Yuan, M., Zou, H.: Regularized parameter estimation in high-dimensional Gaussian mixture models. Neural Comput. 23(6), 1605–1622 (2011)
Schwarz, G.: Estimating the dimension of a model. Ann. Stat. 6(2), 461–464 (1978)
Scrucca, L.: GA: A package for genetic algorithms in R. J. Stat. Softw. 53(4), 1–37 (2013)
Scrucca, L.: Genetic algorithms for subset selection in model-based clustering. In: Celebi, M.E., Aydin, K. (eds.) Unsupervised Learning Algorithms, pp. 55–70. Springer, Berlin (2016)
Scrucca, L.: On some extensions to GA package: hybrid optimisation, parallelisation and Islands evolution. R J. 9(1), 187–206 (2017)
Scrucca, L., Raftery, A.E.: Improved initialisation of model-based clustering using Gaussian hierarchical partitions. Adv. Data Anal. Classif. 9(4), 447–460 (2015)
Scrucca, L., Fop, M., Murphy, T.B., Raftery, A.E.: mclust 5: Clustering, classification and density estimation using Gaussian finite mixture models. R J. 8(1), 289–317 (2016)
Sharapov, R.R., Lapshin, A.V.: Convergence of genetic algorithms. Pattern Recogn. Image Anal. 16(3), 392–397 (2006)
Shen, X., Ye, J.: Adaptive model selection. J. Am. Stat. Assoc. 97(457), 210–221 (2002)
Talluri, R., Baladandayuthapani, V., Mallick, B.K.: Bayesian sparse graphical models and their mixtures. Stat 3(1), 109–125 (2014)
Tan, K.M.: hglasso: Learning graphical models with hubs. R package version 12. (2014) https://CRAN.R-project.org/package=hglasso
Thiesson, B., Meek, C., Chickering, D.M., Heckerman, D.: Learning mixtures of DAG models. In: Proceedings of the Fourteenth Conference on Uncertainty in Artificial Intelligence, pp 504–513 (1997)
Titterington, D., Smith, A., Makov, U.: Statistical Analysis of Finite Mixture Distributions. Wiley, London (1985)
Wang, H.: Scaling it up: Stochastic search structure learning in graphical models. Bayesian Anal. 10(2), 351–377 (2015)
Wermuth, N., Cox, D., Marchetti, G.M.: Covariance chains. Bernoulli 12(5), 841–862 (2006)
Whittaker, J.: Graphical Models in Applied Multivariate Statistics. Wiley, London (1990)
Wiegand, R.E.: Performance of using multiple stepwise algorithms for variable selection. Stat. Med. 29(15), 1647–1659 (2010)
Wu, C.F.J.: On the convergence properties of the EM algorithm. Ann. Stat. 11(1), 95–103 (1983)
Xie, B., Pan, W., Shen, X.: Variable selection in penalized model-based clustering via regularization on grouped parameters. Biometrics 64(3), 921–930 (2008)
Yuan, M., Lin, Y.: Model selection and estimation in the Gaussian graphical model. Biometrika 94(1), 19–35 (2007)
Zhou, H., Pan, W., Shen, X.: Penalized model-based clustering with unconstrained covariance matrices. Electr. J. Stat. 3, 1473–1496 (2009)
Zhou, S., RÃijtimann, P., Xu, M., BÃijhlmann, P.: High-dimensional covariance estimation based on Gaussian graphical models. J. Mach. Learn. Res. 12, 2975–3026 (2011)
Zhu, Y., Shen, X., Pan, W.: Structural pursuit over multiple undirected graphs. J. Am. Stat. Assoc. 109(508), 1683–1696 (2014)
Zou, H., Hastie, T., Tibshirani, R.: On the “degrees of freedom” of the lasso. Ann. Stat. 35(5), 2173–2192 (2007)
Acknowledgements
We thank the editor and the anonymous referees for their valuable comments, which substantially improved the quality of the work. Michael Fop’s and Thomas Brendan Murphy’s research was supported by the Science Foundation Ireland funded Insight Research Centre (SFI/12/RC/2289). Luca Scrucca received the support of “Fondo Ricerca di Base, 2015” from Università degli Studi di Perugia for the project “Parallel genetic algorithms with applications in statistical estimation and evaluation”.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendix A: Iterative conditional fitting algorithm
The ICF algorithm (Chaudhuri et al. 2007) is employed to estimate a sparse covariance matrix given a certain structure of association. In this appendix, we present the algorithm in application to Gaussian mixture model estimation and we extend it to allow for Bayesian regularization of the covariance matrix.
Given a graph \({\mathcal {G}}_k = ({\mathcal {V}}, {\mathcal {E}}_k)\), to find the corresponding sparse covariance matrix under the constraint of being positive definite we need to maximize the objective function:
Let us make use of the following conventions: subscript [j, h] denotes element (j, h) of a matrix, a negative index such as \(-j\) denotes that row or column j has been removed, subscript \([\,,j]\) (or \([j,\,]\)) denotes that column (or row) j has been selected. Moreover, we denote with s(j) the set of indexes corresponding to the variables connected to variable \(X_j\) in the graph, i.e. the positions of the non zero entries in the covariance matrix for \(X_j\). Following Chaudhuri et al. (2007), the ICF algorithm is implemented as follows:
-
1.
Set the iteration counter \(r=0\). Initialize the covariance matrix \({\hat{{{\varvec{\Sigma }}}}}^{(0)}_k = \text {diag}({\mathbf {S}}_k)\).
-
2.
For \(j = (1,\, \ldots ,\, V)\)
-
2a
compute \(\varvec{\varOmega }_k^{(r)} = ({\hat{{{\varvec{\Sigma }}}}}^{(r)}_{k[-j,-j]})^{-1}\)
-
2b
compute the covariance terms estimates
$$\begin{aligned} {\hat{{{\varvec{\Sigma }}}}}^{(r)}_{k[j,s(j)]}= & {} \left( {\mathbf {S}}_{k[j,-j]}\,\varvec{\varOmega }^{(r)}_{k[\,,s(j)]} \right) \,\\&\quad \left( \varvec{\varOmega }^{(r)}_{k[s(j),\,]} {\mathbf {S}}_{k[-j,-j]} \varvec{\varOmega }^{(r)}_{k[\,,s(j)]} \right) \end{aligned}$$ -
2c
compute \(\lambda _j {=} {\mathbf {S}}_{k[j,j]} - {\hat{{{\varvec{\Sigma }}}}}^{(r)}_{k[j,s(j)]} \left( {\mathbf {S}}_{k[j,-j]}\,\varvec{\varOmega }^{(r)}_{k[\,,s(j)]} \right) ^{\!\top }\)
-
2d
compute the variance term estimate
$$\begin{aligned} {\hat{{{\varvec{\Sigma }}}}}^{(r)}_{k[j,j]} = \lambda _j + {\hat{{{\varvec{\Sigma }}}}}^{(r)}_{k[j,s(j)]} \varvec{\varOmega }^{(r)}_{k[s(j),s(j)]} {\hat{{{\varvec{\Sigma }}}}}^{(r)}_{k[s(j),j]} \end{aligned}$$
-
2a
-
3.
Set \({\hat{{{\varvec{\Sigma }}}}}^{(r+1)}_k = {\hat{{{\varvec{\Sigma }}}}}_k^{(r)}\), increment \(r = r + 1\) and return to (2).
The algorithm stops when the increase in the objective function is less than a pre-specified tolerance. The covariance matrix in output has zero entries corresponding to the graph structure and is guaranteed of being positive definite.
In the case of Bayesian regularization, the objective function becomes:
where
The shape of the objective function corresponds to the one not regularized. Therefore, the same algorithm can be applied replacing \(N_k\) and \({\mathbf {S}}_k\) with \({\tilde{N}}_k\) and \(\tilde{{\mathbf {S}}}_k\).
Appendix B: Initialization of the S-EM algorithm
The S-EM algorithm requires two initialization steps: initialization of cluster allocations and initialization of the graph structure search. For the first task we use the Gaussian model-based hierarchical clustering approach of Scrucca and Raftery (2015), which has been shown to yield good starting points, be computationally efficient and work well in practice. For initialization of the graph structure search we use the following approach. Let \({\mathbf {R}}_k\) be the correlation matrix for component k, computed as:
where \({\mathbf {U}}_k\) is a diagonal matrix whose elements are \({\mathbf {S}}_{k,[j,j]}^{-1/2}\) for \(j=1,\ldots ,V\), i.e. the within component sample standard deviations. A sound strategy is to initialize the search for the optimal association structure by looking at the most correlated variables. Therefore, we define the adjacency matrix \({\mathbf {A}}_k\) whose off-diagonal elements \(a_{jhk}\) are given by:
where \(r_{jhk}\) is an off-diagonal element of \({\mathbf {R}}_k\) and \(\rho \) is a threshold value. In practice, we define a vector of values for \(\rho \) ranging from 0.4 to 1. For each value of \(\rho \), the related adjacency matrix is derived and the corresponding sparse covariance matrix is estimated using the ICF algorithm. Then the different adjacency matrices are ranked according to their value of the objective function in (5). Subsequently the structure search starts from the adjacency matrix at the top of the rank.
Appendix C: Details of simulation experiments
This appendix section describes the various simulated data scenarios considered in Sect. 5 of the paper.
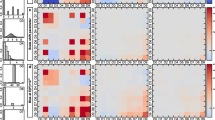
Scenario 1: In this setting we consider a structure with a single block of associated variables of size \(\left\lfloor {\frac{V}{2}}\right\rfloor \). The groups are differentiated by the position of the block, top corner, center and bottom corner respectively. Figure 3 displays an example of such structure for \(V=20\). To generate the covariance matrices, first we generate a \(V\times V\) matrix with all entries equal to 0.9 and diagonal 1. Then we use it as input of the ICF algorithm to estimate the corresponding covariance matrix with the given structure.
Scenario 2: For this scenario, the graphs are generated at random from an Erdős–Rényi model. The groups are characterized by different probabilities of connection, 0.3, 0.2 and 0.1 respectively. Figure 4 presents an example of a collection of structures of association for \(V=20\). Starting from a \(V\times V\) matrix with all entries equal to 0.9 and diagonal 1, we employ the ICF algorithm to estimate the corresponding sparse covariance matrix. In the simulated data experiment of Part III, we consider connection probabilities equal to 0.10, 0.05 and 0.03.
Scenario 3: This scenario is characterized by hubs, i.e. highly connected variables. Each cluster has \(\frac{V}{2}\) such hubs. The graph structures and the corresponding covariance matrices are generated randomly using the R package hglasso. (Tan 2014). The three groups have different sparsity levels, respectively 0.7, 0.8 and 0.9. Figure 5 presents an example of this type of graphs for \(V=20\). We point out that the method implemented in the package poses strict constraints on the covariance matrix and often some connected variables have weak correlations, making difficult to infer the association structure.
Scenario 4: Here the groups have structures of different types: block diagonal, random connections and Toeplitz type. For the first group we consider a block diagonal matrix with blocks of size 5. Regarding the second, the graph is generated at random from an Erdős–Rényi model with parameter 0.2. In both cases, we start from a \(V\times V\) matrix with all entries equal to 0.9 and diagonal 1, and then we employ the ICF algorithm to estimate the corresponding sparse covariance matrices. For the Toeplitz matrix we take \(\sigma _{j,\,j-1} = \sigma _{j-1,\,j} = 0.5\) for \(j=2,\,\ldots ,\,V\). Figure 6 depicts an example of these graph configurations for \(V=20\). In the simulated data experiment of Part III, we consider an Erdős–Rényi model with parameter 0.05 and a block diagonal matrix with 5 blocks of size 20; the Toeplitz matrix is generated as before.
Rights and permissions
About this article
Cite this article
Fop, M., Murphy, T.B. & Scrucca, L. Model-based clustering with sparse covariance matrices. Stat Comput 29, 791–819 (2019). https://doi.org/10.1007/s11222-018-9838-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11222-018-9838-y