Abstract
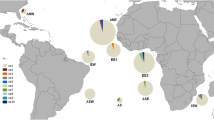
Sharks of the genus Rhizoprionodon are among the most important predators along the coastal marine ecosystems, and they represent an important economic resource for the small-scale fisheries. To properly manage and conserve exploited shark species, detailed analyses of their population structure are needed. To evaluate the gene flow and levels of the genetic diversity among populations of the Caribbean sharpnose shark R. porosus, we identified the nucleotide sequence based on collections (n = 321 specimens) from 10 different areas, including the Caribbean Sea and several locations along the entire Brazilian coast. The analysis of 802 nucleotides from the mitochondrial DNA control region revealed 53 distinct haplotypes. The majority of these haplotypes were restricted to their collection locales with a significant genetic structure detected among the overall populations (Φ ST = 0.237, P < 0.0001). The data suggest a population division with two distinct management units in the western Atlantic. These management units are likely separated by the Equatorial Current. The strong population structure in R. porosus indicates that regional populations, if depleted, will not recover swiftly through immigration.



Similar content being viewed by others
References
Aljanabi SM, Martinez I (1997) Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucleic Acids Res 25:4692–4693
Avise JC (2000) Phylogeography: The history and formation of species. Harvard University Press, Cambridge
Baum J, Myers RA, Kehler DG, Worm B, Harley SJ, Doherty PA (2003) Collapse and conservation of shark populations in the northwest Atlantic. Science 299:389–392
Beheregaray LB, Sunnucks P, Briscoe DA (2002) A rapid fish radiation associated with the last sealevel changes in southern Brazil: the silverside Odontesthes perugiae complex. Proc Roy Soc Lond Ser B Biol Sci 269:65–73
Brunner PC, Douglas MR, Osinov A, Wilson CC, Bernatchez L (2001) Holartic phylogeography of Arctic charr (Salvelinus alpines L.) inferred from mitochondrial DNA sequences. Evol 55:573–586
Castillo-Géniz JL, Márques-Farias JF, Rodriguez De La Cruz MC, Cortés C, CidDel Prado A (1998) TheMexican artisanal shark fishery in the Gulf of Mexico: towards a regulated fishery. Mar Freshw Res 49:611–620
Castro ALF, Stewart BS, Wilson SG (2007) Population genetic structure of Earth′s largest fish, the whale shark (Rhincodon typus). Mol Ecol 16:5183–5192
Chabot CL, Allen LG (2009) Global population structure of the tope (Galeorhinus galeus) inferred by mitochondrial control region sequence data. Mol Ecol 18:545–552
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate genegenealogies. Mol Ecol 9:1657–1660
Cody TJ, Avent RM (1980) Assessment of botton longline fishing off the central Texas coast. Manage. Data Sertex Texas Parks Wildlife Department 32
Compagno LJV (1984) FAO Species Catalogue. Vol 4. Parts 1 and 2, Sharks of the world. FAO Fisheries Synopsis 125
Crottini A, Andreone F, Kosuch J (2007) Fossorial but widespread: the phylogeography of the common spadefoot toad (Pelobates fuscus), and the role of the Po Valley as a major source of genetic variability. Mol Ecol 16:2734–2754
Duncan KM, Martin AP, Bowen BW, De Couet GH (2006) Global phylogeography of the scalloped hammerhead shark (Sphyrna lewini). Mol Ecol 15:2239–2251
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2005) ARLEQUIN ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinf Online 1:47–50
Fu YX (1996) Estimating the age of the common ancestor of a DNA sample using the number of segregating sites. Genetics 144:829–838
Gadig OBF (2001) Tubarões da Costa Brasileira. Tese de Doutorado, Unesp, Campus de Rio Claro, São Paulo 343
Gilbert CR (1967) A revision of the hammerhead sharks (Family Sphyrnidae). Proceedings of the U.S. National Museum 119:1–88
Grace M, Henwood T (1997) Assessment of the distribution and abundance of coastal sharks in the U.S. Gulf of Mexico and Eastern Seaboard, 1995 and 1996. Mar Fish Rev 59(4):23–32
Grant WS, Waples RS (2000) Spatial and temporal scales of genetic variability in marine and anadromous species: implications for fisheries oceanography. In: Harrison P, Parsons TR (eds) Fisheries oceanography: an integrative approach to fisheries ecology and management. Oxford, Blackwell, pp 61–93
Graves JE (1998) Molecular insights into the population structure of cosmopolitan marine fishes. J Hered 89:427–437
Grunwald C, Stabile J, Waldman JR, Gross R, Wirgin I (2002) Population genetics of shortnose sturgeon Acipenser brevirostrum based on mitochondrial DNA control region sequences. Mol Ecol 11:1885–1898
Harpending RC (1994) Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 66:591–600
Heist EJ (2004) Genetics of sharks, skates, and rays. In: Carrier JC, Musick JA, Heithaus MR (eds) Biology of sharks and their relatives. CRC Press, New York, pp 471–485
Heist EJ, Musick JA, Graves JE (1996) Genetic population structure of the shortfin mako (Isurus oxyrinchus) inferred from restriction fragment length polymorphism analysis of mitochondrial DNA. Can J Fish Aquatic Scie 53:583–588
Kasim HM (1991) Shark fishery of Veraval coast with special reference to population dynamics of Scoliodon laticaudus (Muller and Henle) and Rhizoprionodon acutus (Ruppell). J Mar Biol Assoc 33:213–228
Keeney DB, Heist EJ (2006) Worldwide phylogeography of the blacktip shark (Carcharhinus limbatus) inferred from mitochondrial DNA reveals isolation of western Atlantic populations coupled with recent Pacific dispersal. Mol Ecol 15:3669–3679
Keeney DB, Heupel MR, Hueter RE, Heist EJ (2005) Microsatellite and mitochondrial DNA analyses of the genetic structure of blacktip shark (Carcharhinus limbatus) nurseries in the north western Atlantic, Gulf of Mexico, and Caribbean Sea. Mol Ecol 14:1911–1923
Lessa R, Quijano SM, Santana FM, Monzini J (2009) Rhizoprionodon porosus. In: IUCN 2009. IUCN Red List of Threatened Species 2
Mantel N (1967) Detection of disease clustering and a generalized regression approach. Cancer Res 27:209
Márquez-Farias JF, Castillo-Géniz LC (1998) Fishery biology and demography of the Atlantic sharpnose shark, Rhizoprionodon terraenovae, in the southern Gulf of Mexico. Fish Res 2:183–198
Mattos SMG, Broadhurst MK, Hazin FHV, Jonnes DM (2001) Reproductive biology of the Caribbean sharpnose shark, Rhizoprionodon porosus, from Northern Brazil. Mar Freshw Res 52:745–752
Mendonça FF, Oliveira C, Gadig OBF, Foresti F (2009) Populations analysis of the Brazilian Sharpnose Shark Rhizoprionodon lalandii (Chondrichthyes: Carcharhinidae) on the São Paulo coast Southern Brazil: inferences from mt DNA sequences. Neotrop Ichth 7(2):213–216
Mendonça FF, Burgess G, Coelho R, Piercy A, Oliveira C, Gadig OBF, Foresti F (2011) Species delimitation in sharpnose sharks (genus Rhizoprionodon) in the western Atlantic Ocean using mitochondrial DNA. Conserv Genetics 12:193–200
Motta FS, Gadig OBF, Namora RC, Braga FM (2005) Size and sex compositions, length–weight relationship, and occurrence of the Brazilian sharpnose shark, Rhizoprionodon lalandii, caught by artisanal fishery from southeastern Brazil. Fish Res 74:116–126
Ostbye K, Bernatchez L, Næsje TF, Himberg M, Hindar K (2005) The evolutionary history of European whitefish (Coregonus lavaretus L.) as inferred from mtDNA phylogeography and Gillraker numbers. Mol Ecol 14:4371–4387
Pardini AT, Jones CS, Noble LR (2001) Sex-biased dispersal of great white sharks. Nature 412:139–140
Planes S, Doherty PJ, Bernardi G (2001) Strong genetic divergence among populations of a marine fish with limited dispersal, Acanthochromis polyacanthus, within the great barrier reef and the coral sea. Evol 55:2263–2273
Portnoy DS, McDowell JR, Heist EJ, Musick JA, Graves JE (2010) World phylogeography and male-mediated gene flow in the sandbar shark, Carcharhinus plumbeus. Mol Ecol 19:1994–2010
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rocha LA (2003) Patterns of distribution and processes of speciation in Brazilian reef Wshes. J Biogeog 30:1161–1171
Rocha LA, Robertson DR, Rocha CR (2005) Recent invasion of the tropical Atlantic by an Indo-Pacific coral reef fish. Mol Ecol 14:3921–3928
Rocha LA, Craig MT, Bowen BW (2007) Phylogeography and the conservation of coral reef fishes. Cor Reefs 26:501–512
Sadowsky V (1967) Selachier aus dem litoral von São Paulo, Brazil. Beitr Neotrop Fauna 5:71–88
Santos S, Schneider H, Sampaio I (2003) Genetic differentiation of Macrodon ancylodon (Sciaenidae, Perciformes) populations in Atlantic coastal waters of South America as revealed by mtDNA analysis. Genet Mol Biol 26:151–161
Santos S, Farias IP, Schneider H, Sampaio I (2006) Population genetic structuring of the king weakfish, Macrodon ancylodon (Sciaenidae), in Atlantic coastal waters of South America: deep genetic divergence without morphological change. Mol Ecol 15:4361–4373
Schneider S, Excoffier L (1999) Estimation of demographic parameters from the distribution of pairwise differences when the mutation rates vary among sites: application to human mitochondrial DNA. Genetics 152:1079–1089
Schrey AW, Heist EJ (2003) Microsatellite analysis of population structure of the shortfin mako (Isurus oxyrinchus). Can J Fish Aquatic Scie 60:670–675
Schultz JK, Feldheim KA, Gruber SH, Ashley MV, McGovern TM, Bowen BW (2008) Global phylogeography and seascape genetics of the lemon sharks (genus Negaprion). Mol Ecol 17(24):5336–5348
Simpfendorfer CA, Milward NE (1993) Utilization of a tropical bay as a nursery area by sharks of the families Carcharhinidae and Sphyrnidae. Environ Biol Fish 37:337–345
Springer VGA (1964) A revision of the Carcharhinidae shark genera Scoliodon, Loxodon. and Rhizoprionodon. Proc US Nat Museum 115:559–632
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotype associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132:619–633
Thompson JD, Higgins DG, Gibson TJ (1994) ClustalW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Valentin JL, André DL, Jacob SA (1987) Hydrobiology in the Cabo Frio (Brazil) upwelling: two dimensional structure and variability during a wind circle. Cont Shelf Res 7:77–88
Waples RS (1998) Evolutionarily significant units, distinct population segments, and the endangered species act: reply to Pennock and Dimmick. Conserv Biol 12:718–721
Waples RS, Gustafson RG, Weitkramp LA (2001) Characterizing diversity in salmon from the Pacific Northwest. J Fish Biol 59:1–41
Acknowledgments
We thank those who provided sharpnose shark species, especially George Burgess, Andrew Piercy, Ernesto Ron, Iracilda Sampaio, Marisa Fagundes Azevedo, Hugo Bornatowski and the Projeto Cação team. This work was supported by the Brazilian agency FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo/Proc. No. 06/588972).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mendonça, F.F., Oliveira, C., Gadig, O.B.F. et al. Phylogeography and genetic population structure of Caribbean sharpnose shark Rhizoprionodon porosus . Rev Fish Biol Fisheries 21, 799–814 (2011). https://doi.org/10.1007/s11160-011-9210-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11160-011-9210-1