Abstract
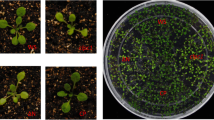
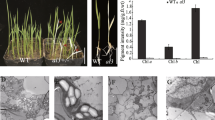
The ATP-dependent Clp protease has been well-characterized in Escherichia coli, but knowledge of its function in higher plants is limited. In bacteria, this two-component protease consists of a Ser-type endopeptidase ClpP, which relies on the ATP-dependent unfolding activity from an Hsp100 molecular chaperone to initiate protein degradation. In the chloroplasts of higher plants, multiple isoforms of the proteolytic subunit exist, with Arabidopsis having five ClpPs and four ClpP-like proteins termed ClpR predicted in its genome. In this work we characterized an Arabidopsis mutant impaired in one subunit of the chloroplast-localized Clp protease core, ClpR1. clpR1-1, a virescent mutant, carries a pre-mature stop codon in the clpR1 gene, resulting in no detectable ClpR1 protein. The accumulation of several chloroplast proteins, as well as most of the chloroplast-localized Clp protease subunits, is inhibited in clpR1-1. Unexpectedly, some plastid-encoded proteins do not accumulate, although their transcripts accumulate to wild-type levels. Maturation of 23S and 4.5S chloroplast ribosomal RNA (cp-rRNA) is delayed in clpR1-1, and both RNAs accumulate as higher molecular weight precursors. Also, chloroplasts in clpR1-1 are smaller than in wild type and have fewer thylakoid membranes with smaller grana stacks. We propose that a ClpR1-containing activity is required for chloroplast development and differentiation and in its absence both are delayed.







Similar content being viewed by others
Abbreviations
- cp-rRNA:
-
Chloroplast ribosomal RNA
- Lhcb2:
-
Light-harvesting Chl a/b binding protein
- RbcL/LSU and RbcS/SSU:
-
Transcript and protein of large and Small subunits of ribulose bisphosphate carboxylase/oxygenase (Rubisco)
- PsaL, PsaD and PsaF:
-
Subunits XI, II and III of photosystem I (PS I)
- Cpn60:
-
Chaperonin 60
- β-ATPase:
-
CF1 β subunit of ATPase
- PsbA and PsbD :
-
Transcripts encoding for D1 and D2 proteins of photosystem II (PSII)
- NF:
-
Norflurazon
- NPQ:
-
Non-photochemical excitation quenching
References
Adam Z, Adamska I, Nakabayashi K, Ostersetzer O, Haussuhl K, Manuell A, Zheng B, Vallon O, Rodermel SR, Shinozaki K, Clarke AK (2001) Chloroplast and mitochondrial proteases in Arabidopsis. A proposed nomenclature. Plant Physiol 125:1912–1918
Adam Z, Clarke AK (2002) Cutting edge of chloroplast proteolysis. Trends Plant Sci 7:451–456
Adam Z, Rudella A, van Wijk KJ (2006) Recent advances in the study of Clp, FtsH and other proteases located in chloroplasts. Curr Opin Plant Biol 9:234–240
Bellaoui M, Keddie JS, Gruissem W (2003) DCL is a plant-specific protein required for plastid ribosomal RNA processing and embryo development. Plant Mol Biol 53:531–543
Bisanz C, Begot L, Carol P, Perez P, Bligny M, Pesey H, Gallois JL, Lerbs-Mache S, Mache R (2003) The Arabidopsis nuclear DAL gene encodes a chloroplast protein which is required for the maturation of the plastid ribosomal RNAs and is essential for chloroplast differentiation. Plant Mol Biol 51:651–663
Bollenbach TJ, Lange H, Gutierrez R, Erhardt M, Stern DB, Gagliardi D (2005) RNR1, a 3′-5′ exoribonuclease belonging to the RNR superfamily, catalyzes 3′ maturation of chloroplast ribosomal RNAs in Arabidopsis thaliana. Nucleic Acids Res 33:2751–2763
Chen G, Bi YR, Li N (2005) EGY1 encodes a membrane-associated and ATP-independent metalloprotease that is required for chloroplast development. Plant J 41:364–375
Chen M, Choi Y, Voytas DF, Rodermel S (2000) Mutations in the Arabidopsis VAR2 locus cause leaf variegation due to the loss of a chloroplast FtsH protease. Plant J 22:303–313
Clarke AK (1999) ATP-dependent Clp proteases in phtosynthetic organisms—A cut above the rest! Ann Bot 83:593–599
Constan D, Froehlich JE, Rangarajan S, Keegstra K (2004) A stromal Hsp100 protein is required for normal chloroplast development and function in Arabidopsis. Plant Physiol 136:3605–3615
Dougan DA, Reid BG, Horwich AL, Bukau B (2002) ClpS, a substrate modulator of the ClpAP machine. Mol Cell 9:673–683
Gottesman S (1996) Proteases and their targets in Escherichia coli. Annu Rev Genet 30:465–506
Grimaud R, Kessel M, Beuron F, Steven AC, Maurizi MR (1998) Enzymatic and structural similarities between the Escherichia coli ATP-dependent proteases, ClpXP and ClpAP. J Biol Chem 273:12476–12481
Halperin T, Ostersetzer O, Adam Z (2001a) ATP-dependent association between subunits of Clp protease in pea chloroplasts. Planta 213:614–619
Halperin T, Zheng B, Itzhaki H, Clarke AK, Adam Z (2001b) Plant mitochondria contain proteolytic and regulatory subunits of the ATP-dependent Clp protease. Plant Mol Biol 45:461–468
Jarvis P, Chen LJ, Li H, Peto CA, Fankhauser C, Chory J (1998) An Arabidopsis mutant defective in the plastid general protein import apparatus. Science 282:100–103
Jarvis P, Robinson C (2004) Mechanisms of protein import and routing in chloroplasts. Curr Biol 14:R1064–1077
Kanazawa A, Kramer DM (2002) In vivo modulation of nonphotochemical exciton quenching (NPQ) by regulation of the chloroplast ATP synthesis. Proc Natl Acad Sci USA 99:12789–12794
Kandror O, Busconi L, Sherman M, Goldberg AL (1994) Rapid degradation of an abnormal protein in Escherichia coli involves the chaperones GroEL and GroES. J Biol Chem 269:23575–23582
Keeler SJ, Boettger CM, Haynes JG, Kuches KA, Johnson MM, Thureen DL, Keeler CL, Jr., Kitto SL (2000) Acquired thermotolerance and expression of the HSP100/ClpB genes of lima bean. Plant Physiol 123:1121–1132
Kishine M, Takabayashi A, Munekage Y, Shikanai T, Endo T, Sato F (2004) Ribosomal RNA processing and an RNase R family member in chloroplasts of Arabidopsis. Plant Mol Biol 55:595–606
Kössel H, Edwards K, Koch W, Langridge P, Schiefermayr E, Schwarz Z, Strittmatter G, Zenke G (1982) Structural and functional analysis of an rRNA operon and its flanking tRNA genes from Zea mays chloroplasts. Nucleic Acids Symp Ser:117–120
Kuroda H, Maliga P (2003) The plastid clpP1 protease gene is essential for plant development. Nature 425:86–89
Li J, Chory J (1997) A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90:929–938
Majeran W, Wollman FA, Vallon O (2000) Evidence for a role of ClpP in the degradation of the chloroplast cytochrome b (6) f complex. Plant Cell 12:137–150
Moreira D, Le Guyader H, Philippe H (2000) The origin of red algae and the evolution of chloroplasts. Nature 405:69–72
Muchizuki N, Brusslan JA, Larkin RM, Nagatani A, Chory J (2001) Arabidopsis genomes uncoupled 5 (gun5) mutant reveals the involvment of Mg-chelatase H subunit in plastid-to nucleus signal transduction. Proc Natl Acad Sci USA 98:2053–2058
Nakabayashi K, Ito M, Kiyosue T, Shinozaki K, Watanabe A (1999) Identification of clp genes expressed in senescing Arabidopsis leaves. Plant Cell Physiol 40:504–514
Nielsen E, Akita M, Davila-Aponte J, Keegstra K (1997) Stable association of chloroplastic precursors with protein translocation complexes that contain proteins from both envelope membranes and a stromal Hsp100 molecular chaperone. Embo J 16:935–946
Nott A, Jung HS, Koussevitzky S, Chory J (2006) Plastid-to-nucleus retrograde signaling. Annu Rev Plant Biol 57:739–759
Park S, Rodermel SR (2004) Mutations in ClpC2/Hsp100 suppress the requirement for FtsH in thylakoid membrane biogenesis. Proc Natl Acad Sci USA 101:12765–12770
Peltier JB, Ripoll DR, Friso G, Rudella A, Cai Y, Ytterberg J, Giacomelli L, Pillardy J, van Wijk KJ (2004) Clp protease complexes from photosynthetic and non-photosynthetic plastids and mitochondria of plants, their predicted three-dimensional structures, and functional implications. J Biol Chem 279:4768–4781
Peltier JB, Ytterberg J, Liberles DA, Roepstorff P, van Wijk KJ (2001) Identification of a 350-kDa ClpP protease complex with 10 different Clp isoforms in chloroplasts of Arabidopsis thaliana. J Biol Chem 276:16318–16327
Puente P, Wei N, Deng XW (1996) Combinatorial interplay of promoter elements constitutes the minimal determinants for light and developmental control of gene expression in Arabidopsis. EMBO J 15:3732–3743
Rodermel S, Haley J, Jiang CZ, Tsai CH, Bogorad L (1996) A mechanism for intergenomic integration: abundance of ribulose bisphosphate carboxylase small-subunit protein influences the translation of the large-subunit mRNA. Proc Natl Acad Sci USA 93:3881–3885
Rudella A, Friso G, Alonso JM, Ecker JR, van Wijk KJ (2006) Downregulation of ClpR2 Leads to Reduced Accumulation of the ClpPRS Protease Complex and Defects in Chloroplast Biogenesis in Arabidopsis. Plant Cell 18:1704–1721
Sakamoto W (2006) Protein degradation machineries in plastids. Annu Rev Plant Biol 57:599–621
Sakamoto W, Tamura T, Hanba-Tomita Y, Murata M (2002) The VAR1 locus of Arabidopsis encodes a chloroplastic FtsH and is responsible for leaf variegation in the mutant alleles. Genes Cells 7:769–780
Schirmer EC, Glover JR, Singer MA, Lindquist S (1996) HSP100/Clp proteins: a common mechanism explains diverse functions. Trends Biochem Sci 21:289–296
Shikanai T, Shimizu K, Ueda K, Nishimura Y, Kuroiwa T, Hashimoto T (2001) The chloroplast clpP gene, encoding a proteolytic subunit of ATP-dependent protease, is indispensable for chloroplast development in tobacco. Plant Cell Physiol 42:264–273
Shunxing J, Hilaire E, Guikema JA (2004) Identification and differential accumulation of two isoforms of the CF1-β subunit under high light stress in Brassica rapa. Plant Physiol Biochem 42:883–890
Sjögren LL, MacDonald TM, Sutinen S, Clarke AK (2004) Inactivation of the clpC1 gene encoding a chloroplast Hsp100 molecular chaperone causes growth retardation, leaf chlorosis, lower photosynthetic activity, and a specific reduction in photosystem content. Plant Physiol 136:4114–4126
Sokolenko A, Lerbs-Mache S, Altschmied L, Herrmann RG (1998) Clp protease complexes and their diversity in chloroplasts. Planta 207:286–295
Sugimoto H, Kusumi K, Tozawa Y, Yazaki J, Kishimoto N, Kikuchi S, Iba K (2004) The virescent-2 mutation inhibits translation of plastid transcripts for the plastid genetic system at an early stage of chloroplast differentiation. Plant Cell Physiol 45:985–996
Takechi K, Sodmergen, Murata M, Motoyoshi F, Sakamoto W (2000) The YELLOW VARIEGATED (VAR2) locus encodes a homologue of FtsH, an ATP-dependent protease in Arabidopsis. Plant Cell Physiol 41:1334–1346
VanderVere PS, Bennett TM, Oblong JE, Lamppa GK (1995) A chloroplast processing enzyme involved in precursor maturation shares a zinc-binding motif with a recently recognized family of metalloendopeptidases. Proc Natl Acad Sci USA 92:7177–7181
Wang J, Hartling JA, Flanagan JM (1997) The structure of ClpP at 2.3 Å resolution suggests a model for ATP-dependent proteolysis. Cell 91:447–456
Zheng B, MacDonald TM, Sutinen S, Hurry V, Clarke AK (2006) A nuclear-encoded ClpP subunit of the chloroplast ATP-dependent Clp protease is essential foe early development in Arabidopsis thaliana. Planta (in press)
Acknowledgement
This work was supported by the Howard Hughes Medical Institute and a grant from the Department of Energy to J.C., the Swedish Research Council for Environmental, Agricultural Sciences and Spatial Planning (Formas) to A.K.C., and an overseas postgraduate scholarship from the Natural Sciences and Engineering Research Council of Canada to T.M.S. S.K. was an EMBO long term fellow (ALTF 118–2000). J.C. is a Howard Hughes Medical Institute Investigator. We thank Takeshi Nakano (Riken Japan) for his help in analyzing chloroplast transcripts, Jason Lim for technical assistance and Olivier Loudet, Chris Schwartz and Jennifer Nemhauser for assisting in the map based cloning of clpR1-1. We thank Paul Sawchenko and the CCMI at Salk Institute for access to microscopy facilities. clpR1-2 seeds and the full length clpR1 cDNA clone were provided by The Arabidopsis Biological Resource Centre (ABRC), Ohio State University (Columbus, Ohio).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Koussevitzky , S., Stanne, T.M., Peto, C.A. et al. An Arabidopsis thaliana virescent mutant reveals a role for ClpR1 in plastid development. Plant Mol Biol 63, 85–96 (2007). https://doi.org/10.1007/s11103-006-9074-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-006-9074-2