Abstract
Background
Glioma is the most common primary brain tumor. Clear classification is crucial for accurate diagnosis and individualized treatment. Histopathological characteristics and genetic alterations have shown to be related to prognosis and treatment response. Germline variants are important components of genetic alterations. However, the distribution of germline variations in glioma patients and their association with survival remain unknown.
Methods
We carried out whole-exome sequencing on 99 cases to explore germline variants in glioma. We also analyzed the association of germline variants with clinicopathological features and other prognostic indicators.
Results
All the glioma cases harbored rare germline variants. Germline ALK variants (gALK-Mut) were identified in 12/99 (12.12%) patients. The gALK-Mut patients had significantly shorter overall survival than germline ALK wildtype (gALK-WT) patients in the all glioma group (99 cases) and the subset of patients with IDH-wildtype glioblastoma (IDH-WT-GBM, 39 cases) (P = 0.013 and 0.027, respectively). The gALK-Mut patients also had higher frequency of BIRC5, PIK3CA and RPN1 somatic mutations than the gALK-WT patients in IDH-WT-GBM. Other confounding factors appeared to contribute to patient survival. The subgroup of patients in IDH-WT-GBM with gALK-Mut/TP53-Mut had worse prognosis than the gALK-WT/TP53-Mut subgroup (P = 0.031); The gALK-Mut/TERT-WT and gALK-Mut/TERT-Mut subgroups both had a worse prognosis than the gALK-WT/TERT-Mut subgroup (P = 0.031 and 0.018, respectively).
Conclusions
Our study revealed ALK variation was an independent indicator of poor prognosis in glioma and IDH-WT-GBM. It could be a promising biomarker and tractable therapeutic target for this deadly disease.






Similar content being viewed by others
Data Availability
Data are available by contacting the corresponding author.
References
Hsieh C-C, Hsu H-S, Chang S-C (2016) Yann-Jang Chen (2016) Circulating Cell-Free DNA Levels Could Predict Oncological Outcomes of Patients Undergoing Esophagectomy for Esophageal Squamous Cell Carcinoma. Int J Mol Sci 17(12):2131
Sanai N, Alvarez-Buylla A, Berger MS (2005) Neural stem cells and the origin of gliomas. N Engl J Med 353(8):811–822
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109
Gilbert MR, Wang M, Aldape KD, Stupp R, Hegi ME, Jaeckle KA, Armstrong TS, Wefel JS, Won M, Blumenthal DT et al (2013) Dose-dense temozolomide for newly diagnosed glioblastoma: a randomized phase III clinical trial. J Clin Oncol 31(32):4085–4091
Aldape K, Brindle KM, Chesler L, Chopra R, Gajjar A, Gilbert MR, Gottardo N, Gutmann DH, Hargrave D, Holland EC et al (2019) Challenges to curing primary brain tumours. Nat Rev Clin Oncol 16(8):509–520
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131(6):803–820
Sun H, Yin L, Li S, Han S, Song G, Liu N, Yan C (2013) Prognostic significance of IDH mutation in adult low-grade gliomas: a meta-analysis. J Neurooncol 113(2):277–284
Aoki K, Nakamura H, Suzuki H, Matsuo K, Kataoka K, Shimamura T, Motomura K, Ohka F, Shiina S, Yamamoto T (2018) Prognostic relevance of genetic alterations in diffuse lower-grade gliomas. Neuro-oncology 20(1):66–77
Weller M, Stupp R, Hegi ME, van den Bent M, Tonn JC, Sanson M, Wick W, Reifenberger G (2012) Personalized care in neuro-oncology coming of age: why we need MGMT and 1p/19q testing for malignant glioma patients in clinical practice. Neuro Oncol 14(4):100–108
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L et al (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352(10):997–1003
Molinaro AM, Taylor JW, Wiencke JK, Wrensch MR (2019) Genetic and molecular epidemiology of adult diffuse glioma. Nat Rev Neurol 15(7):405–417
Iyevleva AG, Imyanitov EN (2016) Cytotoxic and targeted therapy for hereditary cancers. Hereditary cancer in clinical practice 14(1):17
Kinnersley B, Labussiere M, Holroyd A, Di Stefano A-L, Broderick P, Vijayakrishnan J, Mokhtari K, Delattre J-Y, Gousias K, Schramm J (2015) Genome-wide association study identifies multiple susceptibility loci for glioma. Nature communications 6(1):1–9
Chatrath A, Kiran M, Kumar P, Ratan A, Dutta A (2019) The Germline Variants rs61757955 and rs34988193 Are Predictive of Survival in Lower Grade Glioma Patients. Mol Cancer Res 17(5):1075–1086
Tutt A, Ellis P, Kilburn L, Gilett C, Pinder S, Abraham J, Barrett S, Barrett-Lee P, Chan S, Cheang M et al (2015) Abstract S3–01 The TNT trial A randomized phase III trial of carboplatin (C) compared with docetaxel (D) for patients with metastatic or recurrent locally advanced triple negative or BRCA1/2 breast cancer (CRUK/07/012). Can Res 75(9):S3-01-S03-01
Zang YS, Dai C, Xu X, Cai X, Wang G, Wei J, Wu A, Sun W, Jiao S, Xu Q (2019) Comprehensive analysis of potential immunotherapy genomic biomarkers in 1000 Chinese patients with cancer. Cancer Med 8(10):4699–4708
Ronellenfitsch MW, Oh JE, Satomi K, Sumi K, Harter PN, Steinbach JP, Felsberg J, Capper D, Voegele C, Durand G et al (2018) CASP9 germline mutation in a family with multiple brain tumors. Brain Pathol 28(1):94–102
Bainbridge MN, Armstrong GN, Gramatges MM, Bertuch AA, Jhangiani SN, Doddapaneni H, Lewis L, Tombrello J, Tsavachidis S, Liu Y et al (2015) Germline mutations in shelterin complex genes are associated with familial glioma. J Natl Cancer Inst 107(1):384
Jacobs DI, Fukumura K, Bainbridge MN, Armstrong GN, Tsavachidis S, Gu X, Doddapaneni HV, Hu J, Jayaseelan JC, Muzny DM et al (2018) Elucidating the molecular pathogenesis of glioma: integrated germline and somatic profiling of a familial glioma case series. Neuro Oncol 20(12):1625–1633
Muskens IS, de Smith AJ, Zhang C, Hansen HM, Morimoto L, Metayer C, Ma X, Walsh KM, Wiemels JL (2020) Germline cancer predisposition variants and pediatric glioma: a population-based study in California. Neuro Oncol 22(6):864–874
Chiarle R, Voena C, Ambrogio C, Piva R, Inghirami G (2008) The anaplastic lymphoma kinase in the pathogenesis of cancer. Nat Rev Cancer 8(1):11–23
Della Corte CM, Viscardi G, Di Liello R, Fasano M, Martinelli E, Troiani T, Ciardiello F, Morgillo F (2018) Role and targeting of anaplastic lymphoma kinase in cancer. Mol Cancer 17(1):30
Mosse YP, Laudenslager M, Longo L, Cole KA, Wood A, Attiyeh EF, Laquaglia MJ, Sennett R, Lynch JE, Perri P et al (2008) Identification of ALK as a major familial neuroblastoma predisposition gene. Nature 455(7215):930–935
Kudo K, Ueno H, Sato T, Kubo K, Kanezaki R, Kobayashi A, Kamio T, Sasaki S, Terui K, Kurose A et al (2018) Two siblings with familial neuroblastoma with distinct clinical phenotypes harboring an ALK germline mutation. Genes Chromosomes Cancer 57(12):665–669
Chaft JE, Dagogo-Jack I, Santini FC, Eng J, Yeap BY, Izar B, Chin E, Jones DR, Kris MG, Shaw AT et al (2018) Clinical outcomes of patients with resected, early-stage ALK-positive lung cancer. Lung Cancer 122:67–71
Bavi P, Jehan Z, Bu R, Prabhakaran S, Al-Sanea N, Al-Dayel F, Al-Assiri M, Al-Halouly T, Sairafi R, Uddin S et al (2013) ALK gene amplification is associated with poor prognosis in colorectal carcinoma. Br J Cancer 109(10):2735–2743
Lastowska M, Trubicka J, Niemira M, Paczkowska-Abdulsalam M, Karkucinska-Wieckowska A, Kaleta M, Drogosiewicz M, Tarasinska M, Perek-Polnik M, Kretowski A et al (2017) ALK Expression Is a Novel Marker for the WNT-activated Type of Pediatric Medulloblastoma and an Indicator of Good Prognosis for Patients. Am J Surg Pathol 41(6):781–787
Su Y, Long X, Song Y, Chen P, Li S, Yang H, Wu P, Wang Y, Bing Z, Cao Z et al (2019) Distribution of ALK Fusion Variants and Correlation with Clinical Outcomes in Chinese Patients with Non-Small Cell Lung Cancer Treated with Crizotinib. Target Oncol 14(2):159–168
Capellades J, Puig J, Domenech S, Pujol T, Oleaga L, Camins A, Majos C, Diaz R, de Quintana C, Teixidor P et al (2018) Is a pretreatment radiological staging system feasible for suggesting the optimal extent of resection and predicting prognosis in glioblastoma? An observational study J Neurooncol 137(2):367–377
Pinto N, Cohn SL, Dolan ME (2012) Using germline genomics to individualize pediatric cancer treatments. Clin Cancer Res 18(10):2791–2800
Scharfe CPI, Tremmel R, Schwab M, Kohlbacher O, Marks DS (2017) Genetic variation in human drug-related genes. Genome Med 9(1):117
Bai H, Harmanci AS, Erson-Omay EZ, Li J, Coskun S, Simon M, Krischek B, Ozduman K, Omay SB, Sorensen EA et al (2016) Integrated genomic characterization of IDH1-mutant glioma malignant progression. Nat Genet 48(1):59–66
Reifenberger G, Wirsching HG, Knobbe-Thomsen CB, Weller M (2017) Advances in the molecular genetics of gliomas - implications for classification and therapy. Nat Rev Clin Oncol 14(7):434–452
Chaft JE, Arcila ME, Paik PK, Lau C, Riely GJ, Pietanza MC, Zakowski MF, Rusch V, Sima CS, Ladanyi M et al (2012) Coexistence of PIK3CA and other oncogene mutations in lung adenocarcinoma-rationale for comprehensive mutation profiling. Mol Cancer Ther 11(2):485–491
Conde M, Michen S, Wiedemuth R, Klink B, Schrock E, Schackert G, Temme A (2017) Chromosomal instability induced by increased BIRC5/Survivin levels affects tumorigenicity of glioma cells. BMC Cancer 17(1):889
Labussiere M, Di Stefano AL, Gleize V, Boisselier B, Giry M, Mangesius S, Bruno A, Paterra R, Marie Y, Rahimian A et al (2014) TERT promoter mutations in gliomas, genetic associations and clinico-pathological correlations. Br J Cancer 111(10):2024–2032
Alidousty C, Baar T, Martelotto LG, Heydt C, Wagener S, Fassunke J, Duerbaum N, Scheel AH, Frank S, Holz B et al (2018) Genetic instability and recurrent MYC amplification in ALK-translocated NSCLC: a central role of TP53 mutations. J Pathol 246(1):67–76
Song P, Zhang F, Li Y, Yang G, Li W, Ying J, Gao S (2019) Concomitant TP53 mutations with response to crizotinib treatment in patients with ALK-rearranged non-small-cell lung cancer. Cancer Med 8(4):1551–1557
Funding
The author(s) received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by LB, NUF H and CL. The first draft of the manuscript was written by LB and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Ethical approval
The study protocol was approved by the Ethics Committee of Huashan hospital.
Consent to participate
All patients provided written informed consent.
Consent for publication
All authors consent to the publication of the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11060_2020_3676_MOESM3_ESM.xlsx
Supplementary file4 Supplementary Table 3. The damage scores of ALK variants were evaluated using PolyPhen-v2 database (XLSX 10 KB)
11060_2020_3676_MOESM4_ESM.pdf
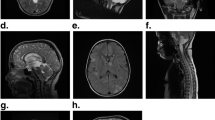
Supplementary file3 Supplementary Figure 1. The expression of ALK between gALK-Mut and gALK-WT patient’s samples. Protein level were measured in 43 patients with glioma, 5 patients with ALK variants and 38 gALK-WT patients as normal control. The scores were calculated based on the percentage of ALK positive cells, 0%-20% positive cells were presented score 0 (A-C); 21%–50% positive cells were presented score 1 (D-F); 51%-80% positive cells were presented score 2 (G-I), as well as 81%-100% positive cells were presented score 3 (J-L). Statistical analysis showed that no significant differences were observed between two groups respectively (M-N) (PDF 162 KB)
11060_2020_3676_MOESM5_ESM.tif
Supplementary file5 Supplementary Figure 2. Summary of two major germline mutated genes identified in 39 IDH-WT-GBM cases. (A) PIK3CA. (B) BIRC5. Each circle with a stem represents a unique somatic variant. The upper labels represent gALK-Mut patients and the lower labels represent gALK-WT patients (TIF 17736 KB)
Rights and permissions
About this article
Cite this article
Bu, L., Hameed, N.U.F., Luo, C. et al. Germline ALK variations are associated with a poor prognosis in glioma and IDH-wildtype glioblastoma. J Neurooncol 152, 27–36 (2021). https://doi.org/10.1007/s11060-020-03676-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03676-5