Abstract
Objectives
The most prevalent sensory disease in humans is deafness. A variety of genes have been linked to hearing loss, which can either be isolated (non-syndromic) or associated with lesions in other organs (syndromic). It has been discovered that WHRN variants are responsible for non-syndromic hearing loss and Usher syndrome type II.
Methods and results
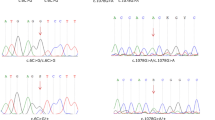
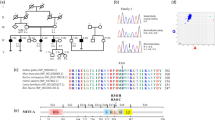
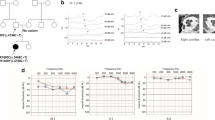
Exome sequencing in a consanguineous Moroccan patient with severe hearing loss identified a single homozygous mutation c.619G > T; p.Ala207Ser in WHRN, encoding a cytoskeletal scaffold protein that binds membrane protein complexes to the cytoskeleton in ocular photoreceptors and ear hair cell stereocilia. Bioinformatics methods and molecular dynamic modeling were able to predict the pathogenic implications of this variation.
Conclusion
We used whole exome sequencing to find a homozygous WHRN gene variant in a Moroccan family. Numerous bioinformatics methods predict that this modification might result in a change in the WHRN protein’s structure.





Similar content being viewed by others
Data Availability
Data will be provided by the authors upon request.
References
Gettelfinger JD, Dahl JP (2018) Syndromic hearing loss: a brief review of Common Presentations and Genetics. J Pediatr Genet 7:1–8. https://doi.org/10.1055/s-0037-1617454
Petrova N, Tebieva I, Kadyshev V et al (2023) Hereditary etiology of non-syndromic sensorineural hearing loss in the Republic of North Ossetia-Alania. PeerJ 11:e14514. https://doi.org/10.7717/peerj.14514
Bakhchane A, Bousfiha A, Charoute H et al (2016) Update of the spectrum of GJB2 gene mutations in 152 moroccan families with autosomal recessive nonsyndromic hearing loss. Eur J Med Genet 59:325–329. https://doi.org/10.1016/j.ejmg.2016.05.002
Aller E, Jaijo T, van Wijk E et al (2010) Sequence variants of the DFNB31 gene among Usher syndrome patients of diverse origin. Mol Vis 16:495–500
Mathur PD, Yang J (2019) Usher syndrome and non-syndromic deafness: functions of different whirlin isoforms in the cochlea, vestibular organs, and retina. Hear Res 375:14–24. https://doi.org/10.1016/j.heares.2019.02.007
Chen Q, Zou J, Shen Z et al (2014) Whirlin and PDZ domain-containing 7 (PDZD7) proteins are both required to form the quaternary protein complex associated with Usher syndrome type 2. J Biol Chem 289:36070–36088. https://doi.org/10.1074/jbc.M114.610535
Audo I, Bujakowska K, Mohand-Saïd S et al (2011) A novel DFNB31 mutation associated with usher type 2 syndrome showing variable degrees of auditory loss in a consanguineous portuguese family. Mol Vis 17:1598–1606
Can T (2014) Introduction to bioinformatics. Methods Mol Biol 1107:51–71. https://doi.org/10.1007/978-1-62703-748-8_4
Porto WF, Franco OL, Alencar SA (2015) Computational analyses and prediction of guanylin deleterious SNPs. Peptides 69:92–102. https://doi.org/10.1016/j.peptides.2015.04.013
Filipe HAL, Loura LMS (2022) Molecular Dynamics Simulations: advances and applications. Molecules 27:2105. https://doi.org/10.3390/molecules27072105
Krieger E, Vriend G (2014) YASARA View - molecular graphics for all devices - from smartphones to workstations. Bioinformatics 30:2981–2982. https://doi.org/10.1093/bioinformatics/btu426
Pires DEV, Ascher DB, Blundell TL (2014) mCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 30:335–342. https://doi.org/10.1093/bioinformatics/btt691
Pandurangan AP, Ochoa-Montaño B, Ascher DB, Blundell TL (2017) SDM: a server for predicting effects of mutations on protein stability. Nucleic Acids Res 45:W229–W235. https://doi.org/10.1093/nar/gkx439
Pires DEV, Ascher DB, Blundell TL (2014) DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res 42:W314–319. https://doi.org/10.1093/nar/gku411
Ashkenazy H, Erez E, Martz E et al (2010) ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res 38:W529–533. https://doi.org/10.1093/nar/gkq399
K L-L SP, K P, et al (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78. https://doi.org/10.1002/prot.22711
Mathur P, Yang J (2015) Usher syndrome: hearing loss, retinal degeneration and associated abnormalities. Biochim Biophys Acta 1852:406–420. https://doi.org/10.1016/j.bbadis.2014.11.020
Mburu P, Mustapha M, Varela A et al (2003) Defects in whirlin, a PDZ domain molecule involved in stereocilia elongation, cause deafness in the whirler mouse and families with DFNB31. Nat Genet 34:421–428. https://doi.org/10.1038/ng1208
Tlili A, Charfedine I, Lahmar I et al (2005) Identification of a novel frameshift mutation in the DFNB31/WHRN gene in a tunisian consanguineous family with hereditary non-syndromic recessive hearing loss. Hum Mutat 25:503. https://doi.org/10.1002/humu.9333
Ebermann I, Scholl HPN, Charbel Issa P et al (2007) A novel gene for Usher syndrome type 2: mutations in the long isoform of whirlin are associated with retinitis pigmentosa and sensorineural hearing loss. Hum Genet 121:203–211. https://doi.org/10.1007/s00439-006-0304-0
G M, G I, S G, et al (2017) Usher’s syndrome type II: a comparative study of genetic mutations and vestibular system evaluation. Otolaryngology–head and neck Surgery: Official Journal of American Academy of Otolaryngology-Head and Neck Surgery 157:. https://doi.org/10.1177/0194599817715235
Yang J, Liu X, Zhao Y et al (2010) Ablation of Whirlin Long Isoform disrupts the USH2 protein complex and causes Vision and hearing loss. PLoS Genet 6:e1000955. https://doi.org/10.1371/journal.pgen.1000955
Acknowledgements
The authors are indebted to the families who contributed to this study. This project was supported by the Institut Pasteur du Maroc (IPM).
Funding
No funds, grants, or other support was received.
Author information
Authors and Affiliations
Contributions
All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The genetic study was approved by the ethics committee for the biomedical research of Rabat (N° 09/19).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
AitRaise, I., Amalou, G., Redouane, S. et al. Novel pathogenic WHRN variant causing hearing loss in a moroccan family. Mol Biol Rep 50, 10663–10669 (2023). https://doi.org/10.1007/s11033-023-08901-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08901-8