Abstract
Background
Melanosomes are lysosome-related organelles that contain melanogenic factors and synthesize melanin as they mature. FYVE finger-containing phosphoinositide kinase (PIKfyve) regulates late endosome and lysosome morphology, vesicle trafficking, and autophagy. In melanocytes, PIKfyve inhibition has been reported to induce hypopigmentation due to impairments in the metabolism of early-stage melanosomes.
Methods and results
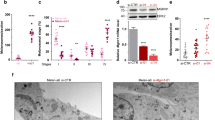
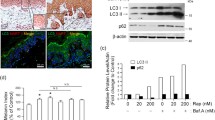
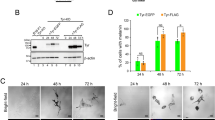
Here, we report a new type of melanosome metabolism: post-PIKfyve inhibition, which was found during the characterization of the endosome/lysosome fluoroprobe GIF-2250. In B16F10 mouse melanoma cells, GIF-2250 highlighted vesicles positive for lysosomal-associated membrane protein 1 (lysosome marker) and other endosome/lysosome markers (CD63 and Rab7/9). When cells were continuously treated with PIKfyve inhibitors, intracellular vacuoles formed, while GIF-2250 fluorescence signals diminished and were diffusely distributed in the vacuoles. After removal of the PIKfyve inhibitors, the GIF-2250 signal intensity was restored, and some GIF-2250-positive vesicles wrapped the melanosomes, which spun at high speed. In addition, intermittent PIKfyve inhibition caused melanin diffusion in the vacuoles and possible leakage into the cytoplasmic compartments, and melanosome degradation was detected by a transmission electron microscope. Melanosome degradation was accompanied by decreased levels of melanin synthesis enzymes and increased levels of the autophagosome maker LC3BII, which is also associated with early melanosomes. However, the protein levels of p62, which is degraded during autophagy, were increased, suggesting an impairment in autophagy flux during intermittent PIKfyve inhibition. Moreover, the autophagy inhibitor 3-methyladenine does not affect these protein levels, suggesting that the melanosome degradation by the intermittent inhibition of PIKfyve is not mediated by canonical autophagy.
Conclusions
In conclusion, disturbance of PIKfyve activity induces melanosome degradation in a canonical autophagy-independent manner.






Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request. The original blots of western analyses are in Supplementary Materials.
References
Raposo G, Marks MS (2007) Melanosomes–dark organelles enlighten endosomal membrane transport. Nat Rev Mol Cell Biol 8:786–797. https://doi.org/10.1038/nrm2258
Bowman SL, Bi-Karchin J, Le L, Marks MS (2019) The road to lysosome-related organelles: insights from Hermansky–Pudlak syndrome and other rare diseases. Traffic 20:404–435. https://doi.org/10.1111/tra.12646
Sitaram A, Marks MS (2012) Mechanisms of protein delivery to melanosomes in pigment cells. Physiology (Bethesda) 27:85–99. https://doi.org/10.1152/physiol.00043.2011
Le L, Sirés-Campos J, Raposo G, Delevoye C, Marks MS (2021) Melanosome biogenesis in the pigmentation of mammalian skin. Integr Comp Biol 61:1517–1545. https://doi.org/10.1093/icb/icab078
Fukuda M (2021) Rab GTPases: key players in melanosome biogenesis, transport, and transfer. Pigment Cell Melanoma Res 34:222–235. https://doi.org/10.1111/pcmr.12931
Yang C, Wang X (2021) Lysosome biogenesis: regulation and functions. J Cell Biol. https://doi.org/10.1083/jcb.202102001
Berson JF, Theos AC, Harper DC, Tenza D, Raposo G, Marks MS (2003) Proprotein convertase cleavage liberates a fibrillogenic fragment of a resident glycoprotein to initiate melanosome biogenesis. J Cell Biol 161:521–533. https://doi.org/10.1083/jcb.200302072
van Niel G, Charrin S, Simoes S et al (2011) The tetraspanin CD63 regulates ESCRT-independent and-dependent endosomal sorting during melanogenesis. Dev Cell 21:708–721. https://doi.org/10.1016/j.devcel.2011.08.019
Raposo G, Tenza D, Murphy DM, Berson JF, Marks MS (2001) Distinct protein sorting and localization to premelanosomes, melanosomes, and lysosomes in pigmented melanocytic cells. J Cell Biol 152:809–824. https://doi.org/10.1083/jcb.152.4.809
Yang Y, Jang GB, Yang X et al (2018) Central role of autophagic UVRAG in melanogenesis and the suntan response. Proc Natl Acad Sci 115:E7728-e7737. https://doi.org/10.1073/pnas.1803303115
Lamark T, Johansen T (2021) Mechanisms of selective autophagy. Annu Rev Cell Dev Biol 37:143–169. https://doi.org/10.1146/annurev-cellbio-120219-035530
Ho H, Ganesan AK (2011) The pleiotropic roles of autophagy regulators in melanogenesis. Pigment Cell Melanoma Res 24:595–604. https://doi.org/10.1111/j.1755-148X.2011.00889.x
Ramkumar A, Murthy D, Raja DA et al (2017) Classical autophagy proteins LC3B and ATG4B facilitate melanosome movement on cytoskeletal tracks. Autophagy 13:1331–1347. https://doi.org/10.1080/15548627.2017.1327509
Klionsky DJ, Abdel-Aziz AK, Abdelfatah S et al (2021) Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition) (1). Autophagy 17:1–382. https://doi.org/10.1080/15548627.2020.1797280
Lee KW, Kim M, Lee SH, Kim KD (2022) The function of autophagy as a regulator of melanin homeostasis. Cells 11:2085. https://doi.org/10.3390/cells11132085
Sbrissa D, Ikonomov OC, Shisheva A (1999) PIKfyve, a mammalian ortholog of yeast Fab1p lipid kinase, synthesizes 5-phosphoinositides: effect of insulin. J Biol Chem 274:21589–21597. https://doi.org/10.1074/jbc.274.31.21589
de Lartigue J, Polson H, Feldman M et al (2009) PIKfyve regulation of endosome-linked pathways. Traffic 10:883–893. https://doi.org/10.1111/j.1600-0854.2009.00915.x
Liggins MC, Flesher JL, Jahid S et al (2018) PIKfyve regulates melanosome biogenesis. PLoS Genet 14:e1007290. https://doi.org/10.1371/journal.pgen.1007290
Bissig C, Croisé P, Heiligenstein X et al (2019) The PIKfyve complex regulates the early melanosome homeostasis required for physiological amyloid formation. J Cell Sci 132:jcs229500. https://doi.org/10.1242/jcs.229500
Hirata Y, Tsunekawa Y, Takahashi M et al (2021) Identification of novel neuroprotective N,N-dimethylaniline derivatives that prevent oxytosis/ferroptosis and localize to late endosomes and lysosomes. Free Radic Biol Med 174:225–235. https://doi.org/10.1016/j.freeradbiomed.2021.08.015
Maeda M, Suzuki M, Takashima S, Sasaki T, Oh-Hashi K, Takemori H (2021) The new live imagers MitoMM1/2 for mitochondrial visualization. Biochem Biophys Res Commun 562:50–54. https://doi.org/10.1016/j.bbrc.2021.05.040
Isogawa K, Asano M, Hayazaki M et al (2021) Thioxothiazolidin derivative, 4-OST, inhibits melanogenesis by enhancing the specific recruitment of tyrosinase-containing vesicles to lysosome. J Cell Biochem 122:667–678. https://doi.org/10.1002/jcb.29895
Watanabe M, Kawaguchi K, Nakamura Y, Furuta K, Takemori H (2021) GIF-2209, an oxindole derivative, accelerates melanogenesis and melanosome secretion via the modification of lysosomes in B16F10 mouse melanoma cells. Molecules 27:177. https://doi.org/10.3390/molecules27010177
Recalcati S, Menotti E, Kühn LC (2001) Peroxisomal targeting of mammalian hydroxyacid oxidase 1 requires the C-terminal tripeptide SKI. J Cell Sci 114:1625–1629. https://doi.org/10.1242/jcs.114.9.1625
Kanamori H, Takemura G, Goto K et al (2015) Autophagic adaptations in diabetic cardiomyopathy differ between type 1 and type 2 diabetes. Autophagy 11:1146–1160. https://doi.org/10.1080/15548627.2015.1051295
Cao XJ, Chen LN, Zhang X et al (2016) A NBD-based simple but effective fluorescent pH probe for imaging of lysosomes in living cells. Anal Chim Acta 920:86–93. https://doi.org/10.1016/j.aca.2016.03.029
Takemori H, Koga K, Kawaguchi K et al (2022) Visualization of mitophagy using LysoKK, a 7-nitro-2,1,3-benzoxadiazole-(arylpropyl)benzylamine derivative. Mitochondrion 62:176–180. https://doi.org/10.1016/j.mito.2021.12.004
Jefferies HB, Cooke FT, Jat P et al (2008) A selective PIKfyve inhibitor blocks PtdIns(3,5)P(2) production and disrupts endomembrane transport and retroviral budding. EMBO Rep 9:164–170. https://doi.org/10.1038/sj.embor.7401155
Giordano F, Bonetti C, Surace EM, Marigo V, Raposo G (2009) The ocular albinism type 1 (OA1) G-protein-coupled receptor functions with MART-1 at early stages of melanogenesis to control melanosome identity and composition. Hum Mol Genet 18:4530–4545. https://doi.org/10.1093/hmg/ddp415
Maulucci G, Chiarpotto M, Papi M, Samengo D, Pani G, Spirito MD (2015) Quantitative analysis of autophagic flux by confocal pH-imaging of autophagic intermediates. Autophagy 11:1905–1916. https://doi.org/10.1080/15548627.2015.1084455
Karabiyik C, Rubinsztein DC (2021) AMPK-activated ULK1 phosphorylates PIKFYVE to drive formation of PtdIns5P-containing autophagosomes during glucose starvation. Autophagy 17:3877–3878. https://doi.org/10.1080/15548627.2021.1961409
Buckley CM, Heath VL, Guého A et al (2019) PIKfyve/Fab1 is required for efficient V-ATPase and hydrolase delivery to phagosomes, phagosomal killing, and restriction of Legionella infection. PLoS Pathog 15:e1007551. https://doi.org/10.1371/journal.ppat.1007551
Isobe Y, Nigorikawa K, Tsurumi G et al (2019) PIKfyve accelerates phagosome acidification through activation of TRPML1 while arrests aberrant vacuolation independent of the Ca2+ channel. J Biochem 165:75–84. https://doi.org/10.1093/jb/mvy084
Kuchitsu Y, Mukai K, Uematsu R et al (2023) STING signalling is terminated through ESCRT-dependent microautophagy of vesicles originating from recycling endosomes. Nat Cell Biol 25:453–466. https://doi.org/10.1038/s41556-023-01098-9
Maciąg D, Dobrowolska E, Sharafan M, Ekiert H, Tomczyk M, Szopa A (2021) Akebia quinata and Akebia trifoliata—a review of phytochemical composition, ethnopharmacological approaches and biological studies. J Ethnopharmacol 280:114486. https://doi.org/10.1016/j.jep.2021.114486
Peng P, Jia D, Cao L et al (2021) Akebia saponin E, as a novel PIKfyve inhibitor, induces lysosome-associated cytoplasmic vacuolation to inhibit proliferation of hepatocellular carcinoma cells. J Ethnopharmacol 266:113446. https://doi.org/10.1016/j.jep.2020.113446
Kim JY, Kim J, Ahn Y et al (2020) Autophagy induction can regulate skin pigmentation by causing melanosome degradation in keratinocytes and melanocytes. Pigment Cell Melanoma Res 33:403–415. https://doi.org/10.1111/pcmr.12838
Zhou D, Ota K, Nardin C et al (2018) Mammalian pigmentation is regulated by a distinct cAMP-dependent mechanism that controls melanosome pH. Sci Signal 11:987. https://doi.org/10.1126/scisignal.aau7987
Koh JY, Kim HN, Hwang JJ, Kim YH, Park SE (2019) Lysosomal dysfunction in proteinopathic neurodegenerative disorders: possible therapeutic roles of cAMP and zinc. Mol Brain 12:18. https://doi.org/10.1186/s13041-019-0439-2
Acknowledgements
We thank Ms. Akiko Tsujimoto and Mr. Yasuaki Hotta for their support of the TEM analyses.
Funding
This work was supported by the Japan Society for the Promotion of Science (JSPS) (Grant No. 20K05703); Japan Science and Technology Agency (JST) A-Step (Grant No. JPMJTM20E8); START University Ecosystem Promotion Type (Supporting Creation of Startup Ecosystem in Startup Cities) (Grant No. JPMJST2183), Japan; the AIST-Gifu Univ. Joint Project; the Nakatani Foundation; the Ogawa Foundation; and the Kobayashi Foundation.
Author information
Authors and Affiliations
Contributions
Experiments were performed by KK, MW, SF, HK, MJI, ST, and MMK GIF-2250 was synthesized by K and KF. Data analysis was performed by KO-H and YH. The manuscript was written by HT. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The title compound is protected by PCT application no. PCT/JP2022/1692, submitted by the authors of this paper. The authors have no other relevant financial or non-financial interests to declare.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Supplementary material 3 (MOV 4202.6 kb)
Supplementary material 4 (MP4 5019.5 kb)
Supplementary material 5 (MOV 8720.6 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kawaguchi, K., Watanabe, M., Furukawa, S. et al. Intermittent inhibition of FYVE finger-containing phosphoinositide kinase induces melanosome degradation in B16F10 melanoma cells. Mol Biol Rep 50, 5917–5930 (2023). https://doi.org/10.1007/s11033-023-08536-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08536-9