Abstract
Background
Biocides are frequently used as preservative, disinfectant and sterilizer against many microorganisms in hospitals, industry and home. However, the reduced susceptibility rate of Pseudomonas aeruginosa (P. aeruginosa) strains to biocides is increasing. The aim of this study was to evaluate the antimicrobial activity of four frequently used biocides against P. aeruginosa and to determine the prevalence of genes involved in biocide resistance.
Methods
A total of 76 clinical isolates of P. aeruginosa strains were used in the present study. The minimum inhibitory concentrations (MICs) of four biocides, i.e. chlorhexidine digluconate, benzalkonium chloride, triclosan and formaldehyde, against P. aeruginosa strains were determined using agar dilution method. In addition, the prevalence of biocide resistance genes was determined using the polymerase chain reaction (PCR) method.
Results
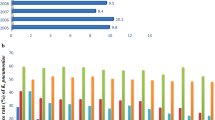
In the present study, the highest MIC90 and MIC95 (epidemiological cut-off) values were observed for benzalkonium chloride (1024 μg/mL), followed by formaldehyde (512 μg/mL), triclosan (512 μg/mL) and chlorhexidine digluconate (64 μg/mL). Furthermore, the prevalence of qacEΔ1, qacE, qacG, fabV, cepA and fabI genes were 73.7% (n = 56), 26.3% (n = 20), 11.8% (n = 9), 84.2% (n = 64), 81.5% (n = 62) and 0% (n = 0), respectively. A significant association was observed between the presence of biocide resistance genes and MICs (p < 0.05). Furthermore, there was no significant association between the presence of biocide resistance genes and antibiotic resistance (p > 0.05), except for levofloxacin and norfloxacin antibiotics and qacE and qacG genes (p < 0.05).
Conclusion
Our results revealed that chlorhexidine digluconate is the most effective biocide against P. aeruginosa isolates in Ardabil hospitals. However, we recommend continuous monitoring of the antimicrobial activity of biocides and the prevalence of biocide-associated resistance genes for a better prevention of microorganism dissemination and infection control in hospitals.
Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on reasonable request.
Abbreviations
- MIC:
-
Minimum inhibitory concentration
- PCR:
-
Polymerase chain reaction
- P. aeruginosa :
-
Pseudomonas aeruginosa
- ICU:
-
Intensive care unit
- MDR:
-
Multi-drug resistant
- CLSI:
-
Clinical and Laboratory Standards Institute
- ECOFF:
-
Epidemiological cut-off value
- ENR:
-
Enoyl-acyl-carrier protein reductase
References
Driscoll JA, Brody SL, Kollef MH (2007) The epidemiology, pathogenesis and treatment of Pseudomonas aeruginosa infections. Drugs 67(3):351–368
Vaez H, Salehi-Abargouei A, Khademi F (2017) Systematic review and meta-analysis of imipenem-resistant Pseudomonas aeruginosa prevalence in Iran. Germs 7(2):86–97
Moradali MF, Ghods S, Rehm BH (2017) Pseudomonas aeruginosa lifestyle: a paradigm for adaptation, survival, and persistence. Front Cell Infect Microbiol 7:39
Ignak S, Nakipoglu Y, Gurler B (2017) Frequency of antiseptic resistance genes in clinical staphycocci and enterococci isolates in Turkey. Antimicrob Resist Infect Control 6(1):1–7
Vásquez-Giraldo DF, Libreros-Zúñiga GA, Crespo-Ortiz MD (2017) Effects of biocide exposure on P. aeruginosa, E. coli and A. baumannii complex isolates from hospital and household environments. Infection 21(4):243–50
Babenko D, Turmuhambetova A, Sandle T, Pestrea SA, Moraru D, CHEŞCĂ A (2017) In silico comparison of different types of MLVA with PFGE based on Pseudomonas aeruginosa genomes. Acta Medica 33:347–352
Wesgate R, Grasha P, Maillard J-Y (2016) Use of a predictive protocol to measure the antimicrobial resistance risks associated with biocidal product usage. Am J Infect Control 44(4):458–464
Ortega-Morente E, Fernández-Fuentes MA, Grande-Burgos MJ, Abriouel H, Pérez-Pulido R, Gálvez A (2013) Biocide tolerance in bacteria. Int J Food Microbiol 162(1):13–25
Murray PR, Rosenthal KS, Pfaller MA (2015) Medical microbiology, 8th edn. Elsevier Health Sciences, UK, pp 272–277
Bridier A, Dubois-Brissonnet F, Greub G, Thomas V, Briandet R (2011) Dynamics of the action of biocides in Pseudomonas aeruginosa biofilms. Antimicrob Agents Chemother 55(6):2648–2654
Mima T, Joshi S, Gomez-Escalada M, Schweizer HP (2007) Identification and characterization of TriABC-OpmH, a triclosan efflux pump of Pseudomonas aeruginosa requiring two membrane fusion proteins. J Bacteriol 189(21):7600–7609
Vikram A, Bomberger JM, Bibby KJ (2015) Efflux as a glutaraldehyde resistance mechanism in Pseudomonas fluorescens and Pseudomonas aeruginosa biofilms. Antimicrob Agents Chemother 59(6):3433–3440
Chuanchuen R, Beinlich K, Hoang TT, Becher A, Karkhoff-Schweizer RR, Schweizer HP (2001) Cross-resistance between triclosan and antibiotics in Pseudomonas aeruginosa is mediated by multidrug efflux pumps: exposure of a susceptible mutant strain to triclosan selects nfxB mutants overexpressing MexCD-OprJ. Antimicrob Agents Chemother 45(2):428–432
Subedi D, Vijay AK, Willcox M (2018) Study of disinfectant resistance genes in ocular isolates of Pseudomonas aeruginosa. Antibiotics 7(4):88
CLSI, 2018. Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Eighth Informational Supplement. CLSI Document M100. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania, USA.
Bazghandi SA, Safarirad S, Arzanlou M, Peeri-Dogaheh H, AliMohammadi H, Khademi F (2021) Prevalence of multidrug-resistant Pseudomonas aeruginosa strains in Ardabil. J Ardabil Univ Med Sci 20(2):280–286
Khademi F, Ashrafi SS, Neyestani Z, Vaez H, Sahebkar A (2021) Prevalence of class I, II and III integrons in multidrug-resistant and carbapenem-resistant Pseudomonas aeruginosa clinical isolates. Gene Rep 25:101407
Delarampour A, Ghalehnoo ZR, Khademi F, Delarampour M, Vaez H (2019) Molecular detection of carbapenem-resistant genes in clinical isolates of Klebsiella pneumoniae. Ann Ig 31(4):349–355
Rizzotti L, Rossi F, Torriani S (2016) Biocide and antibiotic resistance of Enterococcus faecalis and Enterococcus faecium isolated from the swine meat chain. Food Microbiol 60:160–164
Chapman JS (2003) Biocide resistance mechanisms. Int Biodeterior. Biodegradation 51(2):133–138
Romão C, Miranda CA, Silva J, Clementino MM, de Filippis I, Asensi M (2011) Presence of qacEΔ1 gene and susceptibility to a hospital biocide in clinical isolates of Pseudomonas aeruginosa resistant to antibiotics. Curr Microbiol 63(1):16–21
Zhu L, Lin J, Ma J, Cronan JE, Wang H (2010) Triclosan resistance of Pseudomonas aeruginosa PAO1 is due to FabV, a triclosan-resistant enoyl-acyl carrier protein reductase. Antimicrob Agents Chemother 54(2):689–698
Heath RJ, Rock CO (2000) A triclosan-resistant bacterial enzyme. Nature 406(6792):145–146
Huang YH, Lin JS, Ma JC, Wang HH (2016) Functional characterization of triclosan-resistant enoyl-acyl-carrier protein reductase (FabV) in Pseudomonas aeruginosa. Front Microbiol 7:1903
Thomas L, Maillard JY, Lambert RJ, Russell AD (2000) Development of resistance to chlorhexidine diacetate in Pseudomonas aeruginosa and the effect of a “residual” concentration. J Hosp Infect 46(4):297–303
Hassan KA, Liu Q, Henderson PJ, Paulsen IT (2015) Homologs of the Acinetobacter baumannii AceI transporter represent a new family of bacterial multidrug efflux systems. MBio 6(1):e01982-e2014
Mendes ET, Ranzani OT, Marchi AP, da Silva MT, Amigo Filho JU, Alves T, Guimarães T, Levin AS, Costa SF (2016) Chlorhexidine bathing for the prevention of colonization and infection with multidrug-resistant microorganisms in a hematopoietic stem cell transplantation unit over a 9-year period: Impact on chlorhexidine susceptibility. Medicine 95(46):1–8
Vijayakumar R, Sandle T, Al-Aboody MS, Alfonaisan MK, Alturaiki W, Mickymaray S, Premanathan M, Alsagaby SA (2018) Distribution of biocide resistant genes and biocides susceptibility in multidrug-resistant Klebsiella pneumoniae, Pseudomonas aeruginosa and Acinetobacter baumannii—A first report from the Kingdom of Saudi Arabia. J Infect Public Health 11(6):812–816
Kücken D, Feucht HH, Kaulfers PM (2000) Association of qacE and qacEΔ1 with multiple resistance to antibiotics and antiseptics in clinical isolates of Gram-negative bacteria. FEMS Microbiol Lett 183(1):95–98
Helal ZH, Khan MI (2015) QacE and QacEΔ1 Genes and Their Correlation to Antibiotics and Biocides Resistance Pseudomonas aeruginosa. Am J Biomed Sci 7(2):52–62
Mahzounieh M, Khoshnood S, Ebrahimi A, Habibian S, Yaghoubian M (2014) Detection of antiseptic-resistance genes in Pseudomonas and Acinetobacter spp. isolated from burn patients. Jundishapur J Nat Pharm Prod 9(2):e15402
McDonnell G, Russell AD (1999) Antiseptics and disinfectants: activity, action, and resistance. Clin Microbiol Rev 12(1):147–179
Acknowledgements
The authors would like to acknowledge the Vice Chancellor for Research and Technology, Ardabil University of Medical Sciences, Ardabil, Iran, due to financial support.
Funding
This research was supported by Ardabil University of Medical Sciences, Iran (Grant Number: 1005761).
Author information
Authors and Affiliations
Contributions
MN, SS and SAB collected the data. FK and HV analyzed the data and led the writing of the manuscript. SH, MA and AS revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
This research was approved by the Research Ethics Committee of Ardabil University of Medical Sciences.
Consent to participate
Informed written consent was given to subjects from whom the samples were obtained for this study.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Namaki, M., Habibzadeh, S., Vaez, H. et al. Prevalence of resistance genes to biocides in antibiotic-resistant Pseudomonas aeruginosa clinical isolates. Mol Biol Rep 49, 2149–2155 (2022). https://doi.org/10.1007/s11033-021-07032-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-07032-2