Abstract
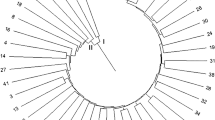
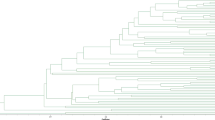
White clover (Trifolium repens L.) is an important perennial legume forage with high productivity and quality. To strengthen the basic research on the genetic characteristics, fingerprint identification and adaptability of white clover germplasm resources, Simple sequence repeat (SSR) molecular markers were applied to 10 white clover cultivars to assess the genetic diversity and related lines of white clover at the molecular level in order to lay a theoretical foundation for the selection of high-quality seeds and cultivars of white clover. A total of 120 different bands were amplified by 29 pairs of SSR primers with good polymorphism, of which 103 (89.5%) were polymorphic. Meanwhile, the PIC of each primer was 0.181–0.588, with an average of 0.329. Analysis of molecular variance revealed that 57% of the genetic variation occurred within cultivars and 43% occurred among cultivars. The results of cluster analysis and the principal coordinate analysis revealed that the parental relationships of the 10 cultivars, with the ‘Purple’ cultivar very distantly related to the other 9 cultivars and the closest parental relationship between ‘Ladino’ and ‘Sulky’. The fingerprints constructed by three representative primers (gtrs679, gtrs319, and gtrs678) have a strong identification ability. In summary, the SSR markers had good polymorphism and could be used for DNA fingerprint analysis of white clover cultivars.





Similar content being viewed by others
References
Griffiths AG, Barrett BA, Simon D, Khan AK, Bickerstaff P, Anderson CB, Franzmayr BK, Hancock KR, Jones CS (2013) An integrated genetic linkage map for white clover (Trifolium repens L.) with alignment to Medicago. BMC Genomics 14(1):388
Jones ES, Hughes LJ, Drayton MC, Abberton MT, Michaelson-Yeates TPT, Bowen C, Forster JW (2003) An SSR and AFLP molecular marker-based genetic map of white clover (Trifolium repens L.). Plant Sci 165(3):531–539
Martínez LE, Cavagnaro PF, Masuelli RW, Zu´n˜iga M (2006) SSR-based assessment of genetic diversity in South American Vitis vinifera varieties. Plant Science 170 (6):0–1044
Brewbaker JL (1954) Incompatibility in Autotetraploid trifolium Repens L. I. Competition and self-compatibility. Genetics 39(3):307–316
Lucas J, Forster J, Smith K, Spangenberg G (2012) Assessment of gene flow in white clover (Trifolium repens L.) under field conditions in Australia using phenotypic and genetic markers. Crop Pasture Sci 63:155–163
Nie G, Ting H, Xiao M, Linkai H, Yan P, Yanhong Y, Zhou L, Xia W, Xinquan Z (2019) Genetic variability evaluation and cultivar identification of tetraploid annual ryegrass using SSR markers. PeerJ 7:e7742. https://doi.org/10.7717/peerj.7742
Humberto RVM, Amalio SV, Octavio M, June S, Corina HK, Celso CR, Zhang T (2013) Analysis and optimization of bulk DNA sampling with binary scoring for germplasm characterization. PLoS ONE 8(11):e79936
Badr A, El-Shazly HH, Mekki L (2012) Genetic diversity in white clover and its progenitors as revealed by DNA fingerprinting. Biol Plant 56(2):283–291
Scarano D, Rao R, Masi P, Corrado G (2015) SSR fingerprint reveals mislabeling in commercial processed tomato products. Food Control 51:397–401
Liu S, An Y, Li F, Li S, Wei C (2018) Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers for genetic studies in tea plant (Camellia sinensis). Mol Breed. https://doi.org/10.1007/s11032-018-0824-z
Tenzer I, Ivanissevich SD, Morgante M, Gessler C (1999) Identification of microsatellite markers and their application to population genetics of Venturia inaequalis. Phytopathology 89(9):748–753
Siew GY, Ng WL, Tan SW, Alitheen NB, Tan SG, Yeap SK (2018) Genetic variation and DNA fingerprinting of durian types in Malaysia using simple sequence repeat (SSR) markers. PeerJ 6:e4266. https://doi.org/10.7717/peerj.4266
Laidò G, Mangini G, Taranto F, Gadaleta A, Blanco A, Cattivelli L, Marone D, Mastrangelo AM, Papa R, Vita PD (2013) Genetic diversity and population structure of tetraploid wheats (Triticum turgidum L.) estimated by SSR, DArT and pedigree data. PLoS ONE 8:e67280
Xie W, Zhang X, Cai H, Huang L, Ma X (2011) Genetic maps of SSR and SRAP markers in diploid orchardgrass (Dactylis glomerata L.) using the pseudo-testcross strategy. Genome 54(3):212–221
Kölliker R, Jones ES, Jahufer MZZ, Forster JW (2001) Bulked AFLP analysis for the assessment of genetic diversity in white clover (Trifolium repens L.). Euphytica 121(3):305–315
Kumar M, Luthra OP, Chawla V, Chaudhary L, Saini N, Poonia A, Kumar R, Singh AP (2006) Identification of RAPD markers linked to the karnal bunt resistance genes in wheat. Biol Plant 50(4):755–758
Forster JW, Jones ES, Kölliker R, Drayton MC, Dupal MP, Guthridge KM, Smith KF, Henry RJ (2001) Application of DNA profiling to an outbreeding forage species.
Tehrani MS, Mardi M, Sahebi J, Catalán P, Díaz-Pérez A (2009) Genetic diversity and structure among Iranian tall fescue populations based on genomic-SSR and EST-SSR marker analysis. Plant Syst Evol 282(1–2):57–70
Jiang L, Zhang XQ, Ma X, Huang LK, Xie WG, Ma YM, Zhao YF (2013) Identification of orchardgrass (Dactylis glomerata L.) cultivars by using simple sequence repeat markers. Genetics Mol Res 12(4):5111–5123
Herrmann D, Flajoulot S, Julier B (2010) Sample size for diversity studies in tetraploid alfalfa (Medicago sativa) based on codominantly coded SSR markers. Euphytica 171(3):441–446
Kongkiatngam P, Waterway MJ, Coulman BE, Fortin MG (1996) Genetic variation among cultivars of red clover (Trifolium pretense L.) detected by RAPD markers amplified from bulk genomic DNA. Euphytica 89(3):355–361
Chandra A (2007) Bulk genomic DNA PCR analysis—a rapid method to estimate genetic relatedness among heterogeneous lucerne (Medicago sativa) cultivars. Cytologia 72(3):363–368
Coleman JS, McConnaughay KDM, Ackerly DD (1994) Interpreting phenotypic variation in plants. Trends Ecol Evol 9(5):187–191. https://doi.org/10.1016/0169-5347(1094)90087-90086
Xian Z, Zhang YJ, Rong Y, Han JG, Fuzeng H, Wang JH, Kai C (2010) Genetic variation of white clover (Trifolium repens L.) collections from China detected by morphological traits, RAPD and SSR. Afr J Biotechnol 9(21):3032–3041
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distance among DNA haplotypes: application to human mitochondria DNA restriction sites. Genetics 131:479–491
Anand D, Prabhu KV, Singh AK (2012) Analysis of molecular diversity and fingerprinting of commercially grown Indian rice hybrids. J Plant Biochem Biotechnol 21(2):173–179
Zhang Q, Li J, Zhao Y, Korban SS, Han Y (2012) Evaluation of genetic diversity in chinese wild apple species along with apple cultivars using SSR markers. Plant Mol Biol Rep 30(3):539–546
Erfani J, Ebadi A, Abdollahi H, Fatahi R (2012) Genetic diversity of some pear cultivars and genotypes using simple sequence repeat (SSR) markers. Plant Mol Biol Rep 30(5):1065–1072
Karaagac E, Yilma S, Cuesta-Marcos A, Vales MI (2014) Molecular analysis of potatoes from the Pacific Northwest tri-state variety development program and selection of markers for practical DNA fingerprinting applications. Am J Potato Res 91(2):195–203
Chen LQ, Shi SL (2015) Genetic diversity of 42 alfalfa accessions revealed by SSR markers. Pratacult Sci 32(03):372–381
Gu XY, Guo Z-h, Ma X, Bai S-q, Zhang X-q, Zhang C-b, Chen S-y, Peng Y, Yan Y-h, Huang L-k (2015) Population genetic variability and structure of Elymus breviaristatus (Poaceae: Triticeae) endemic to Qinghai-Tibetan Plateau inferred from SSR markers. Biochem Syst Ecol 58:247–256
Hameed U, Pan YB, Muhammad K, Afghan S, Iqbal J (2012) Use of simple sequence repeat markers for DNA fingerprinting and diversity analysis of sugarcane (Saccharum spp) cultivars resistant and susceptible to red rot. Genet Mol Res 11(2):1195–1204
Ghérardi M, Mangin B, Goffinet B, Bonnet D, Huguet T (1998) A method to measure genetic distance between allogamous populations of alfalfa (Medicago sativa) using RAPD molecular markers. Theoret Appl Genet 96(3–4):406–412
Ulloa O, Ortega F, Campos H (2003) Analysis of genetic diversity in red clover (Trifolium pratenser, L.) breeding populations as revealed by RAPD genetic markers. Genome 46(4):529–535
Larson SR, Jones TA, Jensen KB (2004) Population structure in Pseudoroegneria spicata (Poaceae: Triticeae) modeled by Bayesian clustering of AFLP genotypes. Am J Bot 91(11):1789–1801
Yu K, Pauls KP (1993) Rapid estimation of genetic relatedness among heterogeneous populations of alfalfa by random amplification of bulked genomic DNA samples. Theoret Appl Genet 86(6):788–794
Dubreuil P, Warburton M, Chastanet M, Hoisington D, Charcosset A (2006) More on the introduction of temperate maize into Europe: Large-scale bulk SSR genotyping and new historical elements. Maydica 51(2):281–291
Kubik C, Sawkins M, Meyer WA, Gaut BS (2001) Genetic diversity in seven perennial ryegrass (Lolium perenne L.) cultivars based on SSR markers. Crop Sci 41:1565–1572
Etten JV, López MRF, Monterroso LGM, Samayoa KMP (2008) Genetic diversity of maize (Zea mays L. ssp. mays ) in communities of the western highlands of Guatemala: geographical patterns and processes. Genet Resour Crop Evol 55(2):303–317
Funding
This study was funded by Modern Agro-industry Technology Research System (CARS-34), the Grassland Basic Resources Investigation Research in China (2017FY100602).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by XZ, GN, SM and CH. The first draft of the manuscript was written by SM, CH and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Sainan Ma and Chongyang Han are the co-first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ma, S., Han, C., Zhou, J. et al. Fingerprint identification of white clover cultivars based on SSR molecular markers. Mol Biol Rep 47, 8513–8521 (2020). https://doi.org/10.1007/s11033-020-05893-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05893-7