Abstract
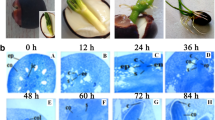
The formation of adventitious roots (ARs) is a key morphological adaptation of cucumber (Cucumis sativus L.) to waterlogging stress. MicroRNAs (miRNAs) constitute a group of non-coding small RNAs (sRNA) that play crucial roles in regulating diverse biological processes, including waterlogging acclimation. However, which specific miRNAs and how they are involved in waterlogging-triggered de novo AR primordia formation are not fully known. Here, Illumina sRNA sequencing was applied to sequence six sRNA libraries generated from the waterlogging-tolerant cucumber Zaoer-N after 48 h of waterlogging and the control. A total of 358 cucumber miRNAs, 312 known and 46 novel, were obtained. Among them, 23 were differentially expressed, with 10 and 13 being up- and downregulated, respectively. A qPCR expression study confirmed that the identified differentially expressed miRNAs were credible. A total of 657 putative miRNA target genes were predicted for the 23 miRNAs using an in silico approach. A gene ontology enrichment analysis revealed that target genes functioning in cell redox homeostasis, cytoskeleton, photosynthesis and cell growth were over-represented. In total, 58 of the 657 target genes showed inverse expression patterns compared with their respective miRNAs through a combined analysis of sRNA- and RNA-sequencing-based transcriptome datasets using the same experimental design. The target gene annotation included a peroxidase, a GDSL esterases/lipase and two heavy metal-associated isoprenylated plant proteins. Our results provide an important framework for understanding the unique miRNA patterns seen in responses to waterlogging and the miRNA-mediated formation of de novo AR primordia in cucumber.





Similar content being viewed by others
References
Shabala S (2011) Physiological and cellular aspects of phytotoxicity tolerance in plants: the role of membrane transporters and implications for crop breeding for waterlogging tolerance. New Phytol 190(2):289–298
Xu X, Ji J, Xu Q, Qi X, Weng Y, Chen X (2018) The major-effect quantitative trait locus CsARN 6.1 encodes an AAA ATP ase domain-containing protein that is associated with waterlogging stress tolerance by promoting adventitious root formation. Plant J 93(5):917–930
Mancuso S, Shabala S (2010) Waterlogging signalling and tolerance in plants. Springer, Berlin
Pearson A, Cogan N, Baillie R, Hand M, Bandaranayake C, Erb S, Wang J, Kearney G, Gendall A, Smith K, Forster J (2011) Identification of QTLs for morphological traits influencing waterlogging tolerance in perennial ryegrass (Lolium perenne L.) Theor Appl Genet 122:609–622
Armstrong W (1971) Radial oxygen losses from intact rice roots as affected by distance from the apex, respiration and waterlogging. Physiol Plant 25:192–197
Medri M, Ferreira A, Kolb R, Bianchini E, Pimenta J, Davanso-Fabro V, Medri C (2007) Morpho-anatomical alterations in plants of Lithraea molleoides (Vell.) Engl. submitted to flooding. Acta Sci Biol Sci 29:15–22
Abiko T, Kotula L, Shiono K, Colmer T, Nakazono M (2012) Enhanced formation of aerenchyma and induction of a barrier to radial oxygen loss in adventitious roots of Zea nicaraguensis contribute to its waterlogging tolerance as compared with maize (Zea mays ssp. mays). Plant, Cell Environ 35(9):1618–1630
Chen T, Yuan F, Song J, Wang B (2016) Nitric oxide participates in waterlogging tolerance through enhanced adventitious root formation in the euhalophyte Suaeda salsa. Funct Plant Biol 43(3):244–253
Rich S, Ludwig M, Colmer T (2008) Photosynthesis in aquatic adventitious roots of the halophytic stem-succulent Tecticornia pergranulata (formerly Halosarcia pergranulata). Plant Cell Environ 31(7):1007–1016
Rich S, Ludwig M, Pedersen O, Colmer T (2011) Aquatic adventitious roots of the wetland plant Meionectes brownii can photosynthesize: implications for root function during flooding. New Phytol 190(2):311–319
Xu X, Chen M, Ji J, Xu Q, Qi X, Chen X (2017) Comparative RNA-seq based transcriptome profiling of waterlogging response in cucumber hypocotyls reveals novel insights into the de novo adventitious root primordia initiation. BMC Plant Biol 17(1):129
Lakehal A, Bellini C (2019) Control of adventitious root formation: insights into synergistic and antagonistic hormonal interactions. Physiol Plant 165(1):90–100
Visser E, Bögemann G, Blom C, Voesenek L (1996) Ethylene accumulation in waterlogged Rumex plants promotes formation of adventitious roots. J Exp Bot 47(3):403–410
Lorbiecke R, Sauter M (1999) Adventitious root growth and cell-cycle induction in deepwater rice. Plant Physiol 119(1):21–30
Steffens B, Kovalev A, Gorb S, Sauter M (2012) Emerging roots alter epidermal cell fate through mechanical and reactive oxygen species signaling. Plant Cell 24(8):3296–3306
Dawood T, Rieu I, Wolters-Arts M, Derksen E, Mariani C, Visser E (2014) Rapid flooding-induced adventitious root development from preformed primordia in Solanum dulcamara. AoB Plants 6:plt058
Vidoz M, Loreti E, Mensuali A, Alpi A, Perata P (2010) Hormonal interplay during adventitious root formation in flooded tomato plants. Plant J 63(4):551–562
Xu X, Wang H, Qi X, Xu Q, Chen X (2014) Waterlogging-induced increase in fermentation and related gene expression in the root of cucumber (Cucumis sativus L.) Sci Hortic 179:388–395
Xu X, Ji J, Ma X, Xu Q, Qi X, Chen X (2016) Comparative proteomic analysis provides insight into the key proteins involved in cucumber (Cucumis sativus L.) adventitious root emergence under waterlogging stress. Front Plant Sci 7:1515
Zhang B, Wang Q (2015) MicroRNA-based biotechnology for plant improvement. J Cell Physiol 230(1):1–15
Garg V, Khan A, Kudapa H, Kale S, Chitikineni A, Sun Q, Sharma M, Li C, Zhang B, Liu X, Kisho K (2019) Integrated transcriptome, small RNA and degradome sequencing approaches provide insights into Ascochyta blight resistance in chickpea. Plant Biotechnol J 17:914–931
Liu Z, Kumari S, Zhang L, Zheng Y, Ware D (2012) Characterization of miRNAs in response to short-term waterlogging in three inbred lines of Zea mays. PLoS ONE 7(6):e39786
Zhai L, Liu Z, Zou X, Jiang Y, Qiu F, Zheng Y, Zhang Z (2013) Genome-wide identification and analysis of microRNA responding to long-term waterlogging in crown roots of maize seedlings. Physiol Plant 147(2):181–193
Jin Q, Xu Y, Mattson N, Li X, Wang B, Zhang X, Jiang H, Liu X, Yang W, Yao D (2017) Identification of submergence-responsive microRNAs and their targets reveals complex miRNA-mediated regulatory networks in lotus (Nelumbo nucifera Gaertn). Front Plant Sci 8:6
Wen M, Shen Y, Shi S, Tang T (2012) miREvo: an integrative microRNA evolutionary analysis platform for next-generation sequencing experiments. BMC Bioinform 13:140
Friedlander M, Mackowiak S, Li N, Chen W, Rajewsky N (2011) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucl Acids Res 40:37–52
Zhou L, Chen J, Li Z, Li X, Hu X, Hang Y, Zhao X, Liang C, Wang Y, Sun L, Shi M, Xu X, Shen F, Chen M, Han Z, Peng Z, Zhai Q, Chen J, Zhang Z, Yang R, Ye J, Guan Z, Yang H, Gui Y, Wang J, Cai Z, Zhang X (2010) Integrated profiling of microRNAs and mRNAs: microRNAs located on Xq27.3 associate with clear cell renal cell carcinoma. PLoS ONE 5:e15224
Love M, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550
Xie F, Jones D, Wang Q, Sun R, Zhang B (2015) Small RNA sequencing identifies miRNA roles in ovule and fibre development. Plant Biotechnol J 13(3):355–369
Yin D, Li S, Shu Q, Gu Z, Wu Q, Feng C, Xu W, Wang L (2018) Identification of microRNAs and long non-coding RNAs involved in fatty acid biosynthesis in tree peony seeds. Gene 666:72–82
Moné Y, Nhim S, Gimenez S, Legeai F, Seninet I, Parrinello H, Nègre N, d’Alençon E (2018) Characterization and expression profiling of microRNAs in response to plant feeding in two host-plant strains of the lepidopteran pest Spodoptera frugiperda. BMC Genom 19(1):804
Martínez G, Forment J, Llave C, Pallás V, Gómez G (2011) High-throughput sequencing, characterization and detection of new and conserved cucumber miRNAs. PLoS ONE 6(5):e19523
Zeng X, Xu Y, Jiang J, Zhang F, Ma L, Wu D, Wang Y, Sun W (2018) Identification of cold stress responsive microRNAs in two winter turnip rape (Brassica rapa L.) by high throughput sequencing. BMC Plant Biol 18(1):52
Carlsbecker A, Lee J, Roberts C, Dettmer J, Lehesranta S, Zhou J, Lindgren O, Moreno-Risueno M, Vatén A, Thitamadee S, Campilho A, Sebastian J, Bowman J, Helariutta Y, Benfey P (2010) Cell signalling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465(7296):316
Singh A, Roy S, Singh S, Das S, Gautam V, Yadav S, Kumar A, Singh A, Samantha S, Sarkar A (2017) Phytohormonal crosstalk modulates the expression of miR166/165 s, target Class III HD-ZIPs, and KANADI genes during root growth in Arabidopsis thaliana. Sci Rep 7(1):3408
Luo N, Yu X, Nie G, Liu J, Jiang Y (2016) Specific peroxidases differentiate Brachypodium distachyon accessions and are associated with drought tolerance traits. Ann Bot 118(2):259–270
Passard F, Tognolli M, De Meyer M, Penel C, Dunand C (2006) Two cell wall associated peroxidases from Arabidopsis influence root elongation. Planta 223(5):965–974
Chepyshko H, Lai C, Huang L, Liu J, Shaw J (2012) Multifunctionality and diversity of GDSL esterase/lipase gene family in rice (Oryza sativa L. japonica) genome: new insights from bioinformatics analysis. BMC Genom 13(1):309
Kim R, Kim H, Shim D, Suh M (2016) Molecular and biochemical characterizations of the monoacylglycerol lipase gene family of Arabidopsis thaliana. Plant J 85(6):758–771
Dykema P, Sipes P, Marie A, Biermann B, Crowell D, Randall S (1999) A new class of proteins capable of binding transition metals. Plant Mol Biol 41(1):139–150
de Abreu-Neto J, Turchetto-Zolet A, de Oliveira L, Zanettini M, Margis-Pinheiro M (2013) Heavy metal-associated isoprenylated plant protein (HIPP): characterization of a family of proteins exclusive to plants. FEBS J 280(7):1604–1616
Gomathi R, Rao P, Chandran K, Selvi A (2015) Adaptive responses of sugarcane to waterlogging stress: an over view. Sugar Tech 17(4):325–338
Acknowledgements
This research was supported by the National Natural Science Foundation of China (Grant Nos. 31171978, 31672176 and 31801883) and the Natural Science Foundation of the Jiangsu Higher Education Institutions of China (18KJB210014). We thank Lesley Benyon, PhD, from Liwen Bianji, Edanz Group China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
XC conceived and designed the experiment. XX and KW performed the experiments. XX and JP interpreted the data and wrote the manuscript. All authors reviewed and approved this submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material
11033_2019_5084_MOESM1_ESM.xlsx
Online Resource 1 (XLSX 11 kb). Distribution of cucumber small RNAs in the six sequencing libraries. NAT, natural antisense transcripts. C1, C2 and C3 represent the three replications of control hypocotyl samples; W1, W2 and W3 represent the three replications of waterlogged hypocotyl samples
11033_2019_5084_MOESM2_ESM.xlsx
Online Resource 2 (XLSX 78 kb). Known and novel miRNAs identified in cucumber hypocotyls from the six libraries. C1, C2 and C3 represent the three replications of control hypocotyl samples; W1, W2 and W3 represent the three replications of waterlogged hypocotyl samples. TPM represents transcripts per kilobase million
11033_2019_5084_MOESM3_ESM.xlsx
Online Resource 3 (XLSX 74 kb). List of cucumber miRNAs and their target genes predicted by the online psRNATarget server
11033_2019_5084_MOESM5_ESM.xlsx
Online Resource 5 (XLSX 17 kb). List of the fragments per kilobase million (FPKM) values of the 58 negatively regulated miRNAs target genes that were identified by RNA-seq. C1, C2 and C3 represent the three replications of control hypocotyl samples; W1, W2 and W3 represent the three replications of waterlogged hypocotyl samples
Rights and permissions
About this article
Cite this article
Xu, X., Wang, K., Pan, J. et al. Small RNA sequencing identifies cucumber miRNA roles in waterlogging-triggered adventitious root primordia formation. Mol Biol Rep 46, 6381–6389 (2019). https://doi.org/10.1007/s11033-019-05084-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-05084-z