Abstract
Introduction
The ability to identify early epigenetic signatures underlying the inheritance of cardiovascular risk, including trans- and intergenerational effects, may help to stratify people before cardiac symptoms occur.
Methods
Prospective and retrospective cohorts and case–control studies focusing on DNA methylation and maternal/paternal effects were searched in Pubmed from 1997 to 2023 by using the following keywords: DNA methylation, genomic imprinting, and network analysis in combination with transgenerational/intergenerational effects.
Results
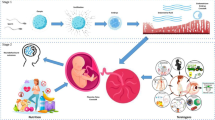
Maternal and paternal exposures to traditional cardiovascular risk factors during critical temporal windows, including the preconceptional period or early pregnancy, may perturb the plasticity of the epigenome (mainly DNA methylation) of the developing fetus especially at imprinted loci, such as the insulin-like growth factor type 2 (IGF2) gene. Thus, the epigenome is akin to a “molecular archive” able to memorize parental environmental insults and predispose an individual to cardiovascular diseases onset in later life. Direct evidence for human transgenerational epigenetic inheritance (at least three generations) of cardiovascular risk is lacking but it is supported by epidemiological studies. Several blood-based association studies showed potential intergenerational epigenetic effects (single-generation studies) which may mediate the transmittance of cardiovascular risk from parents to offspring.
Discussion
In this narrative review, we discuss some relevant examples of trans- and intergenerational epigenetic associations with cardiovascular risk. In our perspective, we propose three network-oriented approaches which may help to clarify the unsolved issues regarding transgenerational epigenetic inheritance of cardiovascular risk and provide potential early biomarkers for primary prevention.
Significance
Many maternal risk factors, including dyslipidemia, hypertension, smoking, diabetes, and obesity have been associated with CVDs risk in the offspring. Paternal factors are emerging as risk factors contributing to transgenerational effects. From a molecular point of view, DNA methylation may be responsible, at least in part, for early modifications during fetal development which set CVD risk trajectories during post-natal life.
AbstractSection What this study adds?Big data availability supports the importance of Network Medicine approaches for integrating environmental exposures, clinical information, and advanced next-generation sequencing platform data over time, representing one of the most potent research paradigms for identifying novel molecular routes underlying the potential transmittance of CVD risk from parents to their offspring. The possible inheritance of DNA methylation signatures from parents to offspring and the ability to detect them using cardiac tissue, or blood as surrogate, may provide useful biomarkers for primary prevention of CVDs.






Source of Figures: The Figures are original. We used the Servier Medical Art by Servier which is licensed under a Creative Commons Attribution 3.0 Unported License (https://smart.servier.com/)
Similar content being viewed by others
Data Availability
Not applicable.
Code Availability
Not applicable.
References
Agha, G., Mendelson, M. M., Ward-Caviness, C. K., Joehanes, R., Huan, T., Gondalia, R., et al. (2019). Blood leukocyte DNA methylation predicts risk of future myocardial infarction and coronary heart disease. Circulation, 140, 645–657. https://doi.org/10.1161/CIRCULATIONAHA.118.039357
Antoun, E., Kitaba, N. T., Titcombe, P., Dalrymple, K. V., Garratt, E. S., Barton, S. J., et al. (2020). Maternal dysglycaemia, changes in the infant’s epigenome modified with a diet and physical activity intervention in pregnancy: Secondary analysis of a randomised control trial. PLoS Medicine, 17, e1003229. https://doi.org/10.1371/journal.pmed.1003229
Barker, D. J., Osmond, C., Forsen, T. J., Kajantie, E., & Eriksson, J. G. (2005). Trajectories of growth among children who have coronary events as adults. New England Journal of Medicine, 353, 1802–1809. https://doi.org/10.1056/NEJMoa044160
Bauer, T., Trump, S., Ishaque, N., Thürmann, L., Gu, L., Bauer, M., et al. (2016). Environment-induced epigenetic reprogramming in genomic regulatory elements in smoking mothers and their children. Molecular Systems Biology, 12, 861. https://doi.org/10.15252/msb.20156520
Benincasa, G., Coscioni, E., & Napoli, C. (2022). Cardiovascular risk factors and molecular routes underlying endothelial dysfunction: Novel opportunities for primary prevention. Biochemical Pharmacology, 202, 115108. https://doi.org/10.1016/j.bcp.2022.115108
Benincasa, G., Mansueto, G., & Napoli, C. (2019). Fluid-based assays and precision medicine of cardiovascular diseases: The ‘hope’ for Pandora’s box? Journal of Clinical Pathology, 72, 785–799. https://doi.org/10.1136/jclinpath-2019-206178
Benincasa, G., Marfella, R., Della Mura, N., Schiano, C., & Napoli, C. (2020). Strengths and opportunities of network medicine in cardiovascular diseases. Circulation Journal, 84, 144–152. https://doi.org/10.1253/circj.CJ-19-0879
Benincasa, G., Maron, B. A., Affinito, O., D’Alto, M., Franzese, M., Argiento, P., et al. (2023). Association between circulating CD4+ T cell methylation signatures of network-oriented SOCS3 gene and hemodynamics in patients suffering pulmonary arterial hypertension. Journal of Cardiovascular Translational Research, 16, 17–30. https://doi.org/10.1007/s12265-022-10294-1
Cardenas, A., Gagné-Ouellet, V., Allard, C., Brisson, D., Perron, P., Bouchard, L., et al. (2018). Placental DNA methylation adaptation to maternal glycemic response in pregnancy. Diabetes, 67, 1673–1683. https://doi.org/10.2337/db18-0123
Cedar, H., & Bergman, Y. (2009). Linking DNA methylation and histone modification: Patterns and paradigms. Nature Reviews Genetics, 10, 295–304. https://doi.org/10.1038/nrg2540
Chatterton, Z., Hartley, B. J., Seok, M. H., Mendelev, N., Chen, S., Milekic, M., et al. (2017). In utero exposure to maternal smoking is associated with DNA methylation alterations and reduced neuronal content in the developing fetal brain. Epigenetics & Chromatin, 10, 4. https://doi.org/10.1186/s13072-017-0111-y
Chen, L., Ge, B., Casale, F. P., Vasquez, L., Kwan, T., Garrido-Martín, D., et al. (2016). Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell, 167, 1398-1414.e24. https://doi.org/10.1016/j.cell.2016.10.026
Chhabra, D., Sharma, S., Kho, A. T., Gaedigk, R., Vyhlidal, C. A., Leeder, J. S., et al. (2014). Fetal lung and placental methylation is associated with in utero nicotine exposure. Epigenetics, 9, 1473–1484. https://doi.org/10.4161/15592294.2014.971593
Cuellar Partida, G., Laurin, C., Ring, S. M., Gaunt, T. R., McRae, A. F., Visscher, P. M., et al. (2018). Genome-wide survey of parent-of-origin effects on DNA methylation identifies candidate imprinted loci in humans. Human Molecular Genetics, 27, 2927–2939. https://doi.org/10.1093/hmg/ddy206
de Nigris, F., Cacciatore, F., Mancini, F. P., Vitale, D. F., Mansueto, G., D’Armiento, F. P., et al. (2018). Epigenetic hallmarks of fetal early atherosclerotic lesions in humans. JAMA Cardiol, 3, 1184–1191. https://doi.org/10.1001/jamacardio.2018.3546
Everson, T. M., Vives-Usano, M., Seyve, E., Cardenas, A., Lacasaña, M., Craig, J. M., et al. (2021). Placental DNA methylation signatures of maternal smoking during pregnancy and potential impacts on fetal growth. Nature Communications, 12, 5095. https://doi.org/10.1038/s41467-021-24558-y
Feil, R., & Fraga, M. F. (2012). Epigenetics and the environment: Emerging patterns and implications. Nature Reviews Genetics, 13, 97–109. https://doi.org/10.1038/nrg3142
Feinberg, A. P. (2018). The key role of epigenetics in human disease prevention and mitigation. New England Journal of Medicine, 378, 1323–1334. https://doi.org/10.1056/NEJMra1402513
Gaunt, T. R., Shihab, H. A., Hemani, G., Min, J. L., Woodward, G., Lyttleton, O., et al. (2016). Systematic identification of genetic influences on methylation across the human life course. Genome Biology, 17, 61. https://doi.org/10.1186/s13059-016-0926-z
Geurtsen, M. L., Jaddoe, V. W. V., Gaillard, R., & Felix, J. F. (2020). Associations of maternal early-pregnancy blood glucose and insulin concentrations with DNA methylation in newborns. Clinical Epigenetics, 12, 134. https://doi.org/10.1186/s13148-020-00924-3
Gonzalez-Rodriguez, P., Cantu, J., O’Neil, D., Seferovic, M. D., Goodspeed, D. M., Suter, M. A., et al. (2016). Alterations in expression of imprinted genes from the H19/IGF2 loci in a multigenerational model of intrauterine growth restriction (IUGR). American Journal of Obstetrics and Gynecology, 214, 625.e1-625.e11. https://doi.org/10.1016/j.ajog.2016.01.194
Guay, S. P., Houde, A. A., Breton, E., Baillargeon, J. P., Perron, P., Gaudet, D., et al. (2020). DNA methylation at LRP1 gene locus mediates the association between maternal total cholesterol changes in pregnancy and cord blood leptin levels. Journal of Developmental Origins of Health and Disease, 11, 369–378. https://doi.org/10.1017/S204017441900076X
Gui, W., Liang, J., Lin, X., Shi, N., Zhu, Y., Tan, B., & Li, H. (2021). Association of genetic variants in IGF2-related genes with risk of metabolic syndrome in the Chinese han population. Front Endocrinol (lausanne), 12, 654747. https://doi.org/10.3389/fendo.2021.654747
Hannon, E., Weedon, M., Bray, N., O’Donovan, M., & Mill, J. (2017). Pleiotropic effects of trait-associated genetic variation on dna methylation: Utility for refining GWAS loci. American Journal of Human Genetics, 100, 954–959. https://doi.org/10.1016/j.ajhg.2017.04.013
Heard, E., & Martienssen, R. A. (2014). Transgenerational epigenetic inheritance: Myths and mechanisms. Cell, 157, 95–109. https://doi.org/10.1016/j.cell.2014.02.045
Hedman, Å. K., Mendelson, M. M., Marioni, R. E., Gustafsson, S., Joehanes, R., Irvin, M. R., et al. (2017). Epigenetic patterns in blood associated with lipid traits predict incident coronary heart disease events and are enriched for results from genome-wide association studies. Circulation. Cardiovascular Genetics, 10, e001487. https://doi.org/10.1161/CIRCGENETICS.116.001487
Heijmans, B. T., Tobi, E. W., Stein, A. D., Putter, H., Blauw, G. J., Susser, E. S., et al. (2008). Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci U S A, 105, 17046–17049. https://doi.org/10.1073/pnas.0806560105
Horsthemke, B. (2018). A critical view on transgenerational epigenetic inheritance in humans. Nature Communications, 9, 2973. https://doi.org/10.1038/s41467-018-05445-5
Howe, C. G., Cox, B., Fore, R., Jungius, J., Kvist, T., Lent, S., et al. (2020). Maternal gestational diabetes mellitus and newborn DNA methylation: Findings from the Pregnancy and Childhood Epigenetics Consortium. Diabetes Care, 43, 98–105. https://doi.org/10.2337/dc19-0524
Huan, T., Joehanes, R., Song, C., Peng, F., Guo, Y., Mendelson, M., et al. (2019). Genome-wide identification of DNA methylation QTLs in whole blood highlights pathways for cardiovascular disease. Nature Communications, 10, 4267. https://doi.org/10.1038/s41467-019-12228-z
Huang, R. C., Galati, J. C., Burrows, S., Beilin, L. J., Li, X., Pennell, C. E., et al. (2012). DNA methylation of the IGF2/H19 imprinting control region and adiposity distribution in young adults. Clinical Epigenetics, 4, 21. https://doi.org/10.1186/1868-7083-4-21
Jahan, N., Naveed, S., Zeshan, M., & Tahir, M. A. (2016). How to conduct a systematic review: A narrative literature review. Cureus, 8, e864. https://doi.org/10.7759/cureus.864
Jones, P. A. (2012). Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nature Reviews Genetics, 13, 484–492. https://doi.org/10.1038/nrg3230. PMID: 22641018.
Joubert, B. R., Felix, J. F., Yousefi, P., Bakulski, K. M., Just, A. C., Breton, C., et al. (2016). DNA methylation in newborns and maternal smoking in pregnancy: Genome-wide Consortium Meta-analysis. American Journal of Human Genetics, 98, 680–696. https://doi.org/10.1016/j.ajhg.2016.02.019
Kachroo, P., Morrow, J. D., Kho, A. T., Vyhlidal, C. A., Silverman, E. K., Weiss, S. T., et al. (2020). Co-methylation analysis in lung tissue identifies pathways for fetal origins of COPD. European Respiratory Journal, 56, 1902347. https://doi.org/10.1183/13993003.02347-2019
Kato, N., Loh, M., Takeuchi, F., Verweij, N., Wang, X., Zhang, W., et al. (2015). Trans-ancestry genome-wide association study identifies 12 genetic loci influencing blood pressure and implicates a role for DNA methylation. Nature Genetics, 47, 1282–1293. https://doi.org/10.1038/ng.3405
Kazmi, N., Sharp, G. C., Reese, S. E., Vehmeijer, F. O., Lahti, J., Page, C. M., et al. (2019). Hypertensive disorders of pregnancy and DNA methylation in newborns. Hypertension, 74, 375–383. https://doi.org/10.1161/HYPERTENSIONAHA.119.12634
Kim, S. S., Dai, C., Hormozdiari, F., van de Geijn, B., Gazal, S., Park, Y., O’Connor, L., et al. (2019). Genes with high network connectivity are enriched for disease heritability. American Journal of Human Genetics, 104, 896–913. https://doi.org/10.1016/j.ajhg.2019.03.020
Kuijjer, M. L., Hsieh, P. H., Quackenbush, J., & Glass, K. (2019). LionessR: Single sample network inference in R. BMC Cancer, 19, 1003. https://doi.org/10.1186/s12885-019-6235-7
Napoli, C., Benincasa, G., Donatelli, F., & Ambrosio, G. (2020a). Precision medicine in distinct heart failure phenotypes: Focus on clinical epigenetics. American Heart Journal, 224, 113–128. https://doi.org/10.1016/j.ahj.2020.03.007
Napoli, C., Benincasa, G., Schiano, C., & Salvatore, M. (2020b). Differential epigenetic factors in the prediction of cardiovascular risk in diabetic patients. Eur Heart J Cardiovasc Pharmacother, 6, 239–247. https://doi.org/10.1093/ehjcvp/pvz062
Napoli, C., Crudele, V., Soricelli, A., Al-Omran, M., Vitale, N., Infante, T., Mancini, F. P., et al. (2012). Primary prevention of atherosclerosis: A clinical challenge for the reversal of epigenetic mechanisms? Circulation, 125, 2363–2373. https://doi.org/10.1161/CIRCULATIONAHA.111.085787
Napoli, C., D’Armiento, F. P., Mancini, F. P., Postiglione, A., Witztum, J. L., Palumbo, G., et al. (1997). Fatty streak formation occurs in human fetal aortas and is greatly enhanced by maternal hypercholesterolemia. Intimal accumulation of low density lipoprotein and its oxidation precede monocyte recruitment into early atherosclerotic lesions. The Journal of Clinical Investigation, 100, 2680–2690. https://doi.org/10.1172/JCI119813
Napoli, C., Glass, C. K., Witztum, J. L., Deutsch, R., D’Armiento, F. P., & Palinski, W. (1999). Influence of maternal hypercholesterolaemia during pregnancy on progression of early atherosclerotic lesions in childhood: Fate of Early Lesions in Children (FELIC) study. Lancet, 354, 1234–1241. https://doi.org/10.1016/S0140-6736(99)02131-5
Napoli, C., Lerman, L. O., de Nigris, F., Gossl, M., Balestrieri, M. L., & Lerman, A. (2006). Rethinking primary prevention of atherosclerosis-related diseases. Circulation, 114, 2517–2527. https://doi.org/10.1161/CIRCULATIONAHA.105.570358
Nordin, M., Bergman, D., Halje, M., Engström, W., & Ward, A. (2014). Epigenetic regulation of the Igf2/H19 gene cluster. Cell Proliferation, 47, 189–199. https://doi.org/10.1111/cpr.12106
Olsson, A. H., Volkov, P., Bacos, K., Dayeh, T., Hall, E., Nilsson, E. A., et al. (2014). Genome-wide associations between genetic and epigenetic variation influence mRNA expression and insulin secretion in human pancreatic islets. PLoS Genetics, 10, e1004735. https://doi.org/10.1371/journal.pgen.1004735
Ouidir, M., Zeng, X., Workalemahu, T., Shrestha, D., Grantz, K. L., Mendola, P., et al. (2020). Early pregnancy dyslipidemia is associated with placental DNA methylation at loci relevant for cardiometabolic diseases. Epigenomics, 2020, 921–934. https://doi.org/10.2217/epi-2019-0293
Perkins, E., Murphy, S. K., Murtha, A. P., Schildkraut, J., Jirtle, R. L., Demark-Wahnefried, W., et al. (2012). Insulin-like growth factor 2/H19 methylation at birth and risk of overweight and obesity in children. Journal of Pediatrics, 161, 31–39. https://doi.org/10.1016/j.jpeds.2012.01.015
Putra, S. E. D., Reichetzeder, C., von Websky, K., Neuber, C., Halle, H., Kleuser, B., et al. (2022). Association between placental global DNA methylation and blood pressure during human pregnancy. Journal of Hypertension, 40, 1002–1009. https://doi.org/10.1097/HJH.0000000000003103
Rauschert, S., Melton, P. E., Burdge, G., Craig, J. M., Godfrey, K. M., Holbrook, J. D., et al. (2019). Maternal smoking during pregnancy induces persistent epigenetic changes into adolescence, independent of postnatal smoke exposure and is associated with cardiometabolic risk. Frontiers in Genetics, 10, 770. https://doi.org/10.3389/fgene.2019.00770
Reed, Z. E., Suderman, M. J., Relton, C. L., Davis, O. S. P., & Hemani, G. (2020). The association of DNA methylation with body mass index: Distinguishing between predictors and biomarkers. Clinical Epigenetics, 12, 50. https://doi.org/10.1186/s13148-020-00841-5
Richmond, R. C., Simpkin, A. J., Woodward, G., Gaunt, T. R., Lyttleton, O., McArdle, W. L., et al. (2015). Prenatal exposure to maternal smoking and offspring DNA methylation across the life course: Findings from the Avon Longitudinal Study of Parents and Children (ALSPAC). Human Molecular Genetics, 24, 2201–2217. https://doi.org/10.1093/hmg/ddu739
Röhl, A., Baek, S. H., Kachroo, P., Morrow, J. D., Tantisira, K., Silverman, E. K., et al. (2022). Protein interaction networks provide insight into fetal origins of chronic obstructive pulmonary disease. Respiratory Research, 23, 69. https://doi.org/10.1186/s12931-022-01963-5
Sarno, F., Benincasa, G., List, M., Barabasi, A. L., Baumbach, J., Ciardiello, F., et al. (2021). Clinical epigenetics settings for cancer and cardiovascular diseases: Real-life applications of network medicine at the bedside. Clinical Epigenetics, 13, 66. https://doi.org/10.1186/s13148-021-01047-z
Schiano, C., D’Armiento, M., Franzese, M., Castaldo, R., Saccone, G., de Nigris, F., et al. (2022). DNA methylation profile of the SREBF2 gene in human fetal aortas. Journal of Vascular Research, 59, 61–68. https://doi.org/10.1159/000518513
Sharp, G. C., Alfano, R., Ghantous, A., Urquiza, J., Rifas-Shiman, S. L., Page, C. M., et al. (2021). Paternal body mass index and offspring DNA methylation: Findings from the PACE consortium. International Journal of Epidemiology, 50, 1297–1315. https://doi.org/10.1093/ije/dyaa267
Silverman, E. K., Schmidt, H. H. H. W., Anastasiadou, E., Altucci, L., Angelini, M., Badimon, L., et al. (2020). Molecular networks in network medicine: Development and applications. Wiley Interdiscip Rev Syst Biol Med, 12, e1489. https://doi.org/10.1002/wsbm.1489
Soubry, A., Murphy, S. K., Wang, F., Huang, Z., Vidal, A. C., Fuemmeler, B. F., et al. (2015). Newborns of obese parents have altered DNA methylation patterns at imprinted genes. International Journal of Obesity, 39, 650–657. https://doi.org/10.1038/ijo.2013.193
Soubry, A., Schildkraut, J. M., Murtha, A., Wang, F., Huang, Z., Bernal, A., et al. (2013). Paternal obesity is associated with IGF2 hypomethylation in newborns: Results from a Newborn Epigenetics Study (NEST) cohort. BMC Medicine, 11, 29. https://doi.org/10.1186/1741-7015-11-29
Teschendorff, A. E., Zhu, T., Breeze, C. E., & Beck, S. (2020). EPISCORE: Cell type deconvolution of bulk tissue DNA methylomes from single-cell RNA-Seq data. Genome Biology, 21, 221. https://doi.org/10.1186/s13059-020-02126-9
Tobi, E. W., Goeman, J. J., Monajemi, R., Gu, H., Putter, H., Zhang, Y., et al. (2015). DNA methylation signatures link prenatal famine exposure to growth and metabolism. Nature Communications, 6, 7740. https://doi.org/10.1038/ncomms8740
Tucci, V., Isles, A. R., Kelsey, G., & Ferguson-Smith, A. C. (2019). Genomic imprinting and physiological processes in mammals. Cell, 176, 952–965. https://doi.org/10.1016/j.cell.2019.01.043
van Otterdijk, S. D., & Michels, K. B. (2016). Transgenerational epigenetic inheritance in mammals: How good is the evidence? The FASEB Journal, 30, 2457–2465. https://doi.org/10.1096/fj.201500083
Vives-Usano, M., Hernandez-Ferrer, C., Maitre, L., Ruiz-Arenas, C., Andrusaityte, S., Borràs, E., et al. (2020). In utero and childhood exposure to tobacco smoke and multi-layer molecular signatures in children. BMC Medicine, 18, 243. https://doi.org/10.1186/s12916-020-01686-8
Volkov, P., Olsson, A. H., Gillberg, L., Jørgensen, S. W., Brøns, C., Eriksson, K. F., et al. (2016). A genome-wide mQTL analysis in human adipose tissue identifies genetic variants associated with DNA methylation, gene expression and metabolic traits. PLoS ONE, 11, e0157776. https://doi.org/10.1371/journal.pone.0157776
Wu, L., Li, N., & Liu, Y. (2023). Association between maternal factors and risk of congenital heart disease in offspring: A systematic review and meta-analysis. Maternal and Child Health Journal, 27, 29–48. https://doi.org/10.1007/s10995-022-03538-8
Xu, R., Hong, X., Zhang, B., Huang, W., Hou, W., Wang, G., et al. (2021). DNA methylation mediates the effect of maternal smoking on offspring birthweight: A birth cohort study of multi-ethnic US mother-newborn pairs. Clinical Epigenetics, 13, 47. https://doi.org/10.1186/s13148-021-01032-6
Zhu, T., Liu, J., Beck, S., Pan, S., Capper, D., Lechner, M., et al. (2022). A pan-tissue DNA methylation atlas enables in silico decomposition of human tissue methylomes at cell-type resolution. Nature Methods, 19, 296–306. https://doi.org/10.1038/s41592-022-01412-7
Funding
None.
Author information
Authors and Affiliations
Contributions
GB and DDM wrote the manuscript, CN critically revised the content.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Benincasa, G., Napoli, C. & DeMeo, D.L. Transgenerational Epigenetic Inheritance of Cardiovascular Diseases: A Network Medicine Perspective. Matern Child Health J 28, 617–630 (2024). https://doi.org/10.1007/s10995-023-03886-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10995-023-03886-z