Abstract
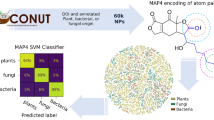
Natural products (NP) or secondary metabolites, as a class of small chemical molecules that are naturally synthesized by chromosomally clustered biosynthesis genes (also called biosynthetic gene clusters, BGCs) encoded enzymes or enzyme complexes, mediates the bioecological interactions between host and microbiota and provides a natural reservoir for screening drug-like therapeutic pharmaceuticals. In this work, we propose a multi-label learning framework to functionally annotate natural products or secondary metabolites solely from their catalytical biosynthetic gene clusters without experimentally conducting NP structural resolutions. All chemical classes and bioactivities constitute the label space, and the sequence domains of biosynthetic gene clusters that catalyse the biosynthesis of natural products constitute the feature space. In this multi-label learning framework, a joint representation of features (BGCs domains) and labels (natural products annotations) is efficiently learnt in an integral and low-dimensional space to accurately define the inter-class boundaries and scale to the learning problem of many imbalanced labels. Computational results on experimental data show that the proposed framework achieves satisfactory multi-label learning performance, and the learnt patterns of BGCs domains are transferrable across bacteria, or even across kingdom, for instance, from bacteria to Arabidopsis thaliana. Lastly, take Arabidopsis thaliana and its rhizosphere microbiome for example, we propose a pipeline combining existing BGCs identification tools and this proposed framework to find and functionally annotate novel natural products for downstream bioecological studies in terms of plant-microbiota-soil interactions and plant environmental adaption.




Similar content being viewed by others
Data Availability
Not applicable.
References
Aghdam SA, Brown AMV (2021) Deep learning approaches for natural product discovery from plant endophytic microbiomes. Environ Microbiome 16:6
Alam K, Hao J, Zhang Y, Li A (2021) Synthetic biology-inspired strategies and tools for engineering of microbial natural product biosynthetic pathways. Biotechnol Adv 49:07759
Atanasov AG, Zotchev SB, Dirsch VM (2021) Natural products in drug discovery: advances and opportunities. Nat Rev Drug Discov 28:1–17
Begani J, Lakhani J, Harwani D (2018) Current strategies to induce secondary metabolites from microbial biosynthetic cryptic gene clusters. Annals Microbiol 68:419–432
Blin K, Medema MH, Kazempour D, Fischbach MA, Breitling R, Takano E, Weber T (2013) antiSMASH 2.0–a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res 41(Web Server issue):W204-12
Blin K, Kim HU, Medema MH, Weber T (2019) Recent development of antiSMASH and other computational approaches to mine secondary metabolite biosynthetic gene clusters. Brief Bioinform 20(4):1103–1113
Blin K, Shaw S, Tong Y, Weber T (2020) Designing sgRNAs for CRISPR-BEST base editing applications with CRISPy-web 2.0. Synth Syst Biotechnol 5:99–102
Blin K, Shaw S, Kloosterman AM, Charlop-Powers Z, Wezel GPV et al (2021a) antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res 49:W29–W35
Blin K, Shaw S, Kautsar SA, Medema MH, Weber T (2021b) The antiSMASH database version 3: increased taxonomic coverage and new query features for modular enzymes. Nucleic Acids Res 49:D639–D643
Blum M, Chang H, Chuguransky S, Grego T, Kandasaamy S et al (2021) The InterPro protein families and domains database: 20 years on. Nucleic Acids Res 49(D1):D344–D354
Breiman L (2001) Random forests. Mach Learn 45:5–32
Capecchi A, Reymond JL (2020) Assigning the Origin of Microbial Natural Products by Chemical Space Map and Machine Learning. Biomolecules 10(10):1385
Capecchi A, Reymond JL (2021) Classifying natural products from plants, fungi or bacteria using the COCONUT database and machine learning. J Chem Inform 13(1):82
Chen TQ, Guestrin C (2016) XGBoost: A scalable tree boosting System. KDD 16:785–794
Chen Y, Stork C, Hirte S, Kirchmair J (2019) NP-Scout: Machine Learning Approach for the Quantification and Visualization of the Natural Product-Likeness of Small Molecules. Biomolecules 9(2):43
Chu J, Vila-Farres X, Inoyama D, Ternei M, Cohen LJ et al (2016) (2016). Discovery of MRSA active antibiotics using primary sequence from the human microbiome. Nat Chem Biol 12:1004–1006
Cimermancic P, Medema MH, Claesen J, Kurita K, Brown W et al (2014) Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158:412–421
Egieyeh S, Syce J, Malan SF, Christoffels A (2018) Predictive classifier models built from natural products with antimalarial bioactivity using machine learning approach. PLoS ONE 13(9):e0204644
Hannigan GD, Prihoda D, Palicka A, Soukup J, Klempir O et al (2019) A deep learning genome-mining strategy for biosynthetic gene cluster prediction. Nucleic Acids Res 47(18):e110
Huang AC, Jiang T, Liu YX, Bai YC, Reed J et al (2019) A specialized metabolic network selectively modulates Arabidopsis root microbiota. Sci 364(eaau6440):6389
Jacoby RP, Koprivova A, Kopriva S (2021) Pinpointing secondary metabolites that shape the composition and function of the plant microbiome. J Exp Bot 72(1):57–69
Kautsar SA, Duran HGS, Blin K, Osbourn A, Medema MH (2017) plantiSMASH: automated identification, annotation and expression analysis of plant biosynthetic gene clusters. Nucleic Acids Res 45(W1):W55–W63
Kautsar SA, Blin K, Shaw S, Navarro-Muñoz JC, Terlouw BR et al (2020) MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res 48:D454–D458
Khaldi N, Seifuddin FT, Turner G et al (2010) SMURF: genomic mapping of fungal secondary metabolite clusters. Fungal Genet Biol 47:736–741
Li MH, Ung PMU, Zajkowski J, Garneau-Tsodikova S, Sherman DH (2009) Automated genome mining for natural products. BMC Bioinformatics 10:185
Li L, Wang H (2016). Towards Label Imbalance in Multi-label Classification with Many Labels. arXiv:1604.01304
Lucaciu R, Pelikan C, Gerner SM, Zioutis C, Köstlbacher S et al (2019) A Bioinformatics Guide to Plant Microbiome Analysis. Front Plant Sci 10:1313
Martín MF, Liras P (1989) Organization and expression of genes involved in the biosynthesis of antibiotics and other secondary metabolites. Annu Rev Microbiol 43:173–206
Martínez-Treviño SH, Uc-Cetina V, Fernández-Herrera María A, Merino Gabriel (2020) Prediction of Natural Product Classes Using Machine Learning and 13C NMR Spectroscopic Data. J Chem Inf Model 60(7):3376–3386
Medema MH, Fischbach MA (2015) Computational approaches to natural product discovery. Nat Chem Biol 11:639–648
Medema MH, Blin K, Cimermancic P, Jager VD, Zakrzewski P et al (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:W339–W346
Mei S, Zhang K (2019) A Multi-Label Learning Framework for Drug Repurposing. Pharmaceutics 11(9):466
Mei S, Zhu H (2015) Multi-label multi-instance transfer learning for simultaneous reconstruction and cross-talk modeling of multiple human signaling pathways. BMC Bioinf 16:417
Milshteyn A, Colosimo DA, Brady SF (2018) Accessing Bioactive Natural Products from the Human Microbiome. Cell Host Microbe 23(6):725–736
Morton JT, Aksenov AA, Nothias LF, Foulds JR, Quinn RA et al (2019) Learning representations of microbe-metabolite interactions. Nat Methods 16(12):1306–1314
Navarro-Muñoz JC, Selem-Mojica N, Mullowney MW, Kautsar SA, Tryon JH et al (2020) A computational framework to explore large-scale biosynthetic diversity. Nat Chem Biol 16:60–68
Nützmann HW, Doerr D, Ramírez-Colmenero A, Sotelo-Fonseca JE, Wegel E et al (2020) Active and repressed biosynthetic gene clusters have spatially distinct chromosome states. Proc Natl Acad Sci U S A 117(24):13800–13809
Palaniappan K, Chen IMA, Chu K, Ratner A, Seshadri R et al (2020) IMG-ABC vol 5.0: an update to the IMG / Atlas of Biosynthetic Gene Clusters Knowledgebase. Nucleic Acids Res 48:D422–D430
Pang Z, Chen J, Wang T, Gao C, Li Z et al (2021) Linking Plant Secondary Metabolites and Plant Microbiomes: A Review. Front Plant Sci 12:621276
Piasecka A, Jedrzejczak-Rey N, Bednarek P (2015) Secondary metabolites in plant innate immunity: conserved function of divergent chemicals. New Phytol 206:948–964
Qiang B, Lai J, Jin H, Zhang L, Liu Z (2021) Target Prediction Model for Natural Products Using Transfer Learning. Int J Mol Sci 22(9):4632
Ren H, Shi C, Zhao H (2020) Computational Tools for Discovering and Engineering Natural Product Biosynthetic Pathways. iSci 23(1):100795
Schlaeppi K, Dombrowski N, Oter RG, van Themaat EVL, Schulze-Leferta P (2014) Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc Natl Acad Sci U S A 111(2):585–592
Schütz V, Frindte K, Cui J, Zhang P, Hacquard S et al (2021) Differential Impact of Plant Secondary Metabolites on the Soil Microbiota. Front Microbiol 12:666010
Sechidis et al (2011) In: Machine learning and knowledge discovery in databases. ECML PKDD 2011. Lect Notes Comput Sci 6913:145–158
Shi MZ, Xie DY (2014) Biosynthesis and Metabolic Engineering of Anthocyanins in Arabidopsis thaliana. Recent Pat Biotechnol 8(1):47–60
Skinnider MA, Johnston CW, Gunabalasingam M, Merwin NJ et al (2020) Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences. Nat Commun 11(1):6058
Smanski MJ, Zhou H, Claesen J, Shen B, Fischbach MA, Voigt CA (2016) Synthetic biology to access and expand nature’s chemical diversity. Nat Rev Microbiol 14:135–149
Starcevic A, Zucko J, Simunkovic J, Long PF, Cullum J, Hranueli D (2008) ClustScan: an integrated program package for the semi-automatic annotation of modular biosynthetic gene clusters and in silico prediction of novel chemical structures. Nucleic Acids Res 36:6882–6892
Tran PN, Yen MR, Chiang CY, Lin HC, Chen PY (2019) Detecting and prioritizing biosynthetic gene clusters for bioactive compounds in bacteria and fungi. Appl Microbiol Biotechnol 103(8):3277–3287
UniProt Consortium (2021) UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 49(D1):D480–D489
van Heel AJ, de Jong A, Song C, Viel JH, Kok J, Kuipers OP (2018) BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res 46(W1):W278–W281
Walker AS, Clardy J (2021) A Machine Learning Bioinformatics Method to Predict Biological Activity from Biosynthetic Gene Clusters. J Chem Inf Model 61(6):2560–2571
Weston J, Bengio S, Usunier N (2011) WSABIE: scaling up to large vocabulary image annotation. Proceed Twenty-Second Int Joint Conf Artif Intell 3:2764–2770
Yu et al (2014) In: Proceedings of the 31st international conference on machine learning, PMLR 32(1):593–601. Beijing, China
Zhang R, Li X, Zhang X, Qin H, Xiao W (2021) Machine learning approaches for elucidating the biological effects of natural products. Nat Prod Rep 38(2):346–361
Funding
None.
Author information
Authors and Affiliations
Contributions
MS conducted the study and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mei, S. A Multi-Label Learning Framework for Predicting Chemical Classes and Biological Activities of Natural Products from Biosynthetic Gene Clusters. J Chem Ecol 49, 681–695 (2023). https://doi.org/10.1007/s10886-023-01452-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10886-023-01452-z