Abstract
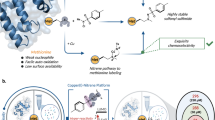
In the presence of formaldehyde and a mild reducing agent, reductive methylation is known to achieve near complete dimethylation of protein amino groups under non-denaturing conditions, in aqueous media. Amino methylation of proteins is employed in mass spectrometric, crystallographic, and NMR studies. Where biosynthetic labeling is prohibitive, amino 13C-methylation provides an attractive option for monitoring folding, kinetics, protein–protein and protein-DNA interactions by NMR. Here, we demonstrate two improvements over traditional 13C-reductive methylation schemes: (1) By judicious choice of stoichiometry and pH, ε-aminos can be preferentially monomethylated. Monomethyl tags are less perturbing and generally exhibit improved resolution over dimethyllysines, and (2) By use of deuterated reducing agents and 13C-formaldehyde, amino groups can be labeled with 13CH2D tags. Use of deutero-13C-formaldehyde affords either 13CHD2, or 13CD3 probes depending on choice of reducing agent. Making use of 13C–2H scalar couplings, we demonstrate a filtering scheme that eliminates natural abundance 13C signal.









Similar content being viewed by others
References
Abe Y, Ueda T, Imoto T (1997) An improved method for preparing lysozyme with chemically C-13-enriched methionine residues using 2-aminothiophenol as a reagent of thiolysis. J Biochem 122:1153–1159
Abraham SJ, Kobayashi T, Solaro RJ, Gaponenko V (2009) Differences in lysine pKa values may be used to improve NMR signal dispersion in reductively methylated proteins. J Biomol NMR 43:239–246
Antos J, Francis M (2006) Transition metal catalyzed methods for site-selective protein modification. Curr Opin Chem Biol 10:253–262
Bokoch MP, Zou Y, Rasmussen SGF, Liu CW, Nygaard R, Rosenbaum DM, Fung JJ, Choi HJ, Thian FS, Kobilka TS, Puglisi JD, Weis WI, Pardo L, Prosser RS, Mueller L, Kobilka BK (2010) Ligand-specific regulation of the extracellular surface of a G-protein-coupled receptor. Nature 463:108–112
Borisenko V, Sansom M, Woolley GA (2000) Protonation of lysine residues inverts cation/anion selectivity in a model channel. Biophys J 78:1335–1348
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRpipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Gerken T, Jentoft J, Jentoft N, Dearborn D (1982) Intramolecular interactions of amino groups in 13C reductively methylated hen egg-white lysozyme. J Biol Chem 257:2894–2900
Goshe MB, Smith RD (2003) Stable isotope-coded proteomic mass spectrometry. Curr Opin Biotechnol 14:101–109
Goux WJ, Teherani J, Sherry AD (1984) Amine inversion in proteins—a C-13-NMR study of proton-exchange and nitrogen inversion rates in n-epsilon, n-epsilon, n-alpha, n-alpha-C-13 tetramethyllysine, n-epsilon, n-epsilon, n-alpha, n-alpha C-13 tetramethyllysine methyl-ester, and reductively methylated concanavalin-A. Biophys Chem 19:363–373
Ikura M, Kay LE, Bax A (1990) A novel-approach for sequential assignment of H-1, C-13, and N-15 spectra of larger proteins—heteronuclear triple-resonance 3-dimensional NMR-spectroscopy—application to calmodulin. Biochemistry 29:4659–4667
Imoto T, Johnson LN, North ACT, Phillips DC, Rupley JA (1972) Lysozyme in enzymes. Academic Press, New York
Johnson BA, Blevins RA (1994) NMR view: a computer-program for the visualization and analysis of NMR data. J Biomol NMR 4:603–614
Joshi N, Whitaker L, Francis M (2004) A three-component Mannich-type reaction for selective tyrosine bioconjugation. J Am Chem Soc 126:15942–15943
Kahsai AW, Rajagopal S, Ahn S, Shukla AK, Sun J, Oas TG, Xiao K, Lefkowitz RJ (2011) Multiple ligand-specific conformations of the beta(2)-adrenergic receptor. Nat Chem Biol 7:692–700
Kalbitzer HR, Rohr G, Nowak E, Goody RS, Kuhn W, Zimmermann H (1992) A new high-sensitivity F-19 probe for labeling cysteine groups of proteins. NMR Biomed 5:347–350
Klein-Seetharaman J, Getmanova E, Loewen M, Reeves P, Khorana H (1999) NMR spectroscopy in studies of light-induced structural changes in mammalian rhodopsin: applicability of solution 19F NMR. Proc Natl Acad Sci USA 96:13744–13749
Korzhnev DM, Mittermaier AK, Kay LE (2005) Cross-correlated spin relaxation effects in methyl H-1 CPMG-based relaxation dispersion experiments: complications and a simple solution. J Biomol NMR 4:337–342
Kumar S, Nussinov R (1999) Salt bridge stability in monomeric proteins. J Mol Biol 293:1241–1255
Kumar S, Tsai CJ, Ma BY, Nussinov R (2000) Contribution of salt bridges toward protein thermostability. J Biomol Struct Dyn 17(suppl 1):79–85
Luchette P, Prosser R, Sanders C (2002) Oxygen as a paramagnetic probe of membrane protein structure by cysteine mutagenesis and F-19 NMR spectroscopy. J Am Chem Soc 124:1778–1781
Macnaughtan MA, Kane AM, Prestegard JH (2005) Mass spectrometry assisted assignment of NMR resonances in reductively C-13-methylated proteins. J Am Chem Soc 127(50):17626–17627
Means GE, Feeney RE (1968) Reductive alkylation of amino groups in proteins. Biochemistry 7:2192–2201
Means GE, Feeney RE (1995) Reductive alkylation of proteins. Anal Biochem 224:1–16
Moore G, Cox M, Crowe D, Osborne M, Rosell F, Bujons J, Barker P, Mauk M, Mauk A (1998) N epsilon, N epsilon-dimethyl-lysine cytochrome c as an NMR probe for lysine involvement in protein–protein complex formation. Biochem J 332:439–449
Moult J, Yonath A, Traub W, Smilansky A, Podjarny A, Rabinovich D, Saya A (1976) Structure of triclinic lysozyme at 2.5 a resolution. J Mol Biol 100:179–195
Muhandiram D, Yamazaki T, Sykes B, Kay L (1995) Measurement of H-2 T-1 and T-1p relaxation-times in uniformly C-13-labeled and fractionally H-2-labeled proteins in solution. J Am Chem Soc 117:11536–11544
Ojida A, Tsutsumi H, Kasagi N, Hamachi I (2005) Suzuki coupling for protein modification. Tetrahedron Lett 46:3301–3305
Oxenoid K, Sonnichsen FD, Sanders CR (2002) Topology and secondary structure of the N-terminal domain of diacylglycerol kinase. Biochemistry 41:12876–12882
Palmer AG, Kroenke CD, Loria JP (2001) Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238
Rayment I (1997) Reductive alkylation of lysine residues to alter crystallization properties of proteins. Methods Enzymol 276:171–179
Rypniewski W, Holden H, Rayment I (1993) Structural consequences of reductive methylation of lysine residues in hen egg white lysozyme: an X-ray analysis at 1.8-Anstrom resolution. Biochemistry 32:9851–9858
Sarakatsannis J, Duan Y (2005) Statistical characterization of salt bridges in proteins. Proteins Struct Funct Bioinf 60:732–739
Sprangers R, Velyvis A, Kay L (2007) Solution NMR of supramolecular complexes: providing new insights into function. Nat Methods 4:697–703
Tieleman DP, Borisenko V, Sansom MSP, Woolley GA (2003) Understanding pH-dependent selectivity of alamethicin K18 channels by computer simulation. Biophys J 84:1464–1469
Tsai CS, Tsai YH, Lauzon G, Cheng ST (1974) Structure and activity of methylated horse liver alcohol-dehydrogenase. Biochemistry 13:440–443
Tugarinov V, Kay LE (2006) A 2H NMR relaxation experiment for the measurement of the time scale of methyl side-chain dynamics in large proteins. J Am Chem Soc 128:12484–12489
Tugarinov V, Ollerenshaw JE, Kay LE (2005) Probing side-chain dynamics in high molecular weight proteins by deuterium NMR spin relaxation: an application to an 82-kDa enzyme. J Am Chem Soc 127:8214–8225
White HD, Rayment I (1993) Kinetic characterization of reductively methylated myosin subfragment 1. Biochemistry 32:9859–9865
Yokoyama S (2003) Protein expression systems for structural genomics and proteomics. Curr Opin Chem Biol 7:39–43
Zhang M, Vogel H (1993) Determination of the side chain pKa values of the lysine residues in calmodulin. J Biol Chem 268:22420
Zhang MJ, Thulin E, Vogel HJ (1994) Reductive methylation and pKa determination of the lysine side-chains in calbindin D-9K. J Protein Chem 13:527–535
Acknowledgments
RSP acknowledges NSERC (Grant number 261980) for a research discovery award. MPB acknowledges support from the Stanford Medical Scientist Training Program.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10858_2012_9664_MOESM1_ESM.doc
Table S1 reports 1H and 13C chemical shifts and linewidths for methylated HEWL.. Figures S2-S7 detail the effect of pH on chemical shifts of mono- and di-methyl lysines. Figure S8 shows the effect of aldehyde stoichiometry on the resulting ratio of mono- to dimethyl lysines for reductively methylated HEWL. Figure S9 and S10 show the effect of aldehyde stoichiometry on the ratio of monoto dimethyl lysines for reductively methylated BSA (DOC 6083 kb)
Rights and permissions
About this article
Cite this article
Larda, S.T., Bokoch, M.P., Evanics, F. et al. Lysine methylation strategies for characterizing protein conformations by NMR. J Biomol NMR 54, 199–209 (2012). https://doi.org/10.1007/s10858-012-9664-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-012-9664-z