Abstract
The most prominent host response to viral infection is the production of type 1 interferons (T1 IFNs). One host regulator of the T1 IFNs is the serine-threonine kinase, tumor progression locus 2 (TPL2). We have previously demonstrated that Tpl2−/− mice succumb to infection with a low-pathogenicity influenza A strain (x31), in association with with increased pulmonary levels of interferon-β (IFN-β), chemokine CCL2, and excessive monocyte and neutrophil pulmonary infiltration. TPL2-dependent overexpression of IFN-β has been implicated in enhanced susceptibility to Mycobacterium tuberculosis; therefore, we examined the role of T1 IFNs in susceptibility of Tpl2−/− mice to influenza. CCL2 overexpression and monocyte recruitment were normalized in Ifnar1−/−Tpl2−/− mice, confirming that TPL2 constrains inflammatory monocyte recruitment via inhibition of the T1 IFN/CCL2 axis. Unexpectedly, excessive neutrophil recruitment in Ifnar1−/− strains was further exacerbated by simultaneous TPL2 genetic ablation in Ifnar1−/−Tpl2−/− by 7 dpi, accompanied by overexpression of neutrophil-regulating cytokines, CXCL1 and IFN-λ. Collectively, our data suggest that TPL2 and T1 IFNs synergize to inhibit neutrophil recruitment. However, treatment with the neutrophil-depleting anti-Ly6G antibody showed only a modest improvement in disease. Analysis of sorted innate immune populations revealed redundant expression of inflammatory mediators among neutrophils, inflammatory monocytes and alveolar macrophages. These findings suggest that targeting a single cell type or mediator may be inadequate to control severe disease characterized by a mixed inflammatory exudate. Future studies will consider TPL2-regulated pathways as potential predictors of severe influenza progression as well as investigate novel methods to modulate TPL2 function during viral infection.
Similar content being viewed by others
Introduction
Respiratory viral infections pose ongoing global challenges due to their rapid evolution and the emergence of new strains with pandemic potential. The ongoing SARS-CoV-2 pandemic has wreaked health, social, and economic devastation globally. However, even seasonal influenza can lead to life-threatening disease in susceptible individuals, including those who are below 6 years of age or above 60 years, as well as people with other risk factors like smoking, heart disease, obesity, pregnancy, and genetic predisposition to interferonopathies [1,2,3,4,5,6]. The variable disease severity experienced by individuals in response to respiratory viral infections highlights the power of the host response to drive disease outcome.
One of the earliest and most prominent host responses to viral infection is the production of antiviral interferons, including type 1 interferons (T1 IFNs) comprising a single IFN-β and thirteen IFN-α subtypes [7, 8] and type 3 interferons (T3 IFNs) comprising up to four species-specific IFN-λs [9, 10]. T1 and T3 IFNs signal via distinct receptors, but both classes of IFNs contribute to the antiviral response by interfering with viral replication via overlapping, yet non-redundant roles [11, 12]. Specifically, T3 IFNs promote early viral control at epithelial barriers without invoking substantial inflammation. In contrast, T1 IFNs are thought to function later on by limiting spread of virus that has escaped T3 IFN control, and in doing so, initiate host inflammatory responses that cause collateral damage to host tissues. Influenza viruses have an enveloped genome of negative sense single-stranded RNA [13], which along with replication intermediates can be sensed by host pathogen recognition receptors (PRRs). The primary PRRs involved in influenza recognition include the RIG-I-like receptor family and Toll-like receptors (TLRs), of which RIG-I activation is the predominant inducer of antiviral interferons (IFNs) [14]. Upon recognition of viral RNA by RIG-I, induction of T1 IFNs [15] leads to activation of TANK-binding kinase 1 (TBK1). TBK1 activates IFN-regulatory factor 3 (IRF3) or TGFβ-activated kinase 1 (TAK1), which ultimately activates NF-κB. Collectively, this leads to IRF3-mediated nuclear transcription of the primary interferons, IFN-β and IFN-α4 [16]. Once secreted, they bind to the heterodimeric interferon alpha receptor (IFNAR1 and IFNAR2) on the cell surface. This leads to phosphorylation of Janus kinases 1 and 2 (JAK1 and JAK2) that, in turn, phosphorylate and activate the signal transducers and activators of transcriptions 1 and 2 (STAT1 and STAT2) [17]. STAT1 and STAT2, together with IRF9, form the interferon-stimulated gene factor 3 (ISGF3) complex. Nuclear translocation of this complex stimulates the expression of IFN-regulatory factor 7 (IRF7), which is required for expression of all other IFN-α subtypes [18]. T1 IFNs also induce other IFN-stimulated genes (ISGs) that collectively mediate the antiviral response [19,20,21,22].
Host factors that control the antiviral immune response make attractive candidates for therapeutics because they are less susceptible to developing viral resistance and have the potential to cross-protect against diverse viruses. One host regulator of the T1 IFN response is the serine-threonine kinase, tumor progression locus 2 (TPL2), also known as MAP3K8 and cancer Osaka thyroid (Cot) [23]. TPL2 is maintained in an inactive state in a trimolecular complex with NF-κB inhibitory protein-1 (NF-κB-1, p105) and A20-binding inhibitor of NF-κB-2 (ABIN-2) [24]. Stimulation of various receptors, including TLRs [25, 26], TNF family receptors [27, 28], IL-1 receptor [29], and some G protein-coupled receptors (GPCR) [30], leads to TPL2 activation downstream of the inhibitor of kappa B kinase (IKK) complex comprising IKKα, IKKβ, and IKKγ [31]. IKK activation leads to NF-κBp105 phosphorylation, ubiquitination, and limited proteosomal degradation that releases TPL2 from its negative regulation [32]. TPL2 kinase activation is subsequently triggered by IKK-mediated phosphorylation of TPL2 [32]. Active TPL2 mediates downstream signaling via multiple pathways, including NF-κB, IRFs, ERK, JNK, and p38 [24, 26, 33], to regulate the production of both pro-inflammatory cytokines and IFNs.
TPL2 differentially regulates T1 IFN production in a context-dependent matter. LPS- and CpG-stimulated Tpl2−/− macrophages and dendritic cells overproduce T1 IFNs compared to WT cells, whereas CpG-stimulated Tpl2−/− plasmacytoid dendritic cells (pDCs) display significantly reduced T1 IFN production [34, 35]. Furthermore, impaired host resistance to both Mycobacterium tuberculosis (M. tuberculosis) and group B Streptococcus has been associated with alterations in the T1 IFN response [36, 37]. Infection of Tpl2−/− mice with M. tuberculosis significantly increased T1 IFN responses. Furthermore, susceptibility of Tpl2−/− mice to M. tuberculosis could be rescued by simultaneously interrupting T1 IFN signaling in Ifnar1−/−Tpl2−/− mice [36], demonstrating the detrimental role of excessive T1 IFNs in the context of TPL2 deficiency during M. tuberculosis infection. Conversely, increased susceptibility of Tpl2−/− mice to group B Streptococcus was associated with a decreased T1 IFN response [37]. Collectively, these studies demonstrate that TPL2-dependent regulation of the T1 IFN response is context dependent and can alter disease pathogenesis.
We have previously shown that Tpl2−/− mice exhibit enhanced morbidity and mortality to influenza infection, with deteriorating clinical symptoms from 7 to 9 dpi [38]. Despite ultimately clearing the virus, Tpl2−/− mice showed hypercytokinemia and excessive pulmonary influx of inflammatory monocytes and neutrophils that correlated with increased expression of T1 IFNs, the interferon-stimulated gene CCL2 (MCP-1), and the inflammatory mediator, nitric oxide synthase 2 (NOS2). Furthermore, we found inefficient regulation of IFN signaling via the positive regulator, signal transducers and activators of transcription 1 (STAT1), and the negative regulator, suppressors of cytokine signaling 1 (SOCS1) [38]. Consequently, influenza-infected Tpl2−/− mice displayed uncontrolled IFN signaling. Considering the dysregulation of IFN signaling in influenza A virus-infected Tpl2−/− mice, the goal of the current study is to determine whether enhanced morbidity and mortality in Tpl2−/− mice during respiratory virus infection are due solely to excessive T1 IFN responses. If so, it may be possible to alleviate severe pathology by abrogation of T1 IFN signaling. Alternatively, additional inflammatory alterations in Tpl2−/− mice may be found to contribute to severe disease.
Results
Interferon Signaling Blockade Is Insufficient to Reverse the Morbidity and Mortality in Influenza-Infected Tpl2 −/− Mice
In order to assess the contribution of T1 IFN signaling to severe disease in influenza-infected Tpl2−/− mice, Tpl2−/− mice were intercrossed with Ifnar1−/− mice to generate Ifnar1−/−Tpl2−/− mice. WT, Tpl2−/−, Ifnar1−/−, and Ifnar1−/−Tpl2−/− mice were infected with 104 pfu influenza A virus strain x31 (IAV x31; H3N2), and their weight loss, clinical symptoms, and survival were monitored over the course of disease (Fig. 1A–C). Clinical symptoms are plotted only through day 9 when peak morbidity and mortality occurred, because loss of the severely ill mice (Fig. 1C) subsequently biased the average clinical scores (Fig. 1B). Similar to what we observed previously [38], WT mice displayed transient weight loss and mild clinical symptoms at 7 dpi and recovered by 9 dpi. In contrast, Tpl2−/− mice showed significantly greater weight loss and more severe clinical symptoms compared to WT mice between 7 and 9 dpi, at which time approximately 60% of Tpl2−/− mice met the humane endpoints of the study and were euthanized (Fig. 1A–C). Ifnar1−/−Tpl2−/− mice showed similarly poor outcomes to influenza infection in terms of weight loss, clinical symptoms, and a trending increase in mortality as shown by a 1-day reduction in mean survival time compared to Tpl2−/− mice (Fig. 1C). Notably, Ifnar1−/− mice also showed trending reductions in mortality compared to the Ifnar1−/−Tpl2−/− mice, with significantly less weight loss and lower clinical scores compared to both Ifnar1−/−Tpl2−/− and Tpl2−/− mice at 9 dpi (Fig. 1A, B). Ifnar1−/− mice that survived beyond 7 dpi recovered their weights similar to WT mice (Fig. 1A). When we stratified the weight loss and clinical score data by sex to evaluate sex as a biological variable, we found that both males and females showed the same trends, with higher weight loss and clinical scores for the Tpl2−/− and Ifnar1−/−Tpl2−/− mice during the later stages of infection (Supplementary Fig. 1). Collectively, these data demonstrate that interferon signaling blockade is insufficient to reverse the heightened morbidity and mortality in influenza-infected Tpl2−/− mice.
Enhanced susceptibility of Tpl2−/− mice to influenza infection is independent of T1 IFN signaling. A Percent weight change and B progression of clinical symptoms of WT (n = 22), Tpl2−/− (n = 29), Ifnar1−/− (n = 40), and Ifnar1−/−Tpl2−/− (n = 28) mice infected with 104 pfu influenza A virus strain x31 through 14 dpi. Data are pooled from 5 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed for each day post infection, with p < 0.05 considered statistically significant. C Survival curve for 14 dpi. Data are pooled from 5 independent experiments. Significance determined with log-rank Mantel-Cox test; *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. Both male and female mice were used for A–C; group averages of male and female mice are shown in the form of diamond symbols.
Increased Inflammatory Monocyte Recruitment to Lungs of Influenza-Infected Tpl2 −/− Mice Is Dependent on T1 IFN Signaling
To gain insight into the contribution of both TPL2 and T1 IFN signaling to inflammatory cellular recruitment that has been linked to severe influenza disease, we first assessed monocyte recruitment in all four strains of mice. Consistent with our previous study [38], we observed an increased influx of monocytes to the lungs of Tpl2−/− mice (Fig. 2A). Notably, this phenotype was reversed in Ifnar1−/−Tpl2−/− mice (Fig. 2A), demonstrating that increased T1 IFN signaling in influenza-infected Tpl2−/− mice is responsible for the increased monocyte recruitment to the lungs. Cytokine and gene expression profiling showed that IFN-β protein and mRNA levels were highest in the lungs of Ifnar1−/−Tpl2−/− (Fig. 2B, D). However, analysis of chemokine CCL2 (MCP-1) showed the highest protein and mRNA levels in lungs of Tpl2−/− mice (Fig. 2C, E). This increased induction of CCL2 protein and mRNA expression by influenza in Tpl2−/− mice was absent in Ifnar1−/−Tpl2−/− mice, consistent with the requirement for IFN signaling for induction of this ISG [39]. Likewise, mRNA expression of other ISGs, including ISG15, IFITM3, and the inducible negative regulator of T1 IFN signaling, SOCS1, were all overexpressed in Tpl2−/− mice, but not Ifnar1−/−Tpl2−/− mice at 7 dpi (Fig. 2F–H), confirming the inability of the Ifnar1−/− and Ifnar1−/−Tpl2−/− strains to transduce T1 IFN signals. These data demonstrate that blockade of T1 IFN signaling in Ifnar1−/−Tpl2−/− mice prevents the excessive monocyte recruitment observed in lungs of Tpl2−/− mice. Therefore, it was unclear why the Ifnar1−/−Tpl2−/− mice remained severely ill.
Enhanced recruitment of inflammatory monocytes and induction of interferon stimulated genes (ISGs) in influenza-infected Tpl2−/− mice is dependent on IFNAR1 signaling at 7 dpi. A WT (n = 20), Tpl2−/− (n = 14), Ifnar1−/− (n = 15), and Ifnar1−/−Tpl2−/− (n = 14) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. The lungs were lavaged, perfused with PBS, digested with collagenase, and interstitial leukocytes were enriched by Percoll density gradient centrifugation. Inflammatory monocytes (Siglec F− CD11bhigh CD11clow Ly6C+) at 7 dpi in the lung tissue of infected mice are shown. Data are pooled from 3 independent experiments. Males are represented as squares and females as circles. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05, **p < 0.01, and ***p < 0.001. WT (n = 22), Tpl2−/− (n = 13), Ifnar1−/− (n = 19), and Ifnar1−/−Tpl2−/− (n = 21) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. The lungs were lavaged and perfused prior to homogenization for analysis of protein levels of B IFN-β and C CCL2 cytokines. Furthermore, the same homogenate was also used for RNA extraction and analysis of gene expression by real-time PCR for D IFN-β and E–H IFN-stimulated genes (ISGs). Squares represent male mice, and circles represent female mice. Data are pooled from 2 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001.
Because T1 IFN signaling is known to mediate viral control, we assessed viral titers in these strains to determine if viral loads were significantly altered. Consistent with our previous study [35], viral titers were modestly increased in Tpl2−/− mice compared to WT mice at 7 dpi (Supplementary Fig. 2A). Additionally, viral titers were only significantly increased in the Ifnar1−/− background if Tpl2 was also ablated, demonstrating that viral loads were higher in the absence of TPL2, irrespective of IFNAR signaling (Supplementary Fig. 2A). However, viral titers did not correlate with morbidity, as measured by weight loss, in influenza-infected Tpl2−/− or Ifnar1−/−Tpl2−/− mice at 7 dpi (Supplemental Fig. 2B).
TPL2 and T1 IFNs Cooperatively Inhibit Neutrophil Recruitment at 7 dpi
Cellular profiling revealed that total numbers of lung alveolar macrophages, natural killer cells, CD4, and CD8 T cells were not altered at 7 dpi by deficiency in either TPL2 or T1 IFN signaling (Supplementary Fig. 3). Besides inflammatory monocytes, the only other cell type that was differentially recruited to the lungs of these mouse strains was neutrophils, with the highest recruitment seen in the Ifnar1−/−Tpl2−/− mice at 7 dpi (Fig. 3A). The fact that neutrophil recruitment was significantly increased in Ifnar1−/−Tpl2−/− compared to Ifnar1−/− or Tpl2−/− mice at 7 dpi suggested a secondary pathway of neutrophil recruitment during influenza infections that is active in the absence of T1 IFN signaling and inhibited by TPL2. In order to delineate this pathway, we examined lung homogenates of mice at 7 dpi and observed increased expression in Ifnar1−/− mice of cytokines previously implicated in neutrophil recruitment [40,41,42,43,44,45], including IFN-γ, IL-6, IL-1β, and G-CSF (Fig. 3B–E). IL-6 and G-CSF were also significantly increased in Ifnar1−/−Tpl2−/− mice compared to WT and Tpl2-/- mice (Fig. 3C, E). Notably, IFN-λ/IL-28 was the only cytokine upregulated solely in the Ifnar1−/−Tpl2−/− mice at the protein (Fig. 3F) and mRNA levels (Supplementary Fig. 4E). Although we did not observe any significant differences in protein expression of the chemokine CXCL1 in the perfused and lavaged lungs of infected mice (Fig. 3G), it was solely upregulated by mRNA expression in the Ifnar1−/−Tpl2−/− mice at 7 dpi (Supplementary Fig. 4F). Because CXCL1 has previously been reported to recruit neutrophils in the absence of interferon signaling during influenza infection [39], we also examined protein expression in intact lungs (without lavage or perfusion) and noted that the expression of CXCL1 is significantly upregulated solely in the lungs of Ifnar1−/−Tpl2−/− mice (Fig. 3H), with no alterations noted for IFN-γ, IL-6, or CCL2 (Fig. 3I–K). Collectively, these data suggest that both TPL2 and T1 IFNs normally inhibit the expression of both CXCL1 and IFN-λ.
IFNAR1−/−Tpl2−/− mice show the highest recruitment of neutrophils and unique overexpression of IFN-λ and CXCL1 at 7 dpi with influenza. A WT (n = 20), Tpl2−/− (n = 14), Ifnar1−/− (n = 15), and Ifnar1−/−Tpl2−/− (n = 14) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. The lungs were lavaged, perfused with PBS, digested with collagenase, and interstitial leukocytes were enriched by Percoll density gradient centrifugation. Neutrophils (Siglec F− CD11bhigh CD11clow Ly6G+) in the tissue of infected mice at 7 dpi are shown. Data are pooled from 3 independent experiments. Males are represented as squares, and females are represented as circles. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05, **p < 0.01, and ***p < 0.001. WT (n = 22), Tpl2−/− (n = 13), Ifnar1−/− (n = 19), and Ifnar1−/−Tpl2−/− (n = 21) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. The lungs were perfused and lavaged prior to extraction and then homogenized for analysis of cytokine expression for B IFN-γ, (C) IL-6, D IL-1β, E G-CSF, F IFN-λ (IL-28), and G CXCL1. Squares represent male mice, and circles represent female mice. Data are pooled from 2 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. WT (n = 17), Tpl2−/− (n = 16), Ifnar1−/− (n = 21), and Ifnar1−/−Tpl2−/− (n = 13) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. Whole lungs (without perfusion or lavage) were homogenized for analysis of cytokine expression for H CXCL1, I IFN-γ, J IL-6, and K CCL2. Squares represent male mice, and circles represent female mice. Data are pooled from 2 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05 and **p < 0.01.
Ifnar1 −/− and Ifnar1 −/− Tpl2 −/− Mice Upregulate CXCL1 Expression and Excessively Recruit Neutrophils to the Lungs as Early as 4 dpi with Influenza
At 7 dpi, we observed the overexpression of IFN-λ and CXCL1 and enhanced recruitment of neutrophils, which are normally involved in the early response to influenza infection [46, 47]. In order to determine if this recruitment occurred earlier and persisted, we examined the cellular recruitment profile of the lungs at 4 dpi. While inflammatory monocytes were not differentially recruited by 4 dpi (Fig. 4A), neutrophils were excessively recruited in both Ifnar1−/− and Ifnar1−/−Tpl2−/− mice by 4 dpi (Fig. 4B). Examination of the cytokine profile of whole lung homogenates showed that CXCL1 expression was significantly upregulated at 4 dpi in the Ifnar1−/−Tpl2−/− mice (Fig. 4C), with no significant changes noted in other cytokines (IFN-γ, IL-6, IFN-λ/IL-28, G-CSF) assessed at the protein level (Fig. 4D, E) or by transcriptional expression (Fig. 4G, H). IFN-β expression was upregulated in the Tpl2−/− mice even at 4 dpi compared to all other strains (Fig. 4I). Accordingly, a significant increase in the expression of the monocyte-recruiting chemokine, CCL2, was also noted in Tpl2−/− mice compared to the Ifnar1−/− and Ifnar1−/−Tpl2−/− mice at the protein (Fig. 4F) and mRNA levels (Fig. 4J). Reduced CCL2 expression in Ifnar1−/− and Ifnar1−/−Tpl2−/− mice is consistent with their block in T1 IFN signaling and failure of these strains to significantly recruit monocytes (Fig. 4A). Collectively, these data suggest that the excessive neutrophil recruitment is observed in both Ifnar1−/− and Ifnar1−/−Tpl2−/− mice by 4 dpi and may be triggered by the induction of CXCL1 in the absence of active T1 IFN signaling.
Neutrophils, but not monocytes, are excessively recruited to the lungs of Ifnar1−/− and Ifnar1−/−Tpl2−/− mouse strains at 4 dpi with upregulation of CXCL1 expression in the absence of T1 IFN signaling. WT (n = 17), Tpl2−/− (n = 9), Ifnar1−/− (n = 8), and Ifnar1−/−Tpl2−/− (n = 17) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 4 dpi, along with uninfected controls. The lungs were lavaged, perfused with PBS, digested with collagenase, and interstitial leukocytes were enriched by Percoll density gradient centrifugation. A Inflammatory monocytes and B neutrophils were examined at 4 dpi along with uninfected controls. Data are pooled from 3 independent experiments. Males are represented as squares, and females are represented as circles. One-way ANOVA with Tukey’s multiple comparison test was performed. *p < 0.05, **p < 0.01, and ***p < 0.001. C–J WT (n = 10), Tpl2−/− (n = 8), Ifnar1−/− (n = 10), and Ifnar1−/−Tpl2−/− (n = 11) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 4 dpi with uninfected controls. The intact lungs were homogenized for analysis of cytokine protein expression for C CXCL1, D IFN-γ, E IL-6, and F CCL2. Furthermore, the same homogenate was also used for RNA extraction and analysis of gene expression by real-time PCR of G IFN-λ3 (IL-28B), H G-CSF, I IFN-β, and J CCL2. Squares represent male mice, and circles represent female mice. Data are pooled from 2 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001.
Anti-Ly6G Treatment only Modestly Improved Survival of Influenza-Infected Ifnar1 −/− Tpl2 −/− Mice
Because neutrophil recruitment to the lungs was dramatically increased in Ifnar1−/−Tpl2−/− mice compared to Tpl2−/− mice by 7 dpi, we hypothesized that a switch from monocyte to neutrophil recruitment induced by IFNAR1 ablation could underlie the severe phenotype of influenza-infected Ifnar1−/−Tpl2−/− mice, rather than a disease rescue. Therefore, we tested whether the administration of the neutrophil-depleting Ly6G antibody (clone 1A8) would ameliorate severe disease in the Ifnar1−/−Tpl2−/− mice. Based on the increased IFN-λ levels in Ifnar1−/−Tpl2−/− mice, we also treated mice with a neutralizing antibody against IFN-λ2/3 (IL-28A/B; clone 244,716). Antibody treatment at 5 and 7 dpi was selected based on the dysregulated cytokine profile and increased morbidity observed in Ifnar1−/−Tpl2−/− mice beginning at 7 dpi (Figs. 1C and 3H); we reasoned that treatment 2 days prior, at 5 dpi, may be sufficient to alter the disease outcome. None of the interventions resulted in a statistically significant improvement in clinical outcome. However, Ly6G treatment of Ifnar1−/−Tpl2−/− mice tended to slightly reduce weight loss (Fig. 5A), decrease average clinical score (Fig. 5B), and increase median survival time by approximately 1.5–2 days compared to isotype control or anti-IFN-λ2/3 treatment, respectively (Fig. 5C). While a modest improvement was noted with Ly6G treatment, it did not substantially reverse the mortality in influenza-infected Ifnar1−/−Tpl2−/− mice. Anti-IFNλ2/3 treatment failed to confer any protection.
Anti-Ly6G treatment is only modestly beneficial for influenza-infected Ifnar1−/−Tpl2−/− mice. Mice were infected intranasally with 104 pfu of influenza A/x31. They were administered either 200 μg isotype control antibody (clone 2A3#BE0089, BioXCell, red line), 200 μg anti-Ly6G antibody (clone 1A8 #BE0075-1, BioXCell, grey line), or 2.5 μg neutralizing anti-IFNλ2/3 antibody (clone 244716 #MAB17892, R&D Systems, dashed black line) intraperitoneally in 200 μl on days 5 and 7 post-infection. Their weight change (A) and clinical scores (B) were recorded daily. Survival (C) was assessed through 12 dpi. Numbers of mice: Ifnar1−/−Tpl2−/− treated with isotype control (n = 4) mice, Ifnar1−/−Tpl2−/− treated with anti-IFNλ2/3 (n = 3), and Ifnar1−/−Tpl2−/− treated with anti-Ly6G (n = 5). Mean and SEM are shown. Data are pooled from 2 independent experiments. Log-rank Mantel-Cox test was used for statistics.
NOS2 Is Overexpressed in Ifnar1 −/− Tpl2 −/− Mice, with Inflammatory Monocytes and Neutrophils Redundantly Contributing to its Expression
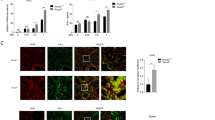
To gain insight into potential drivers of morbidity in influenza-infected Tpl2−/− and Ifnar1−/−Tpl2−/− mice, we examined the lung expression levels of effector molecules implicated in influenza-induced immunopathology, including NOS2, myeloperoxidase (MPO), and TNF-related apoptosis-inducing ligand (TRAIL) [48,49,50]. NOS2 and MPO catalyze the production of reactive nitrogen and reactive oxygen species, respectively [48, 50]. TRAIL, a death receptor ligand, induces cell death via apoptosis [51]. Collectively, these effector molecules constitute part of the innate immune response to pathogens, but they also contribute to tissue damage. Among them, only NOS2 was significantly upregulated in whole lung homogenates of both Ifnar1−/− and Ifnar1−/−Tpl2−/− strains at 7 dpi, although MPO followed a similar trend (Fig. 6A–C). In contrast, TRAIL (encoded by Tnfsf10) trended higher in Tpl2−/− mice compared to the other strains (Fig. 6C). Nitric oxide is expressed in lungs of humans [52, 53], mice [50, 54], and even chickens [55] upon infection with highly pathogenic influenza viruses and is implicated in pulmonary damage. Concomitant with the excessive neutrophil recruitment, this suggested that the neutrophils may partially contribute to severe pathology in Ifnar1−/−Tpl2−/− mice via their NOS2 over-expression (Fig. 6B). However, NOS2 expression alone is insufficient to explain the severe phenotype in Ifnar1−/−Tpl2−/− mice compared to Ifnar1−/− mice (Fig. 1A–C).
The inflammatory mediator NOS2 is overexpressed in the lungs in Ifnar1−/−Tpl2−/− mice, with neutrophils and inflammatory monocytes redundantly contributing to its expression. WT (n = 22), Tpl2−/− (n = 13), Ifnar1−/− (n = 19), and Ifnar1−/−Tpl2−/− (n = 21) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. The lungs were homogenized for RNA extraction and analysis of gene expression by real-time PCR for A MPO, B NOS2, and C TNFSF10. Squares represent male mice, and circles represent female mice. Data are pooled from 2 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed with *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. WT (n = 8), Tpl2−/− (n = 9), Ifnar1−/− (n = 6), and Ifnar1−/−Tpl2−/− (n = 6) mice were infected intranasally with 104 pfu of influenza A/x31 and euthanized at 7 dpi. Their lungs were digested with collagenase, and the following leukocytes populations were sorted: D, G, J, M alveolar macrophages (AM; Siglec Fhigh CD11bint), E, H, K, N neutrophils (Neut; Siglec F− CD11bhigh CD11clow Ly6G+), and F, I, L, O inflammatory monocytes (IM; Siglec F− CD11bhigh CD11clow Ly6C+). Gene expression was analyzed by qPCR, and data were normalized for each cell type relative to the actin endogenous control and the WT strain for a given cell type as baseline, which was designated a value of 1. Data are pooled from 3 independent experiments. One-way ANOVA with Tukey’s multiple comparison test was performed. *p < 0.05 and **p < 0.01. Female mice were used for these experiments.
To examine the cellular roles for both TPL2 and T1 IFNs in influenza-induced inflammation, we next assessed sorted populations of innate immune cells for their expression of cytokines and chemokines known to promote monocyte and neutrophil recruitment. All four strains of mice were infected with influenza A/x31 for 7 days, and lungs were digested to isolate alveolar macrophages, neutrophils, and inflammatory monocytes by fluorescence-activated cell sorting to analyze their gene expression profiles. Gene expression was first normalized to the WT genotype for each cell type, so that a fold change attributed to genotype could be determined. CCL2 has a higher fold expression in Tpl2−/− monocytes and neutrophils compared to the other genotypes, with no differential expression noted in alveolar macrophages (Fig. 6D–F). This expression is consistent with increased CCL2 expression, secondary to increased IFN-β, observed in lungs of Tpl2−/− mice (Fig. 2C, E). There were no differences in the expression fold change of the neutrophil-recruiting chemokine CXCL1 across genotypes of all three cell types examined (Fig. 6G–I). Although not reaching a statistically significant level, there was a trending decrease in IL-1β expression in Tpl2−/− alveolar macrophages and inflammatory monocytes compared to WT, consistent with the TPL2-dependent expression of IL-1β by macrophages and dendritic cells noted previously [56]. Interestingly, simultaneous deletion of both IFNAR1 and TPL2 reversed the TPL2-dependent defect in IL-1β production in both alveolar macrophages and inflammatory monocytes, leading to significantly elevated levels of IL-1β in Ifnar1−/−Tpl2−/− cells, compared to WT or Tpl2−/− cells (Fig. 6J–L). This finding suggests that increased T1 IFN signaling may be indirectly responsible for the impaired IL-1β expression in Tpl2−/− macrophages. No differences were noted across genotypes for the inflammatory mediator NOS2 in either alveolar macrophages or neutrophils (Fig. 6M, N). However, Ifnar1−/− and Ifnar1−/−Tpl2−/− inflammatory monocytes showed a higher fold change of NOS2 expression compared to WT or Tpl2−/− inflammatory monocytes (Fig. 6O). Collectively, these data show an overexpression of CCL2 within Tpl2−/− inflammatory monocytes and neutrophils, whereas the induction of IL-1β and NOS2 was more pronounced in Ifnar1−/−Tpl2−/− inflammatory monocytes (and also alveolar macrophages for IL-1β).
When the relative expression is computed with the WT alveolar macrophages as the baseline to allow comparison across all cell types, there is a relatively even contribution of the different cell types to the overall CCL2 expression (Supplementary Fig. 5). Increased CCL2 expression was noted in Tpl2−/− neutrophils compared to Ifnar1−/−Tpl2−/− neutrophils, with a similar trending increase in the Tpl2−/− inflammatory monocytes. Alveolar macrophages contribute significantly more to the lung CXCL1 expression in both WT and Tpl2−/− mice (Supplementary Fig. 5), whereas neutrophils contribute significantly more to lung IL-1β expression in both WT and Tpl2−/− mice, compared to the other cells (Supplementary Fig. 5). Notably, NOS2 expression appears to be relatively evenly derived from both neutrophils and inflammatory monocytes (despite increased recruitment of neutrophils), with neither playing a predominant role (Supplementary Fig. 5). Collectively, these data do not implicate a single cell type in the expression of the prominent pro-inflammatory mediator, NOS2, but rather suggest that monocytes, neutrophils, and even alveolar macrophages exhibit overlapping and redundant expression of NOS2. The redundant contribution to expression of inflammatory mediators and effector molecules by multiple innate immune cell types may contribute to the poor efficacy of anti-Ly6G treatment in Ifnar1−/−Tpl2−/− mice. Furthermore, the overexpression of additional inflammatory mediators in Ifnar1−/−Tpl2−/− mice, including IL-1β, IFNγ, IL-6, and IFN-λ (Fig. 3B–D), may contribute to damage and morbidity via neutrophil-independent mechanisms.
Discussion
In a previous study of Tpl2−/− mice infected with M. tuberculosis, the authors observed overexpression of IFN-β, similar to what we have previously reported in influenza-infected Tpl2−/− mice [36, 38]. During M. tuberculosis infection, T1 IFNs are known to impair bacterial control [57]. Further assessment of Ifnar1−/−Tpl2−/− mice revealed efficient bacterial control despite the absence of TPL2 when T1 IFN signaling was also prevented [36]. By extension, our initial hypothesis was that Ifnar1−/−Tpl2−/− mice would show better outcomes during influenza infection than Tpl2−/− mice due to the absence of the excessive T1 IFN, as in the case M. tuberculosis infection.
Using Ifnar1−/−Tpl2−/−, mice we addressed genetically whether increased T1 IFN expression exacerbated disease in Tpl2−/− mice by enhancing inflammatory cellular recruitment. Unexpectedly, Ifnar1−/−Tpl2−/− mice experienced similar or more severe disease with faster progression to morbidity than Tpl2−/− mice. Importantly, the increased disease severity could not simply be attributed to loss of viral control in the absence of T1 IFN signaling, as there was no difference in viral load between WT and Ifnar1−/− mice. Viral titers also did not correlate with morbidity in the severely affected strains, Tpl2−/− and Ifnar1−/−Tpl2−/−. Follow-up cellular studies demonstrated that while recruitment of inflammatory monocytes was reversed in Ifnar1−/−Tpl2−/− mice, a switch from primarily monocytic to neutrophilic pulmonary infiltration occurred in Ifnar1−/−Tpl2−/− mice. This switch was triggered by the absence of early interferon signaling (at 4 dpi). Despite the excessive neutrophil recruitment induced by the absence of T1 IFN signaling in both Ifnar1−/− and Ifnar1−/−Tpl2−/− mice at 4 dpi, by 7 dpi neutrophil recruitment was highest in the Ifnar1−/−Tpl2−/− mice. The upregulation of pro-inflammatory and neutrophil-recruiting cytokines (IFN-γ, IL-6, IL-1β, and G-CSF) observed in Ifnar1−/− mice at 7 dpi could be what drives the 40% mortality observed in the Ifnar1−/− mice. Notably, Ifnar1−/− mice still maintained a higher survival rate, reduced weight loss, and lower clinical scores compared to Ifnar1−/−Tpl2−/− mice. This suggested that other TPL2-dependent and IFNAR1-independent (or IFNAR1-inhibited) cytokines play vital roles in the morbidity observed at later stages in Ifnar1−/−Tpl2−/− mice. Consistent with this hypothesis, we noted increased protein levels of IFN-λ and CXCL1 only in the absence of both TPL2 and T1 IFN signaling (Fig. 7). These data suggest that both TPL2 and T1 IFN normally inhibit the expression of the neutrophil-regulating cytokines, CXCL1[39] and IFN-λ[58], which likely contribute to synergistic neutrophil recruitment and increased morbidity and mortality in Ifnar1−/−Tpl2−/− mice.
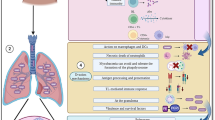
TPL2- and IFN-dependent regulation of influenza A virus inflammation. In the lungs of influenza-infected Tpl2−/− mice at 7 dpi, upregulation of the IFNs and chemokines (CCL2, CCL3) leads to recruitment and retention of inflammatory monocytes and neutrophils, respectively, that collectively lead to lung damage likely via a combination of neutrophil and monocyte-derived effectors, such as NOS2, TRAIL, and MPO. In influenza-infected Ifnar1−/− mice at 7 dpi, overexpression of IFNAR-suppressed cytokines, including IL-1β, G-CSF, IL-6, and IFN-γ, leads primarily to a neutrophilic pulmonary infiltrate. In the context of influenza infection, IFNAR1 deficiency causes some morbidity, although not to the same extent as experienced by Tpl2−/− mice. Both IFNAR1 and TPL2 cooperate to suppress neutrophil chemotactic cytokines, CXCL1 and IFN-λ. Accordingly, Ifnar1−/−Tpl2−/− mice exhibit the highest levels of CXCL1 and IFN-λ expression along with the highest neutrophil recruitment. Concomitant upregulation of other pro-inflammatory mediators, IL-1β, G-CSF, IL-6, and IFN-γ, due to abrogated IFNAR signaling, also likely contributes to inflammation in Ifnar1−/−Tpl2−/− mice via neutrophil-independent mechanisms.
One aspect of inflammation that this study highlights is the role of overactive IFN signaling in Tpl2−/− mice that promotes inflammatory monocyte recruitment directly via the IFN-β/CCL2 axis. In this regard, we attribute the better survival of Ifnar1−/− mice compared to Tpl2−/− mice to the increased monocytic infiltration in Tpl2−/− mice. Monocyte recruitment was increased solely in Tpl2−/− mice, and this increased recruitment was associated with increased CCL2 expression (Fig. 2A, C). Increased CCL2 expression is also evident in both the monocyte and neutrophil sorted cell populations (Fig. 6E, F), suggesting that these cells feed-forward to promote increased inflammatory monocyte recruitment. It is also notable that TRAIL levels (encoded by Tnfsf10) trend highest in the lungs of influenza-infected Tpl2−/− mice. It is possible that increased monocytes present in influenza-infected Tpl2−/− mice promote inflammation and pulmonary damage through TRAIL or other monocytic mediators that were not examined. The upregulation of IFN-β at both the protein and mRNA levels in the Ifnar1−/− strains (Fig. 2B, D) may initially appear paradoxical. However, unlike other IFN-αs and ISGs, IFN-β does not require feedback through the IFNAR1 for its expression [16, 59, 60]. Accordingly, none of the other ISGs was upregulated in the absence of IFNAR1, confirming that T1 IFN signaling was abrogated in the Ifnar1−/− strains. Moreover, similar to our previous study wherein IFN-β was overexpressed in Tpl2−/− mice at 7 dpi [38], IFN-β is also upregulated in Ifnar1−/−Tpl2−/− mice compared to Ifnar1−/− mice, further confirming that TPL2 negatively regulates IFN-β production during influenza infection.
CXCL1 has been shown to be upregulated and to stimulate neutrophil recruitment in the absence of IFNAR signaling during influenza infection [39]. However, due to the multitude of cytokines upregulated in the absence of T1 IFN signaling, increased CXCL1 expression could be an indirect by-product of regulation by IFN-γ, IL-6, IL-1β, G-CSF, or IFN-λ [41,42,43,44, 58]. To explore this possibility, we examined the lung cytokine profile at 4 dpi. At this time point, CXCL1 was only over-expressed in the absence of T1 IFN signaling in Ifnar1−/− strains. Notably, overexpression of CXCL1, but none of the other cytokines under consideration, at 4 dpi led us to conclude that increased CXCL1 may be the initial driving force for enhanced neutrophil recruitment in Ifnar1−/−Tpl2−/− mice. Increased T1 IFN signaling suppresses CXCL1 expression in Tpl2−/− mice, whereas the absence of T1 IFN signaling leads to overexpression of CXCL1 in Ifnar1−/−Tpl2−/− at 7 dpi.
IFN-λ was solely upregulated in Ifnar1−/−Tpl2−/− mice compared to all other strains. Primarily known for being an early response cytokine [47], it is interesting that IFN-λ was upregulated in the late stages of influenza infection, similar to IFN-β. Along with T1 IFNs, IFN-λ expression is involved in viral control, especially in reducing the spread of virus from upper respiratory tract to the lungs and transmission to a naive host [61, 62]. However, despite having similar functions, the receptor for IFN-λ is distinct from the T1 IFNs and its expression is restricted to epithelium and neutrophils [9, 58]. Specifically, IFN-λ expression has been shown to promote optimal antiviral responses from neutrophils, prevent excessive inflammation, and induce better viral control from the epithelium during influenza infection [58]. In another study, overexpression of IFN-λ induced higher proliferation of mature NK cells in spleen, lung, and lymph nodes that were better able to defend against lethal influenza infection [63]. Although T1 IFNs and IFN-λ share an overlapping infection-induced transcriptional signature [12], IFN-λ appears to have more anti-inflammatory functions. In a previous study by our lab, we observed reduced levels of IFN-λ in both bronchoalveolar lavage fluid and lung homogenates in the absence of TPL2 at 3 dpi, irrespective of T1 IFN signaling [35]. IFN-λ expression was also downregulated in Tpl2−/− pDCs, which are early producers of IFN-λ. This could contribute to the dysregulation of viral control seen from the very early stages in Tpl2−/− strains [35], consistent with the importance of IFN-λ expression in early viral control [58, 62]. Recent studies have highlighted a new pro-inflammatory role for IFN-λ in the later stages of influenza infection via inhibition of lung tissue repair [64]. Therefore, it is possible that late-stage expression of IFN-λ contributes to the excessive morbidity and mortality observed in Ifnar1−/−Tpl2−/− mice by preventing lung tissue repair, although we did not see any beneficial effect of blocking IFN-λ in the Ifnar1−/−Tpl2−/− mice to support this idea.
IL-1β has been implicated in neutrophil recruitment [43] and was therefore also examined. Unexpectedly, IL-1β was significantly overexpressed in influenza-infected Ifnar1−/−Tpl2−/− mice. This was unexpected because IL-1β overexpression was not observed in influenza-infected Tpl2−/− mice at 7 dpi in our previous study [38]. Furthermore, IL-1β expression has been shown to be TPL2-dependent in vitro in LPS-stimulated macrophages and in vivo in response to Listeria monocytogenes infection [56]. IL-1β overexpression in Ifnar1−/−Tpl2−/− mice in the current study suggests that T1 IFN overexpression in Tpl2−/− mice may indirectly suppress IL-1β expression.
In order to examine the cellular source of the various inflammatory mediators examined here, sorted neutrophils, inflammatory monocytes, and alveolar macrophages were specifically analyzed. While inflammatory monocytes and neutrophils were selected based on their higher recruitment numbers, alveolar macrophages were considered because they are potent producers of T1 IFN during influenza infection [65, 66], and their influenza-induced depletion [67] increases infection susceptibility for alveolar epithelial cells [68]. While CCL2 expression is upregulated in both Tpl2−/− inflammatory monocytes and neutrophils, NOS2 is upregulated in the IFNAR1-deficient inflammatory monocytes despite their lower recruitment numbers compared to neutrophils. This suggests that neutrophils and inflammatory monocytes might redundantly compensate for each other in promoting inflammation, particularly in the absence of TPL2 or T1 IFN signaling. This has also been seen in other studies, wherein the absence of Ifnar1−/− signaling led to higher expression of NOS2 in both Ly6Chi and Ly6Clo mononuclear subsets [41]. Notably, anti-Ly6G treatment afforded a slight survival advantage to Ifnar1−/−Tpl2−/− mice compared to isotype control or IFN-λ treatment, although it was unable to prevent death. This could be explained by the redundancy in NOS2 expression (and potentially other inflammatory mediators) by Ifnar1−/−Tpl2−/− monocytes and neutrophils, in which case neutrophils could be compensated by monocytes. Furthermore, the overexpression of IFN-λ, IL-1β, IFNγ, and IL-6 in Ifnar1−/−Tpl2−/− mice may contribute to inflammation and morbidity via neutrophil-independent mechanisms. However, we also cannot exclude the possibility that earlier treatment may be necessary to further improve clinical benefit.
Overall, this study highlights the regulation of cytokines, chemokines, and cellular recruitment under the control of TPL2. Prominent among these is the negative regulation of the T1 IFNs, which in turn positively and negatively regulate CCL2 and CXCL1 expression, respectively. This dichotomy leads to a T1 IFN-induced switch from the pathologic increase in pulmonary recruitment of inflammatory monocytes in Tpl2−/− mice to neutrophils in Ifnar1−/−Tpl2−/− mice. Tissue damage may be further exacerbated by dysregulated IFN-λ, as noted above. Future studies will consider TPL2-regulated pathways as potential predictors of severe influenza progression as well as investigate novel methods to modulate TPL2 function during viral infection. Additionally, influenza-infected Tpl2−/− mice may represent a novel preclinical model to test for intervention strategies.
Materials and Methods
Ethics Statement and Study Design
All animal experiments were performed in accordance with “The Guide for Care and Use of Laboratory Animals” and were approved by the University of Georgia Institutional Animal Care and Use Committee (IACUC). Data in this study are reported in accordance with ARRIVE guidelines. Six- to 8-week-old male and female mice (WT, Tpl2−/−, Ifnar1−/−, and Ifnar1−/−Tpl2−/−) were randomized based on age and gender to groups for infection and analysis such that these variables were equally distributed between groups. Exclusion criteria were based solely on the human endpoints of the study as detailed below. All data are reported; no outliers were excluded. The number of mice (n) and the number of replicate experiments are reported for each data set. Outcome measures included lung virus titers, lung gene expression, lung cytokine protein expression, analysis of lung cellular composition, and clinical features.
Mice and Influenza Infections
Wild-type (WT) C57BL/6 (JAX strain #000664) and Rag1−/− mice (strain #002216, strain name B6.129S7-Rag1tm1Mom/J) were bred in-house. Tpl2−/− mice backcrossed at least nine generations onto the C57BL/6 strain were kindly provided by Dr. Philip Tsichlis [25]. Ifnar1−/− mice (B6.129S2-Ifnar1tm1Agt/Mmjax; #032045-JAX) were kindly provided by Dr. Biao He (University of Georgia). Tpl2−/− mice were intercrossed with Ifnar1−/− mice to produce Ifnar1−/−Tpl2−/− mice. Animals were housed in microisolator cages in the Central Animal Facility of the College of Veterinary Medicine.
Age-matched, 6- to 8-week-old WT, Tpl2−/−, Ifnar1−/−, and Ifnar1−/−Tpl2−/− mice were anesthetized with approximately 250 mg/kg of 2% weight/volume avertin (2,2,2- Tribromoethanol, Sigma) followed by intranasal instillation of 50 µl PBS containing 104 pfu of influenza A/HKX31 (H3N2, hereafter referred to as x31). The stock virus was propagated in specific pathogen-free eggs (Poultry Diagnostics and Research Center, UGA) and then titered as described previously [38].
Mice were studied for their susceptibility to infection by measuring daily weight loss and clinical scores according to the following index: piloerection, 1 point; hunched posture, 2 points; rapid breathing, 3 points. Mice with a cumulative score of 5 or that had lost 30% of their initial body weight were humanely euthanized.
Tissue Collection and Analysis for Virus Titers, Cytokines, and Gene Expression
Mice were sacrificed at 7 dpi. The lungs were lavaged with 1 ml of PBS instilled twice into the lungs and then perfused with 10 ml of PBS injected directly into the right ventricle of the heart. Alternatively, some whole lungs were homogenized without lavage or perfusion, where noted. Lungs were harvested into 1 ml of PBS and homogenized in a bead mill homogenizer (Qiagen Tissue Lyser II) at 25 Hz for 2–4 min. The homogenate was centrifuged at 500 × g for 5 min, and the pre-cleared homogenate was (1) directly aliquoted for viral titer assessment with the titration protocol described previously [38], (2) lysed in TRK tissue lysis buffer (E.Z.N.A Omega Bio-Tek) for analysis of gene expression, or (3) further centrifuged at 5000 × g for 5 min to clarify the homogenate for cytokine analysis by ELISA. For gene expression analysis, RNA was extracted, converted to cDNA using a high capacity RNA-to-cDNA kit (Thermo Fisher), and assessed for gene expression. Sensifast Probe Hi-ROX kit (BIO-82020 Bioline) and the following probes (Applied Biosystems) were used: IFN-β1 (Mm00439552), IFN-γ (Mm0116813), IFN-λ3 (Mm00663660), IFITM3 (Mm00847057), IL-6 (Mm00446190), IL-1β (Mm00434228), ISG15 (Mm01705338), CCL2 (Mm00441242), CXCL1 (Mm04207460), TNFSF10 (Mm01283606), NOS2 (Mm00440502), MPO (Mm01298424), and SOCS1 (Mm01342740). The qPCR was run on a StepOne Plus instrument (Applied Biosystems) and gene expression performed using the ΔΔCT method as describe previously [38] relative to an internal actin control and normalized to the WT samples set to a value of 1. Cytokine quantitation was performed using a mouse inflammation cytometric bead array (CBA, analytes IL-6, IFN-γ, MCP-1, TNF, IL-10, and IL-12p70, Becton Dickenson), Mouse ProcartaPlex (analytes IL-1β, IL-28, GM-CSF, Invitrogen), standard murine ABTS ELISA Development kit (CXCL1, IFN-γ, IL-6, CCL2, and IL-1β, Peprotech), and LumiKine Xpress kits (IFN-β, Invivogen).
Cellular Analysis
At 4 and 7 dpi, we isolated lung cells as previously described [38] and assessed lung cellular composition. Cells were stained at 4°C for 20 min with fluorescently labeled antibodies against the following cell surface markers purchased from eBiosciences unless otherwise noted: Siglec F PE (clone IRNM44N), CD11b APC (clone M1/70), CD11c Efluor 450 (clone N418), Ly6C PE-Cy7 (clone HK1.4), Ly6G PerCP-Cy5.5 (Tonbo, clone 1A8), CD45.2 FITC (clone 104) [Stain 1]; TCRαβ FITC (clone H57-597), TCRγδ APC (clone eBioGL3), CD4 PE-Cy7 (clone GK1.5), CD8 Efluor 450 (clone 53–6.7), DX5 PE (clone DX5), CD45.2 PerCP-Cy5.5 (clone 104) [Stain 2]. The cells were fixed with 1% formalin and analyzed on the LSR II flow cytometer (BD Biosciences). CD45.2-gated hematopoietic-derived leukocyte populations were characterized as follows: inflammatory monocytes (IM; Siglec F− CD11bhigh CD11clow Ly6C+), neutrophils (Siglec F− CD11bhigh CD11clow Ly6G+), alveolar macrophages (AM; Siglec Fhigh CD11bint), eosinophils (Siglec Fhigh CD11bhigh), NK cells (αβ TCR− DX5+), CD4 T cells (αβ TCR+ CD4+), CD8 T cells (αβ TCR+ CD8+), and γδ T cells (αβ TCR− γδ TCR+).
Cellular Sort for Gene Expression Analysis
At 7 dpi, lungs from all four genotypes of mice were harvested, digested, and single-cell suspensions were generated as described above. Cells were stained with stain 1 above, and inflammatory monocytes, neutrophils, and alveolar macrophages were isolated by fluorescence-activated cell sorting (FACS) using the same gating strategy on a MoFlo Astrios (BD Biosciences). Sorted populations of cells were lysed in TRK lysis buffer; RNA was extracted and converted into cDNA. Gene expression was quantified as described above with the results expressed relative to the actin internal control and the WT sample for each cell type set to 1 using the ΔΔCT method. Alternatively, to facilitate comparisons across cell types, gene expression was also calculated relative to the actin internal control and the WT alveolar macrophages (AM) as the baseline.
Therapeutic Antibody Interventions to Treat Influenza
Age-matched, 6- to 8-week-old, WT, Tpl2−/−, Ifnar1−/−, and Ifnar1−/−Tpl2−/− mice were infected intranasally with 104 pfu of mouse-adapted influenza A/x31 virus. On days 5 and 7 post infection, the mice were administered intraperitoneally (i.p.) either 200 µg isotype control antibody (clone 2A3#BE0089, BioXCell), 200 µg anti-Ly6G antibody (clone 1A8 #BE0075-1, BioXCell), or 2.5 µg anti-IFN-λ2/3 antibody (clone 244,716 #MAB17892, R&D Systems). Mice were weighed and scored for clinical symptoms throughout the course of infection and treatment.
Statistical Analysis
P values were calculated with GraphPad PRISM software version 9.2.0(332) using one-way ANOVA with Tukey’s multiple comparisons test. Data represent means ± SEM. Survival data are graphed as Kaplan–Meier plots using GraphPad PRISM software, and p values were determined by Mantel-Cox test. Differences were considered statistically significant if p ≤ 0.05. Gaussian correlation was performed to calculate the coefficient based on Pearson’s correlation test with a two-tailed test. Simple linear regression analysis was also performed to analyze the best fit value or the slope and intercept to see if the two variables being compared for a particular genotype correlated or not (as seen by the straight dashed line). Additionally, the confidence interval was set at 95% (as seen by the curved dashed line).
Availability of Data and Materials
Any resources generated from this project will be made available to the scientific community upon publication as required by the NIH. Individual transgenic alleles are commercially available or available upon request through MTA from the original source. New, compound knock-out strains will be made available with permission from the license holders of the individual alleles. All data are available within the document. No large data sets were generated.
References
Monto, A.S., S. Gravenstein, M. Elliott, M. Colopy, and J. Schweinle. 2000. Clinical signs and symptoms predicting influenza infection. Archives of Internal Medicine 160: 3243–3247. https://doi.org/10.1001/archinte.160.21.3243.
Neuzil, Kathleen Maletic, George W. Reed, Edward F. Mitchel, and Marie R. Griffin. 1999. Influenza-associated morbidity and mortality in young and middle-aged women. Journal of the American Medical Association 281: 901–907. https://doi.org/10.1001/jama.281.10.901.
Martínez, Ana, Núria. Soldevila, Arantxa Romero-Tamarit, Núria. Torner, Pere Godoy, Cristina Rius, Mireia Jané, and Àngela. Domínguez. 2019. Risk factors associated with severe outcomes in adult hospitalized patients according to influenza type and subtype. PLoS ONE 14: 1–15. https://doi.org/10.1371/journal.pone.0210353.
Walker, Tiffany A, Ben Waite, Mark G Thompson, Colin McArthur, Conroy Wong, Michael G Baker, Tim Wood, et al. 2020. Risk of severe influenza among adults with chronic medical conditions. The Journal of infectious diseases 221. United States: 183–190. https://doi.org/10.1093/infdis/jiz570.
Mertz, Dominik, Calvin Ka Fung Lo, Lyubov Lytvyn, Justin R. Ortiz, Mark Loeb, Li Wei Ang, Mehta Asmita Anlikumar, et al. 2019. Pregnancy as a risk factor for severe influenza infection: An individual participant data meta-analysis. BMC Infectious Diseases 19. BMC Infectious Diseases: 1–10. https://doi.org/10.1186/s12879-019-4318-3.
Miyashita, Koichi, Eiji Nakatani, Hironao Hozumi, Yoko Sato, Yoshiki Miyachi, and Takafumi Suda. 2021. Risk factors for pneumonia and death in adult patients with seasonal influenza and establishment of prediction scores: A population-based study. Open Forum Infectious Diseases 8: 1–8. https://doi.org/10.1093/ofid/ofab068.
Platanias, Leonidas C. 2005. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nature Reviews Immunology 5: 375–386. https://doi.org/10.1038/nri1604.
Gibbert, K., J.F. Schlaak, D. Yang, and U. Dittmer. 2013. IFN-α subtypes: Distinct biological activities in anti-viral therapy. British Journal of Pharmacology 168: 1048–1058. https://doi.org/10.1111/bph.12010.
Kotenko, Sergei V., Grant Gallagher, Vitaliy V. Baurin, Anita Lewis-Antes, Meiling Shen, Nital K. Shah, Jerome A. Langer, Faruk Sheikh, Harold Dickensheets, and Raymond P. Donnelly. 2003. IFN-λs mediate antiviral protection through a distinct class II cytokine receptor complex. Nature Immunology 4: 69–77. https://doi.org/10.1038/ni875.
Prokunina-Olsson, Ludmila, Brian Muchmore, Wei Tang, Ruth M. Pfeiffer, Heiyoung Park, Harold Dickensheets, Dianna Hergott, et al. 2013. A variant upstream of IFNL3 (IL28B) creating a new interferon gene IFNL4 is associated with impaired clearance of hepatitis C virus. Nature genetics 45: 164–171. https://doi.org/10.1038/ng.2521.
Davidson, Sophia, Teresa M McCabe, Stefania Crotta, Hans Henrik Gad, Edith M Hessel, Soren Beinke, Rune Hartmann, and Andreas Wack. 2016. IFN λ is a potent anti‐influenza therapeutic without the inflammatory side effects of IFN α treatment . EMBO Molecular Medicine 8: 1099–1112. https://doi.org/10.15252/emmm.201606413.
Crotta, Stefania, Sophia Davidson, Tanel Mahlakoiv, Christophe J. Desmet, Matthew R. Buckwalter, Matthew L. Albert, Peter Staeheli, and Andreas Wack. 2013. Type I and type III interferons drive redundant amplification loops to induce a transcriptional signature in influenza-infected airway epithelia. PLoS Pathogens 9. https://doi.org/10.1371/journal.ppat.1003773
Bouvier, Nicole M. xand Palese*, Peter. 2008. The biology of influenza viruses. Vaccine 26: D49–D53. https://doi.org/10.1002/ejoc.201701499
Goffic, Le., Julien Pothlichet Ronan, Damien Vitour, Takashi Fujita, Eliane Meurs, Michel Chignard, and Mustapha Si-Tahar. 2007. Cutting edge: Influenza A virus activates TLR3-dependent inflammatory and RIG-I-dependent antiviral responses in human lung epithelial cells. The Journal of Immunology 178: 3368–3372. https://doi.org/10.4049/jimmunol.178.6.3368.
Pommerenke, Claudia, Esther Wilk, Barkha Srivastava, Annika Schulze, Natalia Novoselova, Robert Geffers, and Klaus Schughart. 2012. Global transcriptome analysis in influenza-infected mouse lungs reveals the kinetics of innate and adaptive host immune responses. PLoS ONE 7. https://doi.org/10.1371/journal.pone.0041169
Honda, Kenya, Akinori Takaoka, and Tadatsugu Taniguchi. 2006. Type I inteferon gene induction by the interferon regulatory factor family of transcription factors. Immunity 25: 349–360. https://doi.org/10.1016/j.immuni.2006.08.009.
Soto, Jorge A., Nicolas M.S.. Gálvez, Catalina A. Andrade, Gaspar A. Pacheco, Karen Bohmwald, Roslye V. Berrios, Susan M. Bueno, and Alexis M. Kalergis. 2020. The role of dendritic cells during infections caused by highly prevalent viruses. Frontiers in Immunology 11: 1–22. https://doi.org/10.3389/fimmu.2020.01513.
Sato, Mitsuharu, Naoki Hata, Masataka Asagiri, Takeo Nakaya, Tadatsugu Taniguchi, and Nobuyuki Tanaka. 1998. Positive feedback regulation of type I IFN genes by the IFN-inducible transcription factor IRF-7. FEBS Letters 441: 106–110. https://doi.org/10.1016/S0014-5793(98)01514-2.
Lee, SangJoon, Akari Ishitsuka, Masayuki Noguchi, Mikako Hirohama, Yuji Fujiyasu, Philipp P Petric, Martin Schwemmle, Peter Staeheli, Kyosuke Nagata, and Atsushi Kawaguchi. 2019. Influenza restriction factor MxA functions as inflammasome sensor in the respiratory epithelium. Science immunology 4. Department of Infection Biology, Faculty of Medicine, University of Tsukuba, Tsukuba, Japan.: American Association for the Advancement of Science. https://doi.org/10.1126/sciimmunol.aau4643.
Poddar, Subhajit, Jennifer L. Hyde, Matthew J. Gorman, Michael Farzan, and Michael S. Diamond. 2016. The interferon-stimulated gene IFITM3 restricts infection and pathogenesis of arthritogenic and encephalitic alphaviruses. Journal of Virology 90: 8780–8794. https://doi.org/10.1128/jvi.00655-16.
Clemens, M J, and B R Williams. 1978. Inhibition of cell-free protein synthesis by pppA2’p5’A2’p5’A: A novel oligonucleotide synthesized by interferon-treated L cell extracts. Cell 13. United States: 565–572. https://doi.org/10.1016/0092-8674(78)90329-x.
Lenschow, Deborah J., Caroline Lai, Natalia Frias-Staheli, Nadia V. Giannakopoulos, Andrew Lutz, Thorsten Wolff, Anna Osiak, et al. 2007. IFN-stimulated gene 15 functions as a critical antiviral molecule against influenza, herpes, and Sindbis viruses. Proceedings of the National Academy of Sciences of the United States of America 104: 1371–1376. https://doi.org/10.1073/pnas.0607038104.
Miyoshi, J., T. Higashi, H. Mukai, T. Ohuchi, and T. Kakunaga. 1991. Structure and transforming potential of the human cot oncogene encoding a putative protein kinase. Molecular and Cellular Biology 11: 4088–4096. https://doi.org/10.1128/mcb.11.8.4088-4096.1991.
Gantke, Thorsten, Srividya Sriskantharajah, and Steven C. Ley. 2011. Regulation and function of TPL-2, an IB kinase-regulated MAP kinase kinase kinase. Cell Research 21. Nature Publishing Group: 131–145. https://doi.org/10.1038/cr.2010.173.
Dumitru, Calin D., Jeffrey D. Ceci, Christos Tsatsanis, Dimitris Kontoyiannis, Konstantinos Stamatakis, Jun Hsiang Lin, Christos Patriotis, et al. 2000. TNF-α induction by LPS is regulated posttranscriptionally via a Tpl2/ERK-dependent pathway. Cell 103: 1071–1083. https://doi.org/10.1016/S0092-8674(00)00210-5.
Kate, Senger, Pham Victoria C., Varfolomeev Eugene, Hackney Jason A., Corzo Cesar A., Collier Jenna, Lau Vivian W C., et al. 2017. The kinase TPL2 activates ERK and p38 signaling to promote neutrophilic inflammation. Science Signaling 10. American Association for the Advancement of Science: eaah4273. https://doi.org/10.1126/scisignal.aah4273.
Pattison, Michael J., Olivia Mitchell, Helen R. Flynn, Chao Sheng Chen, Huei Ting Yang, Ben Addi Hakem, Stefan Boeing, Ambrosius P. Snijders, and Steven C. Ley. 2016. TLR and TNF-R1 activation of the MKK3/MKK6-p38α axis in macrophages is mediated by TPL-2 kinase. Biochemical Journal 473: 2845–2861. https://doi.org/10.1042/BCJ20160502.
Eliopoulos, Aristides G., Chun Chi Wang, Calin D. Dumitru, and Philip N. Tsichlis. 2003. Tpl2 transduces CD40 and TNF signals that activate ERK and regulates IgE induction by CD40. EMBO Journal 22: 3855–3864. https://doi.org/10.1093/emboj/cdg386.
Stafford, Margaret, Nick Morrice, Mark Peggie, and Philip Cohen. 2006. Interleukin-1 stimulated activation of the cot catalytic subunit through the phosphorylation of THR290 and SER62. FEBS letters 580: 4010–4014. https://doi.org/10.1016/j.febslet.2006.06.004.
Maria, Hatziapostolou, Koukos Georgios, Polytarchou Christos, Kottakis Filippos, Serebrennikova Oksana, Kuliopulos Athan, and Tsichlis Philip N. 2011. Tumor progression locus 2 mediates signal-induced increases in cytoplasmic calcium and cell migration. Science Signaling 4. American Association for the Advancement of Science: ra55–ra55. https://doi.org/10.1126/scisignal.2002006.
Israël, Alain. 2010. The IKK complex, a central regulator of NF-kappaB activation. Cold Spring Harbor perspectives in biology 2: 1–14. https://doi.org/10.1101/cshperspect.a000158.
Waterfield, Michael, Wei Jin, William Reiley, Minying Zhang, and Shao-Cong. Sun. 2004. IκB kinase is an essential component of the Tpl2 signaling pathway. Molecular and Cellular Biology 24: 6040–6048. https://doi.org/10.1128/mcb.24.13.6040-6048.2004.
Tsatsanis, Christos, Katerina Vaporidi, Vassiliki Zacharioudaki, Ariadne Androulidaki, Yuri Sykulev, Andrew N. Margioris, and Philip N. Tsichlis. 2008. Tpl2 and ERK transduce antiproliferative T cell receptor signals and inhibit transformation of chronically stimulated T cells. Proceedings of the National Academy of Sciences of the United States of America 105: 2987–2992. https://doi.org/10.1073/pnas.0708381104.
Kaiser, Frank, Dorthe Cook, Stamatia Papoutsopoulou, Ricardo Rajsbaum, Wu. Xuemei, Huei Ting Yang, Susan Grant, et al. 2009. TPL-2 negatively regulates interferon-β production in macrophages and myeloid dendritic cells. Journal of Experimental Medicine 206: 1863–1871. https://doi.org/10.1084/jem.20091059.
Kuriakose, Teneema, Ralph A. Tripp, and Wendy T. Watford. 2015. Tumor progression locus 2 promotes induction of IFNλ, interferon stimulated genes and antigen-specific CD8+ T cell responses and protects against influenza virus. PLoS Pathogens 11: 1–22. https://doi.org/10.1371/journal.ppat.1005038.
McNab, Finlay W., John Ewbank, Ricardo Rajsbaum, Evangelos Stavropoulos, Anna Martirosyan, Paul S. Redford, Wu. Xuemei, et al. 2013. TPL-2–ERK1/2 signaling promotes host resistance against intracellular bacterial infection by negative regulation of type I IFN production. The Journal of Immunology 191: 1732–1743. https://doi.org/10.4049/jimmunol.1300146.
Xiao, Nengming, Celine Eidenschenk, Philippe Krebs, Katharina Brandl, Amanda L. Blasius, Yu. Xia, Kevin Khovananth, Nora G. Smart, and Bruce Beutler. 2009. The Tpl2 mutation sluggish impairs type I IFN production and increases susceptibility to group B streptococcal disease. The Journal of Immunology 183: 7975–7983. https://doi.org/10.4049/jimmunol.0902718.
Latha, Krishna, Katelyn F Jamison, and Wendy T Watford. 2021. Tpl2 ablation leads to hypercytokinemia and excessive cellular infiltration to the lungs during late stages of influenza infection . Frontiers in Immunology .
Seo, Sang Uk, Hyung Joon Kwon, Hyun Jeong Ko, Young Ho Byun, Baik Lin Seong, Satoshi Uematsu, Shizuo Akira, and Mi Na Kweon. 2011. Type I interferon signaling regulates Ly6Chi monocytes and neutrophils during acute viral pneumonia in mice. PLoS Pathogens 7. https://doi.org/10.1371/journal.ppat.1001304
Lauder, Sarah N., Emma Jones, Kathryn Smart, Anja Bloom, Anwen S. Williams, James P. Hindley, Beatrice Ondondo, et al. 2013. Interleukin-6 limits influenza-induced inflammation and protects against fatal lung pathology. European Journal of Immunology 43: 2613–2625. https://doi.org/10.1002/eji.201243018.
Stifter, Sebastian A., Nayan Bhattacharyya, Roman Pillay, Manuela Flórido, James A. Triccas, Warwick J. Britton, and Carl G. Feng. 2016. Functional interplay between type I and II interferons is essential to limit influenza A virus-induced tissue inflammation. PLoS Pathogens 12: 1–20. https://doi.org/10.1371/journal.ppat.1005378.
Kaplanski, Gilles, Valérie Marin, Félix. Montero-Julian, Alberto Mantovani, and Catherine Farnarier. 2003. IL-6: A regulator of the transition from neutrophil to monocyte recruitment during inflammation. Trends in Immunology 24: 25–29. https://doi.org/10.1016/S1471-4906(02)00013-3.
Rodriguez, Angeline E., Christopher Bogart, Christopher M. Gilbert, Jonathan A. McCullers, Amber M. Smith, Thirumala Devi Kanneganti, and Christopher R. Lupfer. 2019. Enhanced IL-1β production is mediated by a TLR2-MYD88-NLRP3 signaling axis during coinfection with influenza A virus and Streptococcus pneumoniae. PLoS ONE 14: 10–13. https://doi.org/10.1371/journal.pone.0212236.
Ishikawa, Hiroki, Toshie Fukui, Satoshi Ino, Hiraku Sasaki, Naoki Awano, Chikara Kohda, and Kazuo Tanaka. 2016. Influenza virus infection causes neutrophil dysfunction through reduced G-CSF production and an increased risk of secondary bacteria infection in the lung. Virology 499. Elsevier: 23–29. https://doi.org/10.1016/j.virol.2016.08.025.
Fielding, Ceri A., Rachel M. McLoughlin, Louise McLeod, Chantal S. Colmont, Meri Najdovska, Dianne Grail, Matthias Ernst, Simon A. Jones, Nicholas Topley, and Brendan J. Jenkins. 2008. IL-6 regulates neutrophil trafficking during acute inflammation via STAT3. The Journal of Immunology 181: 2189–2195. https://doi.org/10.4049/jimmunol.181.3.2189.
Tate, Michelle D., Andrew G. Brooks, and Patrick C. Reading. 2008. The role of neutrophils in the upper and lower respiratory tract during influenza virus infection of mice. Respiratory Research 9: 1–13. https://doi.org/10.1186/1465-9921-9-57.
Jewell, Nancy A., Troy Cline, Sara E. Mertz, Sergey V. Smirnov, Emilio Flaño, Christian Schindler, Jessica L. Grieves, Russell K. Durbin, Sergei V. Kotenko, and Joan E. Durbin. 2010. Lambda interferon is the predominant interferon induced by influenza A virus infection in vivo. Journal of Virology 84: 11515–11522. https://doi.org/10.1128/jvi.01703-09.
Yamamoto, K, T Miyoshi-Koshio, Y Utsuki, S Mizuno, and K Suzuki. 1991. Virucidal activity and viral protein modification by myeloperoxidase: A candidate for defense factor of human polymorphonuclear leukocytes against influenza virus infection. The Journal of infectious diseases 164. United States: 8–14. https://doi.org/10.1093/infdis/164.1.8.
Colletti, L.M., D.G. Remick, G.D. Burtch, S.L. Kunkel, R.M. Strieter, and D.A. Campbell. 1990. Role of tumor necrosis factor-α in the pathophysiologic alterations after hepatic ischemia/reperfusion injury in the rat. Journal of Clinical Investigation 85: 1936–1943. https://doi.org/10.1172/JCI114656.
Perrone, Lucy A., Jessica A. Belser, Debra A. Wadford, Jacqueline M. Katz, and Terrence M. Tumpey. 2013. Inducible nitric oxide contributes to viral pathogenesis following highly pathogenic influenza virus infection in mice. Journal of Infectious Diseases 207: 1576–1584. https://doi.org/10.1093/infdis/jit062.
Ishikawa, Eri, Masatoshi Nakazawa, Masahiro Yoshinari, and Mutsuhiko Minami. 2005. Role of tumor necrosis factor-related apoptosis-inducing ligand in immune response to influenza virus infection in mice. Journal of virology 79: 7658–7663. https://doi.org/10.1128/JVI.79.12.7658-7663.2005.
Mian, I., and Asad. 2012. Nitric oxide metabolites as biomarkers for influenza-like acute respiratory infections presenting to the emergency room. The Open Respiratory Medicine Journal 6: 127–134. https://doi.org/10.2174/1874306401206010127.
Nin, N, C Sánchez-Rodríguez, L S Ver, P Cardinal, A Ferruelo, L Soto, A Deicas, et al. 2012. Lung histopathological findings in fatal pandemic influenza A (H1N1). Medicina Intensiva 36: 24–31. https://doi.org/10.1016/j.medin.2011.10.005.
Coates, Bria M., Kelly L. Staricha, Clarissa M. Koch, Yuan Cheng, Dale K. Shumaker, G.R. Scott Budinger, Harris Perlman, Alexander V. Misharin, and Karen M. Ridge. 2018. Inflammatory monocytes drive influenza A virus–Mediated lung injury in juvenile mice. The Journal of Immunology 200: 2391–2404. https://doi.org/10.4049/jimmunol.1701543.
Burggraaf, Simon, John Bingham, Jean Payne, Wayne G. Kimpton, John W. Lowenthal, and Andrew G.D. Bean. 2011. Increased inducible nitric oxide synthase expression in organs is associated with a higher severity of H5N1 influenza virus infection. PLoS ONE 6: 1–12. https://doi.org/10.1371/journal.pone.0014561.
Mielke, Lisa A, Karen L Elkins, Lai Wei, Robyn Starr, Philip N Tsichlis, John J O'Shea, and Wendy T Watford. 2009. Tumor progression locus 2 (Map3k8) is critical for host defense against Listeria monocytogenes and IL-1β production. The Journal of Immunology 183: 7984 LP – 7993. https://doi.org/10.4049/jimmunol.0901336.
Redford, Paul S., Katrin D. Mayer-Barber, Finlay W. McNab, Evangelos Stavropoulos, Andreas Wack, Alan Sher, and Anne O’Garra. 2014. Influenza A virus impairs control of mycobacterium tuberculosis coinfection through a type i interferon receptor-dependent pathway. Journal of Infectious Diseases 209: 270–274. https://doi.org/10.1093/infdis/jit424.
Galani, Ioanna E., Vasiliki Triantafyllia, Evridiki Evangelia Eleminiadou, Ourania Koltsida, Athanasios Stavropoulos, Maria Manioudaki, Dimitris Thanos, et al. 2017. Interferon-λ mediates non-redundant front-line antiviral protection against influenza virus infection without compromising host fitness. Immunity 46. Elsevier Inc.: 875–890.e6. https://doi.org/10.1016/j.immuni.2017.04.025.
Sato, Mitsuharu, Hirofumi Suemori, Naoki Hata, Masataka Asagiri, Kouetsu Ogasawara, Kazuki Nakao, Takeo Nakaya, et al. 2000. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-α/β gene induction. Immunity 13: 539–548. https://doi.org/10.1016/S1074-7613(00)00053-4.
Taniguchi, Tadatsugu, and Akinori Takaoka. 2002. The interferon-α/β system in antiviral responses: A multimodal machinery of gene regulation by the IRF family of transcription factors. Current Opinion in Immunology 14: 111–116. https://doi.org/10.1016/S0952-7915(01)00305-3.
Mordstein, Markus, Eva Neugebauer, Vanessa Ditt, Birthe Jessen, Toni Rieger, Valeria Falcone, Frederic Sorgeloos, et al. 2010. Lambda interferon renders epithelial cells of the respiratory and gastrointestinal tracts resistant to viral infections. Journal of Virology 84: 5670–5677. https://doi.org/10.1128/jvi.00272-10.
Klinkhammer, Jonas, Daniel Schnepf, Liang Ye, Marilena Schwaderlapp, Hans Henrik Gad, Rune Hartmann, Dominique Garcin, Tanel Mahlakõiv, and Peter Staeheli. 2018. IFN-λ prevents influenza virus spread from the upper airways to the lungs and limits virus transmission. eLife 7: 1–18. https://doi.org/10.7554/eLife.33354.
Wang, Yanshi, Tingting Li, Yongyan Chen, Haiming Wei, Rui Sun, and Zhigang Tian. 2017. Involvement of NK Cells in IL-28B–Mediated Immunity against Influenza Virus Infection. The Journal of Immunology 199: 1012–1020. https://doi.org/10.4049/jimmunol.1601430.
Major, Jack, Stefania Crotta, Miriam Llorian, Teresa M. McCabe, Hans H. Gad, Simon L. Priestnall, Rune Harmann, and Andreas Wack. 2020. Type I and III interferons disrupt lung epithelial repair during recovery from viral infection. Science 369: 712–717.
Kumagai Y, Takeuchi O, Kato H, Kumar H, Matsui K, Morii E, Aozasa K, Kawai T, Akira S. 2007. Alveolar macrophages are the primary interferon-alpha producer in pulmonary infection with RNA viruses. Immunity. Aug;27(2):240–252. https://doi.org/10.1016/j.immuni.2007.07.013. PMID: 17723216.
Högner K, Wolff T, Pleschka S, Plog S, Gruber AD, Kalinke U, Walmrath HD, Bodner J, Gattenlöhner S, Lewe-Schlosser P, Matrosovich M, Seeger W, Lohmeyer J, Herold S. Macrophage-expressed IFN-β contributes to apoptotic alveolar epithelial cell injury in severe influenza virus pneumonia. PLoS Pathog. 2013 Feb;9(2):e1003188. https://doi.org/10.1371/journal.ppat.1003188. Epub 2013 Feb 28. Erratum in: PLoS Pathog. 2016 Jun;12(6):e1005716. PMID: 23468627; PMCID: PMC3585175.
Ghoneim, Hazem E., Paul G. Thomas, and Jonathan A. McCullers. 2013. Depletion of alveolar macrophages during influenza infection facilitates bacterial superinfections. The Journal of Immunology 191: 1250–1259. https://doi.org/10.4049/jimmunol.1300014.
Cardani, Amber, Adam Boulton, Taeg S. Kim, and Thomas J. Braciale. 2017. Alveolar macrophages prevent lethal influenza pneumonia by inhibiting infection of type-1 alveolar epithelial cells. PLoS Pathogens 13: 1–25. https://doi.org/10.1371/journal.ppat.1006140.
Acknowledgements
We are thankful to Denise Fahey and Kimberly Oliva for technical support in the study. We would also like to thank University Research Animal Resources at the UGA Coverdell Rodent Vivarium for excellent care of animals. Additionally, we are grateful to Jamie Barber and Julie Nelson for their support and maintenance of the instruments at the College of Veterinary Medicine Cytometry Core Facility and CTEGD Cytometry Shared Resource Laboratory. We also thank Donald Harn and Biao He for use of laboratory equipment.
Funding
Research reported in this publication was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number R21AI147003-01 to W.T.W. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
KL and WW conceptualized and designed the experiments. KL, YP, and SR performed the experiments and analyzed the data. KL and WW wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
All experiments involving mice were performed according to the University of Georgia Guidelines for Laboratory Animals and were approved by the UGA Institutional Animal Care and Use Committee (#A2021 03–026-A3). No human subjects were used in the study.
Consent for Publication
Not applicable. No human subjects were used in the study.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Latha, K., Patel, Y., Rao, S. et al. The Influenza-Induced Pulmonary Inflammatory Exudate in Susceptible Tpl2-Deficient Mice Is Dictated by Type I IFN Signaling. Inflammation 46, 322–341 (2023). https://doi.org/10.1007/s10753-022-01736-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10753-022-01736-8