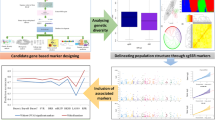
Abstract
The study objective was to evaluate the application of factor analysis (FA) on genomic prediction considering different density marker panels. The FA transforms phenotype traits in latent variables (factor scores), called pseudo-phenotype in this study. The Genomic Best Linear Unbiased Prediction method was applied to the Oriza sativa L phenotype traits. The dataset contains twenty-two phenotypic traits and 36,901 SNPs (Single Nucleotide Polymorphism) from 413 genotypes. The results obtained indicate that combining the factor analysis and the genomic prediction with different density marker panels was efficient. The analysis presented similar values for predictive ability, considering the phenotypes and pseudo-phenotypes (in both analyses, there was variation between 0.60 and 0.80), high agreement of SNPs with major effects, and high agreement between the best and worst selected individuals considering phenotypes and pseudo-phenotypes analysis.






Similar content being viewed by others
Availability of data and material
Not applicable
Code availability
Not applicable
References
Akinwale MG, Gregorio G, Nwilene F, Akinyele BO, Ogunbayo SA, Odiyi AC (2011) Heritability and correlation coefficient analysis for yield and its components in rice (Oryza sativa L.). Afr J Plant Sci 5:207–212
Alvarenga AB, Veroneze R, Oliveira HR, Marques DBD, Lopes PS, Silva FF, Brito LF (2020) Comparing alternative single-step GBLUP approaches and training population designs for genomic evaluation of crossbred animals. Front Genet 11:1–19. https://doi.org/10.3389/fgene.2020.00263
Ammiraju JS, Luo M, Goicoechea JL, Wang W, Kudrna D, Mueller C, Talag J, Kim H, Sisneros NB, Blackmon B, Fang E, Tomkins JB, Brar D, MacKill D, McCouch S, Kurata N, Lambert G, Galbraith DW, Arumuganathan K, Rao K, Walling JG, Gill N, Yu Y, SanMiguel P, Soderlund C, Jackson S, Wing RA (2006) The Oryza bacterial artificial chromosome library resource: construction and analysis of 12 deep-coverage large-insert BAC libraries that represent the 10 genome types of the genus Oryza. Genome Res 16(1):140–147. https://doi.org/10.1101/gr.3766306
Azevedo CF, de Resende MDV, Silva FF, Viana JMS, Valente MSF, Resende MFR Jr, Munoz P (2015) Ridge, Lasso and Bayesian additive-dominance genomic models. BMC Genet 16:105. https://doi.org/10.1186/s12863-015-0264-2
Azevedo CF, Nascimento M, Fontes VC, Silva FF, Resende MDV, Cruz CD (2019) GenomicLand: software for genome-wide association studies and genomic prediction. Acta Sci Agro 41:e45361. https://doi.org/10.4025/actasciagron.v41i1.45361
Baertschi C, Cao TV, Bartholomé J, Ospina Y, Quintero C, Frouin J, Bouvet JM, Grenier C (2021) Impact of early genomic prediction for recurrent selection in an upland rice synthetic population. G3 Genes Gen Genet 11(12):jkab320. https://doi.org/10.1093/g3journal/jkab320
Batista LG, Gaynor RC, Margarido GRA, Byrne T, Amer P, Gorjanc G, Hickey JM (2021) Long-term comparison between index selection and optimal independent culling in plant breeding programs with genomic prediction. PLoS ONE 16(5):e0235554. https://doi.org/10.1371/journal.pone.0235554
Cerny BA, Kaiser HF (1977) A study of a measure of sampling adequacy for factor-analytic correlation matrices. Multivar Behav Res 12(1):43–47. https://doi.org/10.1207/s15327906mbr1201_3
Ceron-Rojas JJ, Crossa J, Arief VN, Basford K, Rutkoski J, Jarquín D, Alvarado G, Beyene Y, Semagn K, DeLacy I (2015) A genomic selection index applied to simulated and real data. Genes Gen Genet 5(10):2155–2164. https://doi.org/10.1534/g3.115.019869
Costa JA, Azevedo CF, Nascimento M, Silva FF, Resende MDV, Nascimento ACC (2020) Genomic prediction with the additive-dominant model by dimensionality reduction methods. Pesqui Agropecu Brasil 55:e01713. https://doi.org/10.1590/s1678-3921.pab2020.v55.01713
Cui Y, Li R, Li G, Zhang F, Zhu T, Zhang Q, Ali J, Li Z, Xu S (2020) Hybrid breeding of rice via genomic selection. Plant Biotechnol J 18(1):57–67. https://doi.org/10.1111/pbi.13170
Dekkers JCM (2021) Multiple trait breeding programs with genotype-by-environment interactions based on reaction norms, with application to genetic improvement of disease resilience. Genet Sel Evol 53:93. https://doi.org/10.1186/s12711-021-00687-2
FAO (2022) Food and Agriculture Organization of the United Nations.
Ferreira DF (2011) Estatística Multivariada. UFLA, Lavras
Frouin J, Labeyrie A, Boisnard A, Sacchi GA, Ahmadi N (2019) Genomic prediction offers the most effective marker assisted breeding approach for ability to prevent arsenic accumulation in rice grains. PLoS ONE 14(6):e0217516. https://doi.org/10.1371/journal.pone.0217516
Grenier C, Cao TV, Ospina Y, Quintero C, Châtel MH, Tohme J, Courtois B, Ahmadi N (2016) Accuracy of genomic selection in a rice synthetic population developed for recurrent selection breeding. PLoS ONE 11(5):e0154976. https://doi.org/10.1371/journal.pone.0136594
Guo Z, Tucker DM, Basten CJ, Gandhi H, Ersoz E, Guo B, Xu Z, Wang D, Gay G (2014) The impact of population structure on genomic prediction in stratified populations. Theor Appl Genet 127:749–762. https://doi.org/10.1007/s00122-013-2255-x
Isidro J, Jannink JL, Akdemir D, Poland J, Heslot N, Sorrells ME (2015) Training set optimization under population structure in genomic selection. Theor Appl Genet 128:145–158. https://doi.org/10.1007/s00122-014-2418-4
James G, Witten D, Hastie T, Tibshirani R (2013) An introduction to statistical learning: with applications in R. Springer, New York. https://doi.org/10.1007/978-1-4614-7138-7
Karaman E, Lund MS, Su G (2020) Multi-trait single-step genomic prediction accounting for heterogeneous (co)variances over the genome. Heredity 124:274–287. https://doi.org/10.1038/s41437-019-0273-4
Kehel Z, Sanchez-Garcia MA, Baouchi AE, Aberkane H, Tsivelikas A, Charles C, Amri A (2020) Predictive characterization for seed morphometric traits for genebank accessions using genomic selection. Front Ecol Evol. https://doi.org/10.3389/fevo.2020.00032
Kriaridou C, Tsairidou S, Houston RD, Robledo D (2020) Genomic prediction using low density marker panels in aquaculture: performance across species, traits, and genotyping platforms. Front Genet 27(11):124. https://doi.org/10.3389/fgene.2020.00124
Labroo MR, Ali J, Aslam MU, de Asis EJ, dela Paz MA, Sevilla MA, Lipka AE, Studer AJ, Rutkoski JE (2021) genomic prediction of yield traits in single-cross hybrid rice (Oryza sativa L). Front Genet 12:692870. https://doi.org/10.3389/fgene.2021.692870
Lee SH, Clark S, van der Werf JHJ (2017) Estimation of genomic prediction accuracy from reference populations with varying degrees of relationship. PLoS ONE 12(12):e0189775. https://doi.org/10.1371/journal.pone.0189775
Liu Z, Sun C, Yan Y, Li G, Li XC, Wu G, Yang N (2021) Design and evaluation of a custom 50K Infinium SNP array for egg-type chickens. Poult Sci 100(5):101044. https://doi.org/10.1016/j.psj.2021.101044
Lopez-Cruz M, de Los Campos G (2021) Optimal breeding-value prediction using a sparse selection index. Genetics 218(1):030. https://doi.org/10.1093/genetics/iyab030
Moeinizade S, Kusmec A, Hu G, Wang L, Schnable PS (2020) Multi-trait genomic selection methods for crop improvement. Genetics 215(4):931–945. https://doi.org/10.1534/genetics.120.303305
Montesinos-López A, Runcie DE, Ibba MI, Pérez-Rodríguez P, Montesinos-López OA, Crespo LA, Bentley AR, Crossa J (2021) Multi-trait genomic-enabled prediction enhances accuracy in multi-year wheat breeding trials. Genes Gen Genet 11(10):jkab270. https://doi.org/10.1093/g3journal/jkab270
NASS (2022) National Agricultural Statistics Service.
Onogi A, Ideta O, Inoshita Y, Ebana K, Yoshioka T, Yamasaki M, Iwata H (2015) Exploring the areas of applicability of whole-genome prediction methods for asian rice (oryza sativa L.). Theor Appl Genet 128:41–53. https://doi.org/10.1007/s00122-014-2411-y
Paixão PTM, Nascimento ACC, Nascimento M, Azevedo CF, Oliveira GF, Silva FL, Caixeta ET (2022) Factor analysis applied in genomic selection studies in the breeding of Coffea canephora. Euphytica 218:42. https://doi.org/10.1007/s10681-022-02998-x
Patterson HD, Thompson R (1971) Recovery of inter-block information when block sizes are unequal. Biometrika 58(3):545–554. https://doi.org/10.2307/2334389
Rencher AC (2002) Methods of multivariate analysis. John Wiley, New York
Resende MFR Jr, Muñoz P, Resende MDV, Garrick DJ, Fernando RL, Davis JM, Jokela EJ, Martin TA, Peter GF, Kirst (2012a) M accuracy of genomic selection methods in a standard data set of loblolly pine (Pinus taeda L.). Genetics 190(4):1503–1510. https://doi.org/10.1534/genetics.111.137026
Resende MDV, Silva FF, Lopes PS, Azevedo CF (2012b) Seleção Genômica Ampla (GWS) via Modelos Mistos (REML/BLUP), Inferência Bayesiana (MCMC), Regressão Aleatória Multivariada (RRM) e Estatística Espacial. Universidade Federal de Viçosa/Departamento de Estatística. Viçosa, Viçosa
Revelle W (2021) psych: procedures for personality and psychological research. Northwestern University, Evanston, Illinois, USA
Spindel J, Begum H, Akdemir D, Virk P, Collard B, Redoña E, Atlin G, Jannink JL, McCouch SR (2015) Genomic selection and association mapping in rice (Oryza sativa): effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines. PLoS Genet 11:1–25. https://doi.org/10.1371/journal.pgen
Spindel JE, Begum H, Akdemir D, Collard B, Redoña E, Jannink JL, McCouch S (2016) Genome-wide prediction models that incorporate de novo GWAS are a powerful new tool for tropical rice improvement. Heredity 116(4):395–408. https://doi.org/10.1038/hdy.2015.113
Sweeney DW, Rooney TE, Sorrells ME (2021) Gain from genomic selection for a selection index in two-row spring barley. Plant Gen 14:e20138. https://doi.org/10.1002/tpg2.20138
Teixeira FRF, Nascimento M, Nascimento ACC, Paixão DM, Azevedo CF, Silva FF, Cruz CC, Lopes PS, Guimarães SEF (2015) Determinação de fatores em características de suínos. Rev Brasil De Biom 33:130–138
Teixeira FRF, Nascimento M, Nascimento ACC, Silva FF, Cruz CC, Azevedo CF, Paixão DM, Barroso LMA, Verardo LL, Resende MDV, Guimarães SEF, Lopes OS (2016) Factor analysis applied to genome prediction for high-dimensional phenotypes in pigs. Genet Mol Res 15:1–10. https://doi.org/10.4238/gmr.15028231
Torres LG, Rodrigues MC, Lima NL, Trindade TFH, Silva FF, Azevedo CF, DeLima RO (2018) Multi-trait multi-environment Bayesian model reveals G x E interaction for nitrogen use efficiency components in tropical maize. PLoS ONE 13(6):e0199492. https://doi.org/10.1371/journal.pone.0199492
Vitezica ZG, Varona L, Legarra A (2013) On the additive and dominant variance and covariance of individuals within the genomic selection scope. Genetics 195(4):1223–1230. https://doi.org/10.1534/genetics.113.155176
Wang X, Li L, Yang Z, Zheng X, Yu S, Xu C, Hu Z (2017) Predicting rice hybrid performance using univariate and multivariate GBLUP models based on North Carolina mating design II. Heredity 118(3):302–310. https://doi.org/10.1038/hdy.2016.87
Wang J, Zhou Z, Zhang Z, Li H, Liu D, Zhang Q, Bradbury PJ, Buckler ES, Zhang Z (2018) Expanding the BLUP alphabet for genomic prediction adaptable to the genetic architectures of complex traits. Heredity 121:648–662. https://doi.org/10.1038/s41437-018-0075-0
Xu Y, Wang X, Ding X, Zheng X, Yang Z, Xu C, Hu Z (2018) Genomic selection of agronomic traits in hybrid rice using an NCII population. Rice 11(1):32. https://doi.org/10.1186/s12284-018-0223-4
Xu Y, Ma K, Zhao Y, Wang X, Zhou K, Yu G, Li C, Li P, Yang Z, Xu C, Xu S (2021) Genomic selection: a breakthrough technology in rice breeding. Crop J 9(3):669–677. https://doi.org/10.1016/j.cj.2021.03.008
Zhang Z, Liu J, Ding X, Bijma P, de Koning D-J, Zhang Q (2010) Best linear unbiased prediction of genomic breeding values using a trait-specific marker-derived relationship matrix. PLoS ONE 5(9):e12648. https://doi.org/10.1371/journal.pone.0012648
Zhang Z, Ding X, Liu J, Zhang Q, Koning DJ (2011) Accuracy of genomic prediction using low-density marker panels. J Dairy Sci 94(7):3642–3650. https://doi.org/10.3168/jds.2010-3917
Zhao K, Tung CW, Eizenga GC, Wright MH, Ali ML, Price AH, Norton GJ, Islam MR, Reynolds A, Mezey J, McClung AM, Bustamante CD, McCouch SR (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:1–10. https://doi.org/10.1371/journal.pgen.1002221
Acknowledgements
The authors are grateful for the fnancial support of the Fundação de Amparo à Pesquisa do Estado de Minas Gerais—FAPEMIG, the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—CAPES and the Conselho Nacional de Desenvolvimento Científico e Tecnológico—CNPq Funding
Funding
The Fundação de Amparo à Pesquisa do Estado de Minas Gerais–FAPEMIG, the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–CAPES and the Conselho Nacional de Desenvolvimento Científico e Tecnológico–CNPq.
Author information
Authors and Affiliations
Contributions
Not applicable.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
The authors have no financial or proprietary interests in any material discussed in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fialho, I.C., Azevedo, C.F., Nascimento, A.C.C. et al. Factor analysis applied in genomic prediction considering different density marker panels in rice. Euphytica 219, 88 (2023). https://doi.org/10.1007/s10681-023-03214-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-023-03214-0