Abstract
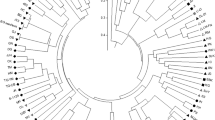
The species Cynara cardunculus includes the globe artichoke (var. scolymus), the cultivated cardoon (var. altilis) and the wild cardoon (var. sylvestris). The three taxa are sexually compatible and originate fertile F 1 progenies, which, given the high heterozygosity of the species, are highly segregating. We report the characterization of two F 1 populations, one bred from a cross between globe artichoke and cultivated cardoon, and the other between globe artichoke and wild cardoon. Both populations featured a wide array of phenotypes in relation to several traits, and some of the newly developed genotypes are of interest for the ornamental market. The two populations were genotyped at 50 microsatellite (SSR) loci: in the globe artichoke × wild cardoon and globe artichoke × cultivated cardoon progenies 116 and 97 alleles were respectively detected. SSR pattern scores were used to produce an UPGMA dendrogram and a PCoA plot. A set of nine SSR loci, evenly dispersed across the genome, was shown to be sufficient to unambiguously identify each segregant. The molecular fingerprinting is useful for establishing the true to type correspondence of propagative materials in nurseries and ensures the effective correspondence between the real and the declared identity of a clone.





Similar content being viewed by others
References
Acquadro A, Portis E, Lanteri S (2003) Isolation of microsatellite loci in artichoke (Cynara cardunculus L. var. scolymus). Mol Ecol Notes 3:37–39
Acquadro A, Portis E, Albertini E, Lanteri S (2005a) M-AFLP-based protocol for microsatellite loci isolation in Cynara cardunculus L. (Asteraceae). Mol Ecol Notes 5:272–274
Acquadro A, Portis E, Lee D, Donini P, Lanteri S (2005b) Development and characterization of microsatellite markers in Cynara cardunculus L. Genome 48:217–225
Acquadro A, Lanteri S, Scaglione D, Arens P, Vosman B, Portis E (2009) Genetic mapping and annotation of genomic microsatellites isolated from globe artichoke. Theor Appl Genet 118(8):1573–1587
Acquadro A, Papanice M, Lanteri S, Bottalico G, Portis E, Campanale A, Finetti-Sialer M, Mascia T, Sumerano P, Gallitelli D (2010) Production and fingerprinting of virus-free clones in a reflowering globe artichoke. Plant Cell Tiss Organ Cult 100(3):329–337
Anido F, Firpo I, Garcia S, Cointry E (1998) Estimation of genetic parameters for yield traits in globe artichoke (Cynara scolymus L.). Euphytica 103:61–66
Barba M, Di Lernia G, Babes G, Citrulli F (2004) Produzione e conservazione di germoplasma di carciofo di tipo romanesco esente da virus. Italus Hortus 11:5–10
Basnitzki J, Zohary D (1994) Breeding of seed planted artichoke. Plant Breed Rev 12:253–269
Cocker H (1967) Il carciofo pianta ornamentale. In: Medica M (ed) I International Congress on Artichoke. Minerva Medica, Bari, pp 313–317
Collard B, Jahufer M, Brouwer J, Pang E (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196
Comino C, Lanteri S, Portis E, Acquadro A, Romani A, Hehn A, Larbat R, Bourgaud F (2007) Isolation and functional characterization of a cDNA coding a hydroxycinnamoyltransferase involved in phenylpropanoid biosynthesis in Cynara cardunculus L. BMC Plant Biol 7:14
Comino C, Hehn A, Moglia A, Menin B, Bourgaud F, Lanteri S, Portis E (2009) The isolation and mapping of a novel hydroxycinnamoyltransferase in the globe artichoke chlorogenic acid pathway. BMC Plant Biol 9:30
Cravero V, Picardi L, Cointry E (2005) An approach for understanding the heredity of two quality traits (head color and tightness) in globe artichoke (Cynara scolymus L.). Genet Mol Biol 28:431–434
Foti S, Mauromicale G, Raccuia S, Fallico B, Fanella F, Maccarone E (1999) Possible alternative utilization of Cynara spp. I. Biomass, grain yield and chemical composition of grain. Ind Crop Prod 10:219–228
Foury C (1969) Étude de la biologie florale de l’artichaut (Cynara scolymus L.). Application a la sélection 2 partie. Étude des descendances obtenues en fécondation contrôlée. Ann Amélior Plantes 19:23–52
Foury C, Aubert S (1977) Observation préliminares sur la présence et la répartition de pigments anthocyaniques dans un mutant d’artichaut (Cynara scolymus L.) à fleurs blanches. Ann Amélior Plantes 27(5):603–612
Holm L, Loeschcke V, Bendixen C (2001) Elucidation of the molecular basis of a null allele in a rainbow trout microsatellite. Mar Biotechnol 3:555–560
Ierna A, Mauromicale G (2010) Cynara cardunculus L. genotypes as a crop for energy purposes in a Mediterranean environment. Biomass Bioenerg 34(5):754–760
Jackson JA, Matthews D (2000) Modified inter-simple sequence repeat PCR protocol for use in conjunction with the LI-COR gene ImagIR(2) DNA analyzer. Biotechniques 28:914–916
Jones A, Stockwell C, Walker D, Avise J (1998) The molecular basis of a microsatellite null allele from the white sands pupfish. J Hered 89:339–342
Lanteri S, Portis E (2008) Globe Artichoke and Cardoon. In: Springer (ed) Vegetables I, vol 1. Springer, New York, pp 49–74
Lanteri S, Di Leo I, Ledda L, Mameli M, Portis E (2001) RAPD variation within and among populations of globe artichoke cultivar ‘Spinoso sardo’. Plant Breeding 120:243–246
Lanteri S, Acquadro A, Saba E, Portis E (2004a) Molecular fingerprinting and evaluation of genetic distances among selected clones of globe artichoke (Cynara cardunculus L. var. scolymus L.). J Hortic Sci Biotech 79:863–870
Lanteri S, Saba E, Cadinu M, Mallica G, Baghino L, Portis E (2004b) Amplified fragment length polymorphism for genetic diversity assessment in globe artichoke. Theor Appl Genet 108:1534–1544
Lanteri S, Acquadro A, Comino C, Mauro R, Mauromicale G, Portis E (2006) A first linkage map of globe artichoke (Cynara cardunculus var. scolymus L.) based on AFLP, S-SAP, M-AFLP and microsatellite markers. Theor Appl Genet 112:1532–1542
Lattanzio V, Kroon PA, Linsalata V, Cardinali A (2009) Globe artichoke: a functional food and source of nutraceutical ingredients. J Funct Food 1:131–144
Lombardo S, Pandino G, Mauromicale G, Knödler M, Carle R, Schieber M (2010) Influence of genotype, harvest time and plant part on polyphenolic composition of globe artichoke [Cynara cardunculus var. scolymus (L.) Fiori]. Food Chem 119:1175–1181
Maccarone E, Fallico B, Fanella F, Mauromicale G, Raccuia S, Foti S (1999) Possible alternative utilization of Cynara spp. II. Chemical characterization of their grain oil. Ind Crop Prod 10(1999):229–237
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Mauro R, Portis E, Acquadro A, Lombardo S, Mauromicale G, Lanteri S (2009) Genetic diversity of globe artichoke landraces from Sicilian small-holdings: implications for evolution and domestication of the species. Conserv Genet 10:431–440
Mauro R, Lombardo S, Longo AMG, Pandino G, Mauromicale G (2011) New cropping designs of globe artichoke for industrial use. Ital J Agron 6:e8
Mauromicale G, Ierna A (2000) Panorama varietale e miglioramento genetico del carciofo. Informatore agrario 26:39–45
Menin B, Comino C, Moglia A, Dolzhenko Y, Portis E, Lanteri S (2010) Identification and mapping of genes related to caffeoylquinic acid synthesis in Cynara cardunculus L. Plant Sci 178:338–347
Pandino G, Courts F, Lombardo S, Mauromicale G, Williamson G (2010) Caffeoylquinic acids and flavonoids in the immature inflorescence of globe artichoke, wild cardoon, and cultivated cardoon. J Agric Food Chem 58:1026–1031
Papanice MA, Campanale A, Bottalico G, Sumerano P, Gallitelli D (2004) Production of virus-free artichoke germplasm cv Brindisino (Cynara scolymus L.; Apulia). Italus Hortus 11(5):11–15
Peakall R, Smouse P (2006) GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pekkinen M, Varvio S, Kulju K, Karkkainen H, Smolander S, Vihera-Aarnio A, Koski V, Sillanpaa M (2005) Linkage map of birch, Betula pendula Roth, based on microsatellites and amplified fragment length polymorphisms. Genome 48:619–625
Pemberton J, Slate J, Bancroft D, Barrett J (1995) Non-amplifying alleles at microsatellite loci—a caution for parentage and population studies. Mol Ecol 4:249–252
Pochard E, Foury C, Chambonet D (1969) Il miglioramento genetico del carciofo. Proceedings the 1 Congresso Internazionale sul carciofo–Bari–Italy, pp 117-155
Porceddu E, Dellacecca V, Bianco V (1976) Classificazione numerica di cultivar di carciofo. Proceedings II International Congress on Artichoke, Ed Minerva Medica, Torino pp 1105-1119
Portis E, Barchi L, Acquadro A, Macua J, Lanteri S (2005a) Genetic diversity assessment in cultivated cardoon by AFLP (Amplified Fragment Length Polymorphism) and microsatellite markers. Plant Breed 124:299–304
Portis E, Acquadro A, Comino C, Mauromicale G, Saba E, Lanteri S (2005b) Genetic structure of island populations of wild cardoon [Cynara cardunculus L. var. sylvestris (Lamk) Fiori] detected by AFLPs and SSRs. Plant Sci 169:199–210
Portis E, Mauromicale G, Barchi L, Mauro R, Lanteri S (2005c) Population structure and genetic variation in autochthonous globe artichoke germplasm from Sicily Island. Plant Sci 168:1591–1598
Portis E, Mauromicale G, Mauro R, Acquadro A, Scaglione D, Lanteri S (2009) Construction of a reference molecular linkage map of globe artichoke (Cynara cardunculus var. scolymus). Theor Appl Genet 120(1):59–70
Rodzen J, May B (2002) Inheritance of microsatellite loci in the white sturgeon (Acipenser transmontanus). Genome 45:1064–1076
Rohlf FJ (1998) NTSYSpc Version 2.0: User Guide. Applied Biostatistics Inc
Shaw P, Turan C, Wright J, O’Connell M, Carvalho G (1999) Microsatellite DNA analysis of population structure in Atlantic herring (Clupea harengus), with direct comparison to allozyme and mtDNA RFLP analyses. Heredity 83:490–499
Smouse P, Peakall R (1999) Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82:561–573
Sneath PHA, Sokal RR (1973) Numerical taxonomy—the principles and practice of numerical classification. W, H. Freeman, San Francisco
Van Oosterhout C, Hutchinson W, Wills D, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes, vol 4. Wiley, New York, pp 535–538
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lanteri, S., Portis, E., Acquadro, A. et al. Morphology and SSR fingerprinting of newly developed Cynara cardunculus genotypes exploitable as ornamentals. Euphytica 184, 311–321 (2012). https://doi.org/10.1007/s10681-011-0509-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-011-0509-8