Abstract
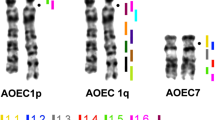
The butterfly lizard (Leiolepis reevesii rubritaeniata) has the diploid chromosome number of 2n = 36, comprising two distinctive components, macrochromosomes and microchromosomes. To clarify the conserved linkage homology between lizard and snake chromosomes and to delineate the process of karyotypic evolution in Squamata, we constructed a cytogenetic map of L. reevesii rubritaeniata with 54 functional genes and compared it with that of the Japanese four-striped rat snake (E. quadrivirgata, 2n = 36). Six pairs of the lizard macrochromosomes were homologous to eight pairs of the snake macrochromosomes. The lizard chromosomes 1, 2, 4, and 6 corresponded to the snake chromosomes 1, 2, 3, and Z, respectively. LRE3p and LRE3q showed the homology with EQU5 and EQU4, respectively, and LRE5p and LRE5q corresponded to EQU7 and EQU6, respectively. These results suggest that the genetic linkages have been highly conserved between the two species and that their karyotypic difference might be caused by the telomere-to-telomere fusion events followed by inactivation of one of two centromeres on the derived dicentric chromosomes in the lineage of L. reevesii rubritaeniata or the centric fission events of the bi-armed macrochromosomes and subsequent centromere repositioning in the lineage of E. quadrivirgata. The homology with L. reevesii rubritaeniata microchromosomes were also identified in the distal regions of EQU1p and 1q, indicating the occurrence of telomere-to-telomere fusions of microchromosomes to the p and q arms of EQU1.





Similar content being viewed by others
Abbreviations
- BrdU:
-
5-Bromodeoxyuridine
- cDNA:
-
Complementary DNA
- FISH:
-
Fluorescence in situ hybridization
- MY:
-
Million years
- MYA:
-
Million years ago
- NF:
-
Fundamental (arm) number
- rRNA:
-
Ribosomal RNA
- RT-PCR:
-
Reverse transcription–polymerase chain reaction
- UV:
-
Ultraviolet
References
Baldini A, Ried T, Shridhar V, Ogura K, D’Aiuto L, Rocchi M, Ward DC (1993) An alphoid DNA sequence conserved in all human and great ape chromosomes: evidence for ancient centromeric sequences at human chromosomal regions 2q21 and 9q13. Hum Genet 90:577–583
Beçak W, Beçak ML (1969) Cytotaxonomy and chromosomal evolution in Serpentes. Cytogenetics 8:247–262
Beçak W, Beçak ML, Nazareth HRS, Ohno S (1964) Close karyological kinship between the reptilian suborder Serpentes and the class Aves. Chromosoma 15:606–617
Bosch HAJ in den, Odierna G, Aprea G et al (2003) Karyological and genetic variation in Middle Eastern lacertid lizards, Lacerta laevis and the Lacerta kulzeri complex: a case of chromosomal allopatric speciation. Chromosome Res 11:165–178
Burt DW (2002) Origin and evolution of avian microchromosomes. Cytogenet Genome Res 96:97–112
Burt DW, Bruley C, Dunn IC et al (1999) The dynamics of chromosome evolution in birds and mammals. Nature 402:411–413
Carbone L, Nergadze SG, Magnani E et al (2006) Evolutionary movement of centromeres in horse, donkey, and zebra. Genomics 87:777–782
Gorman GC (1973) The chromosomes of the Reptilia, a cytotaxonomic interpretation. In: Chiarelli AB, Cappana E (eds) Cytotaxonomy and vertebrate evolution. Academic, New York, pp 349–424
Groenen MAM, Cheng HH, Bumstead N et al (2000) A consensus linkage map of the chicken genome. Genome Res 10:137–147
Ijdo JW, Baldini A, Ward DC, Reeders ST, Wells RA (1991) Origin of human chromosome 2: an ancestral telomere–telomere fusion. Proc Natl Acad Sci USA 88:9051–9055
International Chicken Genome Sequencing Consortium (2004) Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432:695–716
Kawagoshi T, Uno Y, Matsubara K, Matsuda Y, Nishida C (2009) The ZW micro-sex chromosomes of the Chinese soft-shelled turtle (Pelodiscus sinensis, Trionychidae, Testudines) have the same origin as chicken chromosome 15. Cytogenet Genome Res 125:125–131
Kawai A, Nishida-Umehara C, Ishijima J, Tsuda Y, Ota H, Matsuda Y (2007) Different origins of bird and reptile sex chromosomes inferred from comparative mapping of chicken Z-linked genes. Cytogenet Genome Res 117:92–102
Kawai A, Ishijima J, Nishida-Umehara C et al (2009) The ZW sex chromosomes of Gekko hokouensis (Gekkonidae, Squamata) represent highly conserved homology with those of avian species. Chromosoma 118:43–51
Kumar S, Hedges SB (1998) A molecular timescale for vertebrate evolution. Nature 392:917–920
Kumazawa Y (2007) Mitochondrial genomes from major lizard families suggest their phylogenetic relationships and ancient radiations. Gene 388:19–26
Matsubara K, Tarui H, Toriba M et al (2006) Evidence for different origin of sex chromosomes in snakes, birds, and mammals and step-wise differentiation of snake sex chromosomes. Proc Natl Acad Sci USA 103:18190–18195
Matsuda Y, Chapman VM (1995) Application of fluorescence in situ hybridization in genome analysis of the mouse. Electrophoresis 16:261–272
Matsuda Y, Nishida-Umehara C, Tarui H et al (2005) Highly conserved linkage homology between birds and turtles: bird and turtle chromosomes are precise counterparts of each other. Chromosome Res 13:601–615
Montefalcone G, Tempesta S, Rocchi M, Archidiacono N (1999) Centromere repositioning. Genome Res 9:1184–1188
Nanda I, Shan Z, Schartl M et al (1999) 300 million years of conserved synteny between chicken Z and human chromosome 9. Nature Genet 21:258–259
Nanda I, Zend-Ajusch E, Shan Z et al (2000) Conserved synteny between the chicken Z sex chromosome and human chromosome 9 includes the male regulatory gene DMRT1: a comparative (re)view on avian sex determination. Cytogenet Cell Genet 89:67–78
Nishida-Umehara C, Tsuda Y, Ishijima J, Ando J, Fujiwara A, Matsuda Y, Griffin DK (2007) The molecular basis of chromosome orthologies and sex chromosomal differentiation in palaeognathous birds. Chromosome Res 15:721–734
Olmo E (1986) A. Reptilia. In: John B (ed) Animal cytogenetics, 4. Chordata 3. Gebrüder Bornträger, Berlin, pp 1–100
Olmo E, Signorino G (2005) Chromorep: a reptile chromosomes database. Retrieved from http://193.206.118.100/professori/chromorep.pdf
Porter C, Hamilton MJ, Sites Jr JW, Baker RJ (1991) Location of ribosomal DNA in chromosomes of Squamate reptiles: systematic and evolutionary implications. Herpetologica 47:271–280
Schmid M, Feichtinger W, Nanda I et al (1994) An extraordinary low diploid chromosome number in the reptile Gonatodes taniae (Squamata, Gekkonidae). J Hered 85:255–260
Schmid M, Nanda I, Hoehn H et al (2005) Second report on chicken genes and chromosomes 2005. Cytogenet Genome Res 109:415–479
Shibusawa M, Nishibori M, Nishida-Umehara C, Tsudzuki M, Masabanda J, Griffin DK, Matsuda Y (2004) Karyotypic evolution in the Galliformes: an examination of the process of karyotypic evolution by comparison of the molecular cytogenetic findings with the molecular phylogeny. Cytogenet Genome Res 106:111–119
Singh L (1972) Evolution of karyotypes in snakes. Chromosoma 38:185–236
Srikulnath K, Matsubara K, Uno Y et al (2009) Karyological characterization of the butterfly lizard (Leiolepis reevesii rubritaeniata, Agamidae, Squamata) by molecular cytogenetic approach. Cytogenet Genome Res 125:213–223
Townsend TM, Larson A, Louis E, Macey JR (2004) Molecular phylogenetics of Squamata: the position of snakes, amphisbaenians, and dibamids, and the root of the squamate tree. Syst Biol 53:735–757
Vidal N, Hedges SB (2009) The molecular evolutionary tree of lizards, snakes, and amphisbaenians. C R Biol 332:129–139
Acknowledgments
This work was supported by a Grant-in-Aid for Scientific Research (no. 16086201) from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Pat Heslop-Harrison
Rights and permissions
About this article
Cite this article
Srikulnath, K., Nishida, C., Matsubara, K. et al. Karyotypic evolution in squamate reptiles: comparative gene mapping revealed highly conserved linkage homology between the butterfly lizard (Leiolepis reevesii rubritaeniata, Agamidae, Lacertilia) and the Japanese four-striped rat snake (Elaphe quadrivirgata, Colubridae, Serpentes). Chromosome Res 17, 975–986 (2009). https://doi.org/10.1007/s10577-009-9101-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-009-9101-7