Abstract
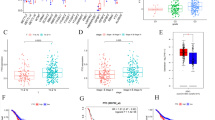
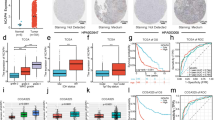
Gliomas are the most common and fatal brain tumors worldwide. Abnormal DNA promoter methylation is an important mechanism for gene loss of tumor suppressors. A long non-coding RNA colorectal adenocarcinoma hypermethylated (CAHM) has been reported to be nearly deleted in glioblastomas (GBMs). Nevertheless, the roles of CAHM in gliomas remain unknown up to now. In the present study, 969 glioma samples downloaded from the CGGA and Gravendeel databases were included. We found that CAHM expression was correlated with glioma grades, molecular subtype, IDH mutation status, and 1q/19p codel status. In glioma cells, CAHM is hypermethylated by DNA methyltransferase1 (DNMT1) and the loss of CAHM expression could be reversed by 5-Aza-2′-deoxycytidine (5-Aza), a specific inhibitor of DNA methyltransferases. Besides, the expression of CAHM was negatively associated with overall survival in both primary and recurrent gliomas. Moreover, the result of Gene Ontology (GO) analysis suggested that CAHM participated in negatively regulating cell development, nervous system development, neurogenesis, and integrin-mediated signaling pathway. Overexpression of CAHM inhibited glioma cell proliferation, clone formation, and invasion. Further exploring results showed that CAHM overexpression suppressed glioma migration and invasion through SPAK/MAPK pathway. Collectively, this study disclosed that CAHM might be a suppressor in gliomas.







Similar content being viewed by others
Data Availability
All data described in this paper are available.
Abbreviations
- CAHM:
-
A long non-coding RNA colorectal adenocarcinoma hypermethylated
- LncRNAs:
-
Long non-coding RNAs
- DNMT1:
-
DNA methyltransferase1
- 5-Aza:
-
5-Aza-2′-deoxycytidine
- LGGs:
-
Lower-grade gliomas
- GBMs:
-
Glioblastomas
- AUC:
-
Area under the curve
- GO:
-
Gene Ontology
References
Bhan A, Soleimani M, Mandal SS (2017) Long noncoding RNA and cancer: a new paradigm. Can Res 77(15):3965–3981. https://doi.org/10.1158/0008-5472.CAN-16-2634
Chew CL, Conos SA, Unal B, Tergaonkar V (2018) Noncoding RNAs: master regulators of inflammatory signaling. Trends Mol Med 24(1):66–84. https://doi.org/10.1016/j.molmed.2017.11.003
Court F, Le Boiteux E, Fogli A, Müller-Barthélémy M, Vaurs-Barrière C, Chautard E, Pereira B, Biau J, Kemeny J, Khalil T, Karayan-Tapon L, Verrelle P, Arnaud P (2019) Transcriptional alterations in glioma result primarily from DNA methylation-independent mechanisms. Genome Res 29(10):1605–1621. https://doi.org/10.1101/gr.249219.119
Ding L, Tian Y, Wang L, Bi M, Teng D, Hong S (2019) Hypermethylated long noncoding RNA MEG3 promotes the progression of gastric cancer. Aging 11(19):8139–8155. https://doi.org/10.18632/aging.102309
Furnari FB, Fenton T, Bachoo RM, Mukasa A, Stommel JM, Stegh A, Hahn WC, Ligon KL, Louis DN, Brennan C, Chin L, DePinho RA, Cavenee WK (2007) Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes Dev 21(21):2683–2710. https://doi.org/10.1101/gad.1596707
Guo Z, Li G, Bian E, Ma CC, Wan J, Zhao B (2017) TGF-β-mediated repression of MST1 by DNMT1 promotes glioma malignancy. Biomed Pharmacother 94:774–780. https://doi.org/10.1016/j.biopha.2017.07.081
Han B, Wang R, Chen Y, Meng X, Wu P, Li Z, Duan C, Li Q, Li Y, Zhao S, Jiang C, Cai J (2019) QKI deficiency maintains glioma stem cell stemness by activating the SHH/GLI1 signaling pathway. Cell Oncol 42(6):801–813. https://doi.org/10.1007/s13402-019-00463-x
Ichimura K, Mungall A, Fiegler H, Pearson D, Dunham I, Carter N, Collins V (2006) Small regions of overlapping deletions on 6q26 in human astrocytic tumours identified using chromosome 6 tile path array-CGH. Oncogene 25(8):1261–1271. https://doi.org/10.1038/sj.onc.1209156
Kerr K, Galler J, Hagen J, Laird P, Laird-Offringa I (2007) The role of DNA methylation in the development and progression of lung adenocarcinoma. Dis Markers 23:5–30. https://doi.org/10.1155/2007/985474
Kim EJ, Kim JS, Lee S, Lee H, Yoon JS, Hong JH, Chun SH, Sun S, Won HS, Hong SA, Kang K, Jo JY, Choi M, Shin DH, Ahn YH, Ko YH (2019) QKI, a miR-200 target gene, suppresses epithelial-to-mesenchymal transition and tumor growth. Int J Cancer 145(6):1585–1595. https://doi.org/10.1002/ijc.32372
Li XX, Wang LJ, Hou J, Liu HY, Wang R, Wang C, Xie W (2020) Identification of long noncoding RNAs as predictors of survival in triple-negative breast cancer based on network analysis. Biomed Res Int 2020:8970340. https://doi.org/10.1155/2020/8970340
Lim M, Xia Y, Bettegowda C, Weller M (2018) Current state of immunotherapy for glioblastoma. Nat Rev Clin Oncol 15(7):422–442. https://doi.org/10.1038/s41571-018-0003-5
Ma JC, Cheng P, Hu Y, Xue YX, Liu YH (2015) Integrin α4 is involved in the regulation of glioma-induced motility of bone marrow mesenchymal stem cells. Oncol Rep 34(2):779–786. https://doi.org/10.3892/or.2015.4012
Mao F, Wang B, Xiao Q, Cheng F, Lei T, Guo D (2017) LRIG proteins in glioma: functional roles, molecular mechanisms, and potential clinical implications. J Neurol Sci 383:56–60. https://doi.org/10.1016/j.jns.2017.10.025
Marchese FP, Raimondi I, Huarte M (2017) The multidimensional mechanisms of long noncoding RNA function. Genome Biol 18(1):206. https://doi.org/10.1186/s13059-017-1348-2
Nir R, Grossman R, Paroush Z, Volk T (2012) Phosphorylation of the drosophila melanogaster RNA-binding protein HOW by MAPK/ERK enhances its dimerization and activity. PLoS Genet 8(3):e1002632. https://doi.org/10.1371/journal.pgen.1002632
Pedersen SK, Mitchell SM, Graham LD, McEvoy A, Thomas ML, Baker RT, Ross JP, Xu ZZ, Ho T, LaPointe LC, Young GP, Molloy PL (2014) CAHM, a long non-coding RNA gene hypermethylated in colorectal neoplasia. Epigenetics 9(8):1071–1082. https://doi.org/10.4161/epi.29046
Prensner JR, Chinnaiyan AM (2011) The emergence of lncRNAs in cancer biology. Cancer Discov 1(5):391–407. https://doi.org/10.1158/2159-8290.CD-11-0209
Romani M, Pistillo MP, Banelli B (2018) Epigenetic targeting of glioblastoma. Front Oncol 8:448. https://doi.org/10.3389/fonc.2018.00448
Shingu T, Ho A, Yuan L, Zhou X, Dai C, Zheng S, Wang Q, Zhong Y, Chang Q, Horner J, Liebelt B, Yao Y, Hu B, Chen Y, Fuller G, Verhaak R, Heimberger A, Hu J (2017) Qki deficiency maintains stemness of glioma stem cells in suboptimal environment by downregulating endolysosomal degradation. Nat Genet 49(1):75–86. https://doi.org/10.1038/ng.3711
Skvortsova K, Stirzaker C, Taberlay P (2019) The DNA methylation landscape in cancer. Essays Biochem 63(6):797–811. https://doi.org/10.1042/ebc20190037
Sun Y, Zhang D, Guo X, Li W, Li C, Luo J, Zhou M, Xue L (2019a) MKK3 modulates JNK-dependent cell migration and invasion. Cell Death Dis 10(3):149. https://doi.org/10.1038/s41419-019-1350-6
Sun Z, Xue H, Wei Y, Wang C, Yu R, Wang C, Wang S, Xu J, Qian M, Meng Q, Li G (2019b) Mucin O-glycosylating enzyme GALNT2 facilitates the malignant character of glioma by activating the EGFR/PI3K/Akt/mTOR axis. Clinical science (London: 1979) 133(10):1167–1184. https://doi.org/10.1042/CS20190145
Wang L, Zhai DS, Ruan BJ, Xu CM, Ye ZC, Lu HY, Jiang YH, Wang ZY, Xiang A, Yang Y, Yuan JL, Lu ZF (2017) Quaking deficiency amplifies inflammation in experimental endotoxemia the aryl hydrocarbon receptor/signal transducer and activator of transcription 1-NF-κB pathway. Front Immunol 8:1754. https://doi.org/10.3389/fimmu.2017.01754
Wen S, Hou Y, Fu L, Xi L, Yang D, Zhao M, Qin Y, Sun K, Teng Y, Liu M (2019) Cancer-associated fibroblast (CAF)-derived IL32 promotes breast cancer cell invasion and metastasis via integrin β3-p38 MAPK signalling. Cancer Lett 442:320–332. https://doi.org/10.1016/j.canlet.2018.10.015
Xiong G, Liu C, Yang G, Feng M, Xu J, Zhao F, You L, Zhou L, Zheng L, Hu Y, Wang X, Zhang T, Zhao Y (2019) Long noncoding RNA GSTM3TV2 upregulates LAT2 and OLR1 by competitively sponging let-7 to promote gemcitabine resistance in pancreatic cancer. J Hematol Oncol 12(1):97. https://doi.org/10.1186/s13045-019-0777-7
Yan J, Jia Y, Chen H, Chen W, Zhou X (2019) Long non-coding RNA PXN-AS1 suppresses pancreatic cancer progression by acting as a competing endogenous RNA of miR-3064 to upregulate PIP4K2B expression. J Exp Clin Cancer Res 38(1):390. https://doi.org/10.1186/s13046-019-1379-5
Ziller MJ, Müller F, Liao J, Zhang Y, Gu H, Bock C, Boyle P, Epstein CB, Bernstein BE, Lengauer T, Gnirke A, Meissner A (2011) Genomic distribution and inter-sample variation of non-CpG methylation across human cell types. PLoS Genet 7(12):e1002389. https://doi.org/10.1371/journal.pgen.1002389
Zou S, Yang J, Guo J, Su Y, He C, Wu J, Yu L, Ding WQ, Zhou J (2018) RAD18 promotes the migration and invasion of esophageal squamous cell cancer via the JNK-MMPs pathway. Cancer Lett 417:65–74. https://doi.org/10.1016/j.canlet.2017.12.034
Funding
This study was funded by the National Natural Science Foundation of China (Grant Nos. 81072066, 81502149).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors confirm that there are no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, Y., Li, Z., Huai, T. et al. DNMT1 Mediated CAHM Repression Promotes Glioma Invasion via SPAK/JNK Pathway. Cell Mol Neurobiol 42, 2643–2653 (2022). https://doi.org/10.1007/s10571-021-01125-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-021-01125-z