-
PDF
- Split View
-
Views
-
Cite
Cite
Chuanzhi Zhang, Junli Zhang, Zhen Kang, Guocheng Du, Xiaobin Yu, Tianwen Wang, Jian Chen, Enhanced production of l-phenylalanine in Corynebacterium glutamicum due to the introduction of Escherichia coli wild-type gene aroH, Journal of Industrial Microbiology and Biotechnology, Volume 40, Issue 6, 1 June 2013, Pages 643–651, https://doi.org/10.1007/s10295-013-1262-x
Close - Share Icon Share
Abstract
Metabolic engineering is a powerful tool which has been widely used for producing valuable products. For improving l-phenylalanine (l-Phe) accumulation in Corynebacterium glutamicum, we have investigated the target genes involved in the biosynthetic pathways. The genes involved in the biosynthesis of l-Phe were found to be strictly regulated genes by feedback inhibition. As a result, overexpression of the native wild-type genes aroF, aroG or pheA resulted in a slight increase of l-Phe. In contrast, overexpression of aroFwt or pheAfbr from E. coli significantly increased l-Phe production. Co-overexpression of aroFwt and pheAfbr improved the titer of l-Phe to 4.46 ± 0.06 g l−1. To further analyze the target enzymes in the aromatic amino acid synthesis pathway between C. glutamicum and E. coli, the wild-type gene aroH from E. coli was overexpressed and evaluated in C. glutamicum. As predicted, upregulation of the wild-type gene aroH resulted in a remarkable increase of l-Phe production. Co-overexpression of the mutated pheAfbr and the wild-type gene aroH resulted in the production of l-Phe up to 4.64 ± 0.09 g l−1. Based on these results we conclude that the wild-type gene aroH from E. coli is an appropriate target gene for pathway engineering in C. glutamicum for the production of aromatic amino acids.
Introduction
l-phenylalanine (l-Phe), one of the essential amino acids for humans and other animals, is widely used in food and pharmaceutical industries [1, 29, 33]. In early industrial processes l-Phe was mainly produced by chemical synthesis. However, because of the very specific demand for the stereo-specific form, consumer’s preference, and various problems with chemical synthesis, the latter is gradually being replaced with bioprocessing, such as microbial fermentation and enzymatic transformation [22]. More recently, metabolic engineering has focused on the Gram-negative strain Escherichia coli and many other well-characterized model microorganisms, and cell factories have been constructed [13, 24, 33]. Although high titers of l-Phe can be produced, the industrial-scale production of l-Phe by genetically engineered E. coli has encountered a lot of problems, especially infection by phage. This has led to the search for alternative robust engineered strains (e.g. Corynebacterium glutamicum).
Since 1957, C. glutamicum, a well-known Gram-positive strain in industrial processes, has been widely used for producing amino acids, especially l-glutamate and l-lysine [9, 26]. The whole genome sequencing of C. glutamicum has led to the successful development of many sophisticated vectors which have been applied in C. glutamicum [6, 18, 23, 30]. In recent years, researchers have taken advantage of the powerful toolbox that has become available (“omics” technologies, metabolic pathway engineering, targeted enzyme modification) and successfully engineered C. glutamicum for producing many high valuable-added products, as described in the excellent review by Becker and Wittmann [2]. Due to the attractive features of genetically engineered C. glutamicum, such as fast growth rate, lack of endotoxin and protease secretion and easy purification step [6], C. glutamicum has also been used for the heterologous expression of recombinant proteins [3, 4, 20]. In addition, C. glutamicum has a distinct ability to simultaneously metabolize pentose and hexose, which makes this microorganism a potential candidate for transforming renewable biomasses to value-added products [9].
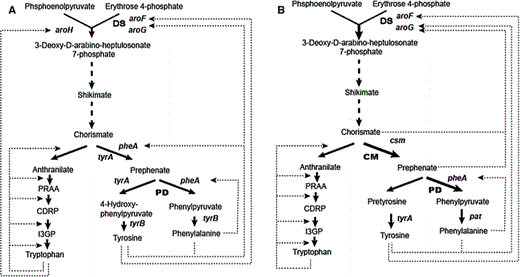
l-phenylalanine (l-Phe) biosynthesis pathway in Escherichia coli (a) and Corynebacterium glutamicum (b). Dotted lines, dashed lines feedback inhibition and repression, respectively. DS 3-deoxy-D-arabinoheptulosonate 7-phosphate synthase, CM chorismate mutase, PD prephenate dehydratase, CDRP 1-(2-carboxyphenylamino)-1-deoxy-D-nbulose-5-phosphate, I3GP indole 3-glycerolphosphate, PRAA n-(5-phospho-β-D-nbosylanthranilate), PRT anthranilate phosphoribosyltransferase
To accumulate l-Phe efficiently in E. coli, the aroG and pheA genes from E. coli were modified by site-mutation to eliminate the specific feedback inhibition and overexpressed to increase the production of l-Phe [1, 8, 31, 33]. In a similar approach, the mutated pheA gene of E. coli was introduced into C. glutamicum KY10694, and a 30 % increase of l-Phe was achieved [17]. In one study, integration of the aroG-pheA tandem genes of E. coli into C. glutamicum resulted in a 1.71-fold increase in l-Phe production, although the highest concentration was also very low (3.97 g l−1) in shake flask cultivation [22]. Shu et al. [24] achieved a high level of l-Phe (23.2 g l−1) by supplying elevated oxygen into 5-l fermentor with the wild-type strain. In a previous study in E. coli in which we analyzed the key enzymes of the shikimate and chorismate pathways in C. glutamicum ATCC 13032, we found that the wild-type gene aroH that is inhibited by l-Trp from E. coli was expressed in C. glutamicum and contributed significantly to increased l-Phe production. By co-overexpression of mutated pheA and the wild-type aroH gene, l-Phe was increased by 13.6-fold. In addition, sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and reverse transcription (RT) PCR data showed that the levels of the mutated pheA and wild-type aroH within one operon were significantly different, suggesting that precise regulation of the target enzymes might be beneficial to the titer of the target end-product, l-Phe.
Materials and methods
Bacterial strains and plasmids
Bacterial strains, plasmids and oligonucleotides used in this study are listed in Tables 1 and 2. E. coli JM109 was applied for molecular cloning and manipulation of plasmids. C. glutamicum ATCC 13032 as the parent strain was engineered for producing l-Phe. Genes aroG, aroF and pheA were amplified from genomic DNA of C. glutamicum ATCC 13032 while the wild-type gene aroH was amplified from genomic DNA of E. coli W3110. After digestion, the fragments were subcloned into pXMJ19 to generate pXMJ19-aroG, pXMJ19-aroF, pXMJ19-pheA and pXMJ19-aroH, respectively. PheAfbr and aroFwt were amplified from plasmid pAP-B03 [33] and subcloned into pXMJ19, resulting in plasmids pXMJ19-pheAfbr and pXMJ19-aroFwt, respectively. To increase the translation efficiency, the Shine-Dalgarno sequence (AGGAGGA) was artificially added in the upstream of starting codon ATG (Table 1).
Strains and plasmids used in this study
| Strains and plasmids . | Relevant properties . | Source or reference . |
|---|---|---|
| Strains | ||
| Escherichia coli JM109 | Wide-type E. coli. | Lab stock |
| E. coli W3110 | In(rrnD-rrnE) E. coli | Lab stock |
| Corynebacterium glutamicum ATCC 13032 | Wide-type C. glutamicum | ATCC |
| C. glutamicum 19G | C. glutamicum ATCC 13032 harboring pXMJ19- aroG | This work |
| C. glutamicum 19F | C. glutamicum ATCC 13032 harboring pXMJ19- aroF | This work |
| C. glutamicum 19Fwt | C. glutamicum ATCC 13032 harboring pXMJ19- aroFwt | This work |
| C. glutamicum 19H | C. glutamicum ATCC 13032 harboring pXMJ19- aroH | This work |
| C. glutamicum 19A | C. glutamicum ATCC 13032 harboring pXMJ19-pheA | This work |
| C. glutamicum 19Afbr | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr | This work |
| C. glutamicum 19AfbrH | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroH | This work |
| C. glutamicum 19AfbrFwt | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroFwt | This work |
| Plasmids | ||
| pMD19-simple Vector | Cloning vector, AmpR | Takara, Dalian, China |
| pXMJ19 | E. coli-C. glutamicum shuttle expression vector | [6] |
| pAP-B03 | Derivative from pACYC177 and pPL450, containing pheAfbr and aroFwt | [33] |
| pXMJ19-aroG | pXMJ19 containing aroG (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroF | pXMJ19 containing aroF gene (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroFwt | pXMJ19 containing aroFwt gene (pAP-B03) | This work |
| pXMJ19-aroH | pXMJ19 containing aroH (E. coli 3110) | This work |
| pXMJ19-pheA | pXMJ19 containing pheA (C. glutamicum ATCC 13032) | This work |
| pXMJ19-pheAfbr | pXMJ19 containing pheAfbr (pAP-B03) | This work |
| pXMJ19-pheAfbr-aroH | pXMJ19 containing pheAfbr (pAP-B03) and aroH (E. coli) | This work |
| pXMJ19-pheAfbr-aroFwt | pXMJ19 containing pheAfbr (pAP-B03) and aroFwt (pAP-B03) | This work |
| Strains and plasmids . | Relevant properties . | Source or reference . |
|---|---|---|
| Strains | ||
| Escherichia coli JM109 | Wide-type E. coli. | Lab stock |
| E. coli W3110 | In(rrnD-rrnE) E. coli | Lab stock |
| Corynebacterium glutamicum ATCC 13032 | Wide-type C. glutamicum | ATCC |
| C. glutamicum 19G | C. glutamicum ATCC 13032 harboring pXMJ19- aroG | This work |
| C. glutamicum 19F | C. glutamicum ATCC 13032 harboring pXMJ19- aroF | This work |
| C. glutamicum 19Fwt | C. glutamicum ATCC 13032 harboring pXMJ19- aroFwt | This work |
| C. glutamicum 19H | C. glutamicum ATCC 13032 harboring pXMJ19- aroH | This work |
| C. glutamicum 19A | C. glutamicum ATCC 13032 harboring pXMJ19-pheA | This work |
| C. glutamicum 19Afbr | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr | This work |
| C. glutamicum 19AfbrH | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroH | This work |
| C. glutamicum 19AfbrFwt | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroFwt | This work |
| Plasmids | ||
| pMD19-simple Vector | Cloning vector, AmpR | Takara, Dalian, China |
| pXMJ19 | E. coli-C. glutamicum shuttle expression vector | [6] |
| pAP-B03 | Derivative from pACYC177 and pPL450, containing pheAfbr and aroFwt | [33] |
| pXMJ19-aroG | pXMJ19 containing aroG (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroF | pXMJ19 containing aroF gene (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroFwt | pXMJ19 containing aroFwt gene (pAP-B03) | This work |
| pXMJ19-aroH | pXMJ19 containing aroH (E. coli 3110) | This work |
| pXMJ19-pheA | pXMJ19 containing pheA (C. glutamicum ATCC 13032) | This work |
| pXMJ19-pheAfbr | pXMJ19 containing pheAfbr (pAP-B03) | This work |
| pXMJ19-pheAfbr-aroH | pXMJ19 containing pheAfbr (pAP-B03) and aroH (E. coli) | This work |
| pXMJ19-pheAfbr-aroFwt | pXMJ19 containing pheAfbr (pAP-B03) and aroFwt (pAP-B03) | This work |
ATCC American type culture collection
Strains and plasmids used in this study
| Strains and plasmids . | Relevant properties . | Source or reference . |
|---|---|---|
| Strains | ||
| Escherichia coli JM109 | Wide-type E. coli. | Lab stock |
| E. coli W3110 | In(rrnD-rrnE) E. coli | Lab stock |
| Corynebacterium glutamicum ATCC 13032 | Wide-type C. glutamicum | ATCC |
| C. glutamicum 19G | C. glutamicum ATCC 13032 harboring pXMJ19- aroG | This work |
| C. glutamicum 19F | C. glutamicum ATCC 13032 harboring pXMJ19- aroF | This work |
| C. glutamicum 19Fwt | C. glutamicum ATCC 13032 harboring pXMJ19- aroFwt | This work |
| C. glutamicum 19H | C. glutamicum ATCC 13032 harboring pXMJ19- aroH | This work |
| C. glutamicum 19A | C. glutamicum ATCC 13032 harboring pXMJ19-pheA | This work |
| C. glutamicum 19Afbr | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr | This work |
| C. glutamicum 19AfbrH | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroH | This work |
| C. glutamicum 19AfbrFwt | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroFwt | This work |
| Plasmids | ||
| pMD19-simple Vector | Cloning vector, AmpR | Takara, Dalian, China |
| pXMJ19 | E. coli-C. glutamicum shuttle expression vector | [6] |
| pAP-B03 | Derivative from pACYC177 and pPL450, containing pheAfbr and aroFwt | [33] |
| pXMJ19-aroG | pXMJ19 containing aroG (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroF | pXMJ19 containing aroF gene (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroFwt | pXMJ19 containing aroFwt gene (pAP-B03) | This work |
| pXMJ19-aroH | pXMJ19 containing aroH (E. coli 3110) | This work |
| pXMJ19-pheA | pXMJ19 containing pheA (C. glutamicum ATCC 13032) | This work |
| pXMJ19-pheAfbr | pXMJ19 containing pheAfbr (pAP-B03) | This work |
| pXMJ19-pheAfbr-aroH | pXMJ19 containing pheAfbr (pAP-B03) and aroH (E. coli) | This work |
| pXMJ19-pheAfbr-aroFwt | pXMJ19 containing pheAfbr (pAP-B03) and aroFwt (pAP-B03) | This work |
| Strains and plasmids . | Relevant properties . | Source or reference . |
|---|---|---|
| Strains | ||
| Escherichia coli JM109 | Wide-type E. coli. | Lab stock |
| E. coli W3110 | In(rrnD-rrnE) E. coli | Lab stock |
| Corynebacterium glutamicum ATCC 13032 | Wide-type C. glutamicum | ATCC |
| C. glutamicum 19G | C. glutamicum ATCC 13032 harboring pXMJ19- aroG | This work |
| C. glutamicum 19F | C. glutamicum ATCC 13032 harboring pXMJ19- aroF | This work |
| C. glutamicum 19Fwt | C. glutamicum ATCC 13032 harboring pXMJ19- aroFwt | This work |
| C. glutamicum 19H | C. glutamicum ATCC 13032 harboring pXMJ19- aroH | This work |
| C. glutamicum 19A | C. glutamicum ATCC 13032 harboring pXMJ19-pheA | This work |
| C. glutamicum 19Afbr | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr | This work |
| C. glutamicum 19AfbrH | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroH | This work |
| C. glutamicum 19AfbrFwt | C. glutamicum ATCC 13032 harboring pXMJ19-pheAfbr-aroFwt | This work |
| Plasmids | ||
| pMD19-simple Vector | Cloning vector, AmpR | Takara, Dalian, China |
| pXMJ19 | E. coli-C. glutamicum shuttle expression vector | [6] |
| pAP-B03 | Derivative from pACYC177 and pPL450, containing pheAfbr and aroFwt | [33] |
| pXMJ19-aroG | pXMJ19 containing aroG (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroF | pXMJ19 containing aroF gene (C. glutamicum ATCC 13032) | This work |
| pXMJ19-aroFwt | pXMJ19 containing aroFwt gene (pAP-B03) | This work |
| pXMJ19-aroH | pXMJ19 containing aroH (E. coli 3110) | This work |
| pXMJ19-pheA | pXMJ19 containing pheA (C. glutamicum ATCC 13032) | This work |
| pXMJ19-pheAfbr | pXMJ19 containing pheAfbr (pAP-B03) | This work |
| pXMJ19-pheAfbr-aroH | pXMJ19 containing pheAfbr (pAP-B03) and aroH (E. coli) | This work |
| pXMJ19-pheAfbr-aroFwt | pXMJ19 containing pheAfbr (pAP-B03) and aroFwt (pAP-B03) | This work |
ATCC American type culture collection
Primers used in this study
| Primers and reverse transcription primers . | Sequencea . |
|---|---|
| Primers | |
| aroG-F | 5′-CTAGTCTAGAAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroG-R | 5′-TCCCCCGGGTTACTTGGCTGCTGCTCGGC-3′ |
| aroF-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroF-R | 5′-CGCGGATCCTTACTTGGCTGCTGCTCGGC-3′ |
| aroFwt-F | 5′-TCCCCGCGGAAAGGAGGACACGCATGCAAAAAGACGCGCTGAATAACG-3′ |
| aroFwt-R | 5′-CGCGGATCCCCGCTCGAGTTAAGCCACGCGAGCCGTCA-3′ |
| aroH-F | 5′-CGCGGATCCAAAGGAGGACACGCATGAACAGAACTGACGAACTCCGTAC-3′ |
| aroH-R | 5′-TCCCCCGGGCCGCTCGAGTCAGAAGCGGGTATCTACCGCA-3′ |
| pheA-F | 5′-CCCAAGCTTAAAGGAGGACACGC ATGAGCGACGCACCAACTGTTG-3′ |
| pheA-R | 5′-TCCCCCGGG CTAGTTAAGTTTCCTTCCTTCGCTTGCT-3′ |
| pheAfbr-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGACATCGGAAAACCCGTTACT-3′ |
| pheAfbr-R | 5′-TGCACTGCAGTCCCCGCGGTCAGGTTGGATCAACAGGCACTA-3′ |
| Primers for RT-PCR | |
| aroH_F | 5′-ACTGACGAACTCCGTACTGC-3′ |
| aroH_R | 5′-TCGCTTATCTTCACCATTCA-3′ |
| aroFwt_F | 5′-AGACGCGCTGAATAACGTAC-3′ |
| aroFwt_R | 5′-ATCGACGAGCATATTCCAGA-3′ |
| pheA_F | 5′-AAGCCCTCTACAAATTTGCC-3′ |
| pheA_R | 5′-GTTGGAGCCCTGGTCAA-3′ |
| pheAfbr_F | 5′-GCGCTGGATGAAAAATTATT-3′ |
| pheAfbr_R | 5′-GCTTTACCGAGCGTAATTAA-3′ |
| Primers and reverse transcription primers . | Sequencea . |
|---|---|
| Primers | |
| aroG-F | 5′-CTAGTCTAGAAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroG-R | 5′-TCCCCCGGGTTACTTGGCTGCTGCTCGGC-3′ |
| aroF-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroF-R | 5′-CGCGGATCCTTACTTGGCTGCTGCTCGGC-3′ |
| aroFwt-F | 5′-TCCCCGCGGAAAGGAGGACACGCATGCAAAAAGACGCGCTGAATAACG-3′ |
| aroFwt-R | 5′-CGCGGATCCCCGCTCGAGTTAAGCCACGCGAGCCGTCA-3′ |
| aroH-F | 5′-CGCGGATCCAAAGGAGGACACGCATGAACAGAACTGACGAACTCCGTAC-3′ |
| aroH-R | 5′-TCCCCCGGGCCGCTCGAGTCAGAAGCGGGTATCTACCGCA-3′ |
| pheA-F | 5′-CCCAAGCTTAAAGGAGGACACGC ATGAGCGACGCACCAACTGTTG-3′ |
| pheA-R | 5′-TCCCCCGGG CTAGTTAAGTTTCCTTCCTTCGCTTGCT-3′ |
| pheAfbr-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGACATCGGAAAACCCGTTACT-3′ |
| pheAfbr-R | 5′-TGCACTGCAGTCCCCGCGGTCAGGTTGGATCAACAGGCACTA-3′ |
| Primers for RT-PCR | |
| aroH_F | 5′-ACTGACGAACTCCGTACTGC-3′ |
| aroH_R | 5′-TCGCTTATCTTCACCATTCA-3′ |
| aroFwt_F | 5′-AGACGCGCTGAATAACGTAC-3′ |
| aroFwt_R | 5′-ATCGACGAGCATATTCCAGA-3′ |
| pheA_F | 5′-AAGCCCTCTACAAATTTGCC-3′ |
| pheA_R | 5′-GTTGGAGCCCTGGTCAA-3′ |
| pheAfbr_F | 5′-GCGCTGGATGAAAAATTATT-3′ |
| pheAfbr_R | 5′-GCTTTACCGAGCGTAATTAA-3′ |
RT reverse transcription
aUnderlining indicates restriction enzyme sites; bold indicates the location of the Shine-Dalgarno sequence added artificially
Primers used in this study
| Primers and reverse transcription primers . | Sequencea . |
|---|---|
| Primers | |
| aroG-F | 5′-CTAGTCTAGAAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroG-R | 5′-TCCCCCGGGTTACTTGGCTGCTGCTCGGC-3′ |
| aroF-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroF-R | 5′-CGCGGATCCTTACTTGGCTGCTGCTCGGC-3′ |
| aroFwt-F | 5′-TCCCCGCGGAAAGGAGGACACGCATGCAAAAAGACGCGCTGAATAACG-3′ |
| aroFwt-R | 5′-CGCGGATCCCCGCTCGAGTTAAGCCACGCGAGCCGTCA-3′ |
| aroH-F | 5′-CGCGGATCCAAAGGAGGACACGCATGAACAGAACTGACGAACTCCGTAC-3′ |
| aroH-R | 5′-TCCCCCGGGCCGCTCGAGTCAGAAGCGGGTATCTACCGCA-3′ |
| pheA-F | 5′-CCCAAGCTTAAAGGAGGACACGC ATGAGCGACGCACCAACTGTTG-3′ |
| pheA-R | 5′-TCCCCCGGG CTAGTTAAGTTTCCTTCCTTCGCTTGCT-3′ |
| pheAfbr-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGACATCGGAAAACCCGTTACT-3′ |
| pheAfbr-R | 5′-TGCACTGCAGTCCCCGCGGTCAGGTTGGATCAACAGGCACTA-3′ |
| Primers for RT-PCR | |
| aroH_F | 5′-ACTGACGAACTCCGTACTGC-3′ |
| aroH_R | 5′-TCGCTTATCTTCACCATTCA-3′ |
| aroFwt_F | 5′-AGACGCGCTGAATAACGTAC-3′ |
| aroFwt_R | 5′-ATCGACGAGCATATTCCAGA-3′ |
| pheA_F | 5′-AAGCCCTCTACAAATTTGCC-3′ |
| pheA_R | 5′-GTTGGAGCCCTGGTCAA-3′ |
| pheAfbr_F | 5′-GCGCTGGATGAAAAATTATT-3′ |
| pheAfbr_R | 5′-GCTTTACCGAGCGTAATTAA-3′ |
| Primers and reverse transcription primers . | Sequencea . |
|---|---|
| Primers | |
| aroG-F | 5′-CTAGTCTAGAAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroG-R | 5′-TCCCCCGGGTTACTTGGCTGCTGCTCGGC-3′ |
| aroF-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGAGTTCTCCAGTCTCACTCGAAAA-3′ |
| aroF-R | 5′-CGCGGATCCTTACTTGGCTGCTGCTCGGC-3′ |
| aroFwt-F | 5′-TCCCCGCGGAAAGGAGGACACGCATGCAAAAAGACGCGCTGAATAACG-3′ |
| aroFwt-R | 5′-CGCGGATCCCCGCTCGAGTTAAGCCACGCGAGCCGTCA-3′ |
| aroH-F | 5′-CGCGGATCCAAAGGAGGACACGCATGAACAGAACTGACGAACTCCGTAC-3′ |
| aroH-R | 5′-TCCCCCGGGCCGCTCGAGTCAGAAGCGGGTATCTACCGCA-3′ |
| pheA-F | 5′-CCCAAGCTTAAAGGAGGACACGC ATGAGCGACGCACCAACTGTTG-3′ |
| pheA-R | 5′-TCCCCCGGG CTAGTTAAGTTTCCTTCCTTCGCTTGCT-3′ |
| pheAfbr-F | 5′-CCCAAGCTTAAAGGAGGACACGCATGACATCGGAAAACCCGTTACT-3′ |
| pheAfbr-R | 5′-TGCACTGCAGTCCCCGCGGTCAGGTTGGATCAACAGGCACTA-3′ |
| Primers for RT-PCR | |
| aroH_F | 5′-ACTGACGAACTCCGTACTGC-3′ |
| aroH_R | 5′-TCGCTTATCTTCACCATTCA-3′ |
| aroFwt_F | 5′-AGACGCGCTGAATAACGTAC-3′ |
| aroFwt_R | 5′-ATCGACGAGCATATTCCAGA-3′ |
| pheA_F | 5′-AAGCCCTCTACAAATTTGCC-3′ |
| pheA_R | 5′-GTTGGAGCCCTGGTCAA-3′ |
| pheAfbr_F | 5′-GCGCTGGATGAAAAATTATT-3′ |
| pheAfbr_R | 5′-GCTTTACCGAGCGTAATTAA-3′ |
RT reverse transcription
aUnderlining indicates restriction enzyme sites; bold indicates the location of the Shine-Dalgarno sequence added artificially
Corynebacterium glutamicum transformation
Corynebacterium glutamicum competent cells for electroporation were prepared by the method described by Xu et al. [30]. Briefly, a colony of C. glutamicum was inoculated into 30 ml of LBG (LB medium with 5 g l−1 glucose) and cultivated overnight for about 16 h at 30 °C with agitation (200 rpm). The overnight cell culture was diluted into Epo medium (10 g l−1 tryptone, 5 g l−1 yeast extract, 10 g l−1 NaCl, 4 g l−1 isonicotinic acid hydrazide, 25 g l−1 glycine, 0.1 % Tween 80) to an optical density at 600 nm (OD600) of 0.3 and cultured at 200 rpm and 30 °C until the OD600 reached 0.9. The culture was then chilled on ice for 10 min, harvested by centrifugation (4,000 g, 10 min) and washed four times with 15 ml ice-cold 10 % (v/v) glycerol; the cells were re-suspended in 0.2 ml 10 % (v/v) glycerol, frozen and stored as aliquots at −70 °C. For electro-transformation, the aliquots of competent cells were first thawed on ice, and 1–5 μl DNA was added. The mixture was transferred to a cold electroporation cuvette (gap 0.1 cm) and electroporated at 1.8 kV using a 10-ms (approx.) pulse. Immediately after electroporation, 1 ml LBHIS media (5 g l−1 tryptone, 5 g l−1 NaCl, 2.5 g l−1 yeast extract, 18.5 g l−1 Brain Heart infusion powder, 91 g l−1 sorbitol) was added to the cuvette, and the contents were first mixed and then transferred to a 2-ml sterile Eppendorf tube. The mixture was then incubated at 30 °C for 2 h before being plated on LBHIS agar containing the appropriate antibiotics.
RNA extraction
Cells were harvested (2,000 g, 5 min) and washed twice with double distilled water and stored at −80 °C until RNA preparation. Total RNA from the untreated control and D-limonene-treated cells with and without exogenous ergosterol was extracted using Trizol reagent (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. The RNA was quantified and checked in a Nanodrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington, DE) at 260 and 280 nm. The integrity of the isolated RNA was verified using an automated electrophoresis system (Bio-Rad, Hercules, CA).
Culture conditions and medium
Corynebacterium glutamicum strains were grown at 30 °C in LBG medium. Where necessary, an antibiotic, such as ampicillin (100 μg ml−1), kanamycin (50 μg ml−1) or chloramphenicol (34 μg ml−1), was supplemented to the medium. A 10 % (v/v) inoculum of an overnight culture (18 h) was used for further studies. To induce the expression of the various genes carried by the plasmids, isopropyl-β-D-thiogalactopyranoside (IPTG) was added to a final concentration of 0.1 mM. Flask cultivations were carried out in 20 ml of medium in a 250-ml Erlenmeyer flask medium at 30 °C with agitation at 200 rpm.
The seed medium contained (g l−1) glucose (35), corn steep liquor (35), (NH4)2SO4 (5), urea (2), KH2PO4 (1), MgSO4·7H2O (0.5); the pH was 7.2. The shake flask fermentation medium contained (g l−1) glucose (90), (NH4)2SO4 (25), corn steep liquor (8), KH2PO4 (1), MgSO4 (0.5), CaCO3 (20); the pH was 7.2.
RT-PCR assay
cDNA was synthesized from 5 μg of total RNA using the Primescript ® RT Reagent Kit Perfect Real Time (Takara, Dalian, China), and the cDNA obtained was used as the template in the subsequent PCR assay. All primer sequences are given in Table 1. The efficiency and specificity of the primers were determined from dilution experiments and melting curves, respectively. RT-PCR experiments were performed using PrimeSTAR® HS DNA Polymerase (Takara), and the parameters for PCR cycling were: denaturation at 95 °C for 5 min, degeneration at 95 °C for 30 s, annealing at 65 °C for 30 s, extension at 72 °C for the associated time (1 kb min−1), final extension at 72 °C for 10 min, and heat preservation at 12 °C.
SDS-PAGE analysis
Analysis by SDS-PAGE was performed in 10 % acrylamide gels, and the proteins were visualized by staining with Coomassie brilliant blue R-250. About 1.0 OD600 of cell culture was obtained after 12 h of induction. Whole cells were harvested at 4,000 g for 10 min and then washed third times with double distilled water. The cells were then resuspended in 40 μl lysozyme (20 mM) and lysed at 37 °C for 1 h, following which 10 μl of the 5× loading buffer was added for the SDS-PAGE analysis.
Analysis of fermentation parameters
The cell concentration was measured at OD600 on a spectrophotometer-722 (Third Analytical Instrument Factory, Shanghai, China) after the appropriate dilution. To determine the concentration of intracellular metabolites, 5 ml of cell culture was rapidly transferred into a centrifuge tube containing 20 ml of precooled glycerol-NaCl (glycerol and 13.5 g l−1 NaCl solution; volume ratio 1:1) and the mixture centrifuged (10,000 g, 3 min, −19 °C). The cells were then resuspended in 1 ml of precooled 50 % methanol solution in a centrifuge tube and the centrifuge tube placed in liquid nitrogen for 2–3 min and then thawed on ice; the freezing–thawing procedures were repeated three times. The supernatant obtained after centrifugation was either directly analyzed by high-performance liquid chromatography (HPLC) to determine the concentration of metabolites or preserved at −80 °C. For shikimate analysis, 5 ml of cell culture was centrifuged (7,000 g, 10 min), and the supernatant filtered through a filtration membrane (diameter 25 mm; pore size 0.22 μm. The filtered sample was analyzed on an Agilent 1200 HPLC system (Agilent Technologies, Santa Clara, CA) which was equipped with a 250 × 4.6-mm ZORBAX SB-Aq column (Agilent Technologies), a standard G 1329A autosampler (Agilent Technologies) and a G13158 diode array detector (DAD) (Agilent Technologies). Na2HPO4 (0.138 mol l−1) and acetonitrile (1 %, v/v) adjusted to pH 2.0 with phosphoric acid were used as the mobile phase at a flow rate of 1.0 ml min−1. The detection wavelength was 210 nm, and the column temperature was maintained at 35 °C.
To determine the concentration of amino acids, 5 ml of cell culture or intracellular supernatant was centrifuged (10,000 g, 10 min) and the supernatant was diluted with trichloroacetic acid (5 g l−1) and filtered through a membrane (pore size 0.22 μm). Amino acids were precolumn derivatized by o-phthaldialdehyde (OPA) and then analyzed on an Agilent 1100 HPLC system (Agilent Technologies) equipped with a reverse-phase column (Zorbax Eclipse-AAA) and an UV detector at 338 nm according to the procedure established by Henderson et al. [11].
Results
Single overexpression of the committed enzymes driving more flux to l-Phe
In C. glutamicum, the synthesis of l-Phe occurs via the shikimate pathway and the branch acid pathway. To evaluate the rate-limiting steps of the l-Phe synthesis pathways, we first examined the committed genes aroG, aroF, pheA, aroFwt and pheAfbr (Table 1). As shown in Table 3, under specific conditions, strains C. glutamicum ATCC 13032, C. glutamicum 19G (aroG), C. glutamicum 19F (aroF), C. glutamicum 19Fwt (aroFwt), C. glutamicum 19A (pheA) and C. glutamicum 19Afbr (pheAfbr) accumulated l-Phe to levels of 0.34 ± 0.03, 0.43 ± 0.01, 0.56 ± 0.06, 0.88 ± 0.06, 0.27 ± 0.06 and 1.34 ± 0.02 g l−1, respectively. Clearly, single overexpression of the mutated gene aroFwt or pheAfbr significantly improved l-Phe production. Compared with l-Phe, l-Tyr as the end byproduct was not increased by the upregulation of aroFwt or pheAfbr; to the contrary, single overexpression of pheAfbr resulted in a decrease of l-Tyr accumulation (Table 3), indicating that more carbon flux was driven to l-Phe.
Comparison of l-phenylalanine, l-tyrosine and shikimate production in different strains
| Strains . | l-Phe (g l−1) . | l-Tyr (g l−1) . | Shikimate (g l−1) . |
|---|---|---|---|
| C. glutamicum ATCC 13032 | 0.34 ± 0.03 | 0.260 ± 0.02 | 0.29 ± 0.08 |
| C. glutamicum 19G | 0.43 ± 0.01 | 0.06 ± 0.05 | 0.69 ± 0.07 |
| C. glutamicum 19F | 0.56 ± 0.06 | 0.37 ± 0.04 | 1.54 ± 0.04 |
| C. glutamicum 19Fwt | 0.88 ± 0.06 | 0.27 ± 0.01 | 5.67 ± 0.05 |
| C. glutamicum 19A | 0.27 ± 0.06 | 0.32 ± 0.02 | 0.32 ± 0.04 |
| C. glutamicum 19Afbr | 1.34 ± 0.02 | 0.04 ± 0.07 | 0.28 ± 0.05 |
| Strains . | l-Phe (g l−1) . | l-Tyr (g l−1) . | Shikimate (g l−1) . |
|---|---|---|---|
| C. glutamicum ATCC 13032 | 0.34 ± 0.03 | 0.260 ± 0.02 | 0.29 ± 0.08 |
| C. glutamicum 19G | 0.43 ± 0.01 | 0.06 ± 0.05 | 0.69 ± 0.07 |
| C. glutamicum 19F | 0.56 ± 0.06 | 0.37 ± 0.04 | 1.54 ± 0.04 |
| C. glutamicum 19Fwt | 0.88 ± 0.06 | 0.27 ± 0.01 | 5.67 ± 0.05 |
| C. glutamicum 19A | 0.27 ± 0.06 | 0.32 ± 0.02 | 0.32 ± 0.04 |
| C. glutamicum 19Afbr | 1.34 ± 0.02 | 0.04 ± 0.07 | 0.28 ± 0.05 |
l-Phe l-phenylalanine, l-Tyr l-tyrosine
Data are presented as the mean ± standard deviation (SD) (n = 3)
Comparison of l-phenylalanine, l-tyrosine and shikimate production in different strains
| Strains . | l-Phe (g l−1) . | l-Tyr (g l−1) . | Shikimate (g l−1) . |
|---|---|---|---|
| C. glutamicum ATCC 13032 | 0.34 ± 0.03 | 0.260 ± 0.02 | 0.29 ± 0.08 |
| C. glutamicum 19G | 0.43 ± 0.01 | 0.06 ± 0.05 | 0.69 ± 0.07 |
| C. glutamicum 19F | 0.56 ± 0.06 | 0.37 ± 0.04 | 1.54 ± 0.04 |
| C. glutamicum 19Fwt | 0.88 ± 0.06 | 0.27 ± 0.01 | 5.67 ± 0.05 |
| C. glutamicum 19A | 0.27 ± 0.06 | 0.32 ± 0.02 | 0.32 ± 0.04 |
| C. glutamicum 19Afbr | 1.34 ± 0.02 | 0.04 ± 0.07 | 0.28 ± 0.05 |
| Strains . | l-Phe (g l−1) . | l-Tyr (g l−1) . | Shikimate (g l−1) . |
|---|---|---|---|
| C. glutamicum ATCC 13032 | 0.34 ± 0.03 | 0.260 ± 0.02 | 0.29 ± 0.08 |
| C. glutamicum 19G | 0.43 ± 0.01 | 0.06 ± 0.05 | 0.69 ± 0.07 |
| C. glutamicum 19F | 0.56 ± 0.06 | 0.37 ± 0.04 | 1.54 ± 0.04 |
| C. glutamicum 19Fwt | 0.88 ± 0.06 | 0.27 ± 0.01 | 5.67 ± 0.05 |
| C. glutamicum 19A | 0.27 ± 0.06 | 0.32 ± 0.02 | 0.32 ± 0.04 |
| C. glutamicum 19Afbr | 1.34 ± 0.02 | 0.04 ± 0.07 | 0.28 ± 0.05 |
l-Phe l-phenylalanine, l-Tyr l-tyrosine
Data are presented as the mean ± standard deviation (SD) (n = 3)
To investigate the effect of disturbance by single overexpression of the above-mentioned genes, shikimate as the committed intermediate was also analyzed. As shown in Table 3, upregulation of the upstream genes aroG, aroF or aroFwt could lead to an apparent accumulation of shikimate which was consistent with the accumulation of l-Phe. Specially, the recombinant C. glutamicum 19Fwt (aroFwt) increased shikimate accumulation to 5.67 ± 0.05 g l−1, suggesting that other key genes must be involved downstream of the aromatic amino acid synthesis. In addition, no l-Trp was detected throughout the cultivation process.
Introduction of the wild-type aroH gene for improving l-Phe production
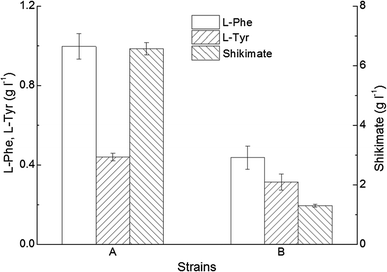
Accumulation of metabolites by the recombinant C. glutamicum 19H (aroH from E. coli) in the absence of l-tryptophan (l-Trp) (a) and following the addition of 1.0 g l−1 l-Trp to the culture (b). Data are presented as the mean ± standard deviation (SD) (n = 3)
Production of l-Phe by overexpression of aroH and pheAfbr
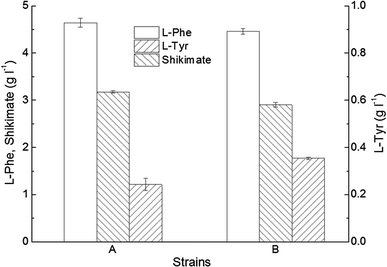
Accumulation of metabolites by co-overexpression of pheAfbr and aroFwt, and pheAfbr and aroH. a The recombinant C. glutamicum 19AfbrH, b the recombinant C. glutamicum 19AfbrFwt. Data are presented as the mean ± SD (n = 3)
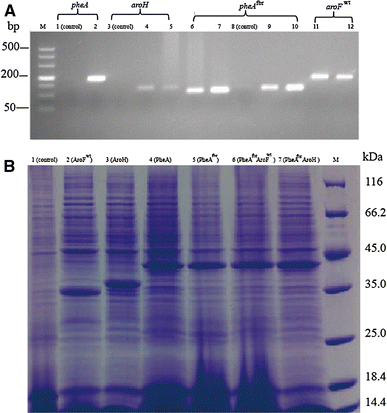
Analysis of the transcript level of key enzymes in recombinant C. glutamicum by reverse transcription (RT)-PCR and sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) with Coomassie brilliant blue staining of protein products. a RT-PCR. Lanes: M DL 500 DNA ladder, 1 control (pheA), 2 C. glutamicum 19A (pheA), 3 control (aroH), 4 C. glutamicum 19H (aroH), 5 C. glutamicum 19AfbrH (aroH), 6 C. glutamicum 19Afbr (pheAfbr), 7 C. glutamicum 19AfbrH (pheAfbr), 8 control (pheAfbr), 9 C. glutamicum 19Afbr (pheAfbr), 10 C. glutamicum 19AfbrFwt (pheAfbr), 11 C. glutamicum 19Fwt (aroFwt), 12 C. glutamicum 19AfbrFwt (aroFwt). b SDS-PAGE. Lanes 1 C. glutamicum ATCC 13032, 2 C. glutamicum 19Fwt, 3 C. glutamicum 19H, 4 C. glutamicum 19A, 5 C. glutamicum 19Afbr, 6 C. glutamicum 19AfbrFwt, 7 C. glutamicum AfbrH, M molecular weight marker
Discussion
In nature, the biosynthetic pathway of amino acids is strictly regulated (generally by feedback inhibition) in C. glutamicum. As a result, modification of the committed enzymes to relieve or eliminate the feedback inhibition by the end-products is crucial to increasing the target compound [29, 32]. In this study, the wild-type aroH gene from E. coli was cloned into C. glutamicum to optimize the shikimate pathway for producing l-Phe.
Many studies have been carried out in E. coli, resulting in a substantial improvement in the production of l-Phe [1, 8, 13, 33]. When we first investigated the committed genes involved in l-Phe production, we found that single overexpression of the native wild-type aroG, aroF and pheA genes had no obvious effect on the yield of l-Phe (Table 3), further demonstrating that the synthesis of l-Phe in C. glutamicum is extremely tightly regulated. To the contrary, when either the mutated aroFwt (E. coli) or pheAfbr (E. coli) gene was introduced into C. glutamicum, the titer of l-Phe was dramatically increased (Table 3), indicating that feedback inhibition to DS (encoded by aroFwt) or CM-PDT [12] (encoded by pheAfbr) had occurred. Subsequent studies on the overexpression of the mutated aroFwt from E. coli in C. glutamicum revealed that shikimate was increased up to a high level (Table 3), demonstrating that other key enzymes are present in downstream of the pathway.
In E. coli, the DS enzyme encoded by aroH is inhibited by feedback from l-Trp, contributing 1 % to DS enzyme activity [13, 27]. In contrast, no aroH gene is naturally present in C. glutamicum, and thus the introduction and overexpression of the wild-type aroH gene from E. coli may be beneficial to the production of l-Phe by C. glutamicum. As expected, introduction of aroH achieved a substantial increase in l-Phe production without l-Trp accumulation, thereby demonstrating the complete inhibition of the native branch pathway for l-Trp synthesis (Fig. 1). The capacity of the aroH gene and the strategy of co-overexpression of the committed enzymes have been studied [19, 29, 32, 33]. As predicted, we found that co-overexpression of the rate-limiting enzymes DS and CM-PDT yielded a high titer of l-Phe (Fig. 3). Moreover, compared with C. glutamicum 19AfbrFwt, C. glutamicum 19AfbrH showed the better property, which revealed that although aroF was mutated, the release of feedback inhibition may be incomplete in vivo. To modulate and balance the pathway flux, we also investigated the expression levels of the key genes at the transcriptional and translational levels, respectively. At the transcriptional level, all of the genes studied were successfully transcribed (Fig. 4a). However, an unbalanced expression level was observed in the SDS-PAGE analysis. When two genes were co-expressed with one operon, the expression level of the gene further away from the promoter declined sharply and no obvious band was detected (Fig. 4b). Similar results have been reported in E. coli [19]. Consequently, to further increase the titer of the targeted product, subtle regulation of the committed genes might be crucial [5, 28].
In conclusion, for a comparative analysis of the committed enzymes involved in the aromatic amino acid synthesis pathway in C. glutamicum and E. coli, we used the wild-type gene aroH from E. coli for improving l-Phe production. As predicted, the introduction and overexpression of this gene in C. glutamicum remarkably increased l-Phe production by C. glutamicum. Compared with aroFwt, the aroH gene from E. coli was shown to be more effective for l-Phe production. Moreover, the observed unbalanced expression levels of the genes within one operon suggested that fine control of the committed enzymes will be very important for improving l-Phe production. The knowledge gained through this study will provide further insights into novel pathway engineering of C. glutamicum for l-Phe production.
Acknowledgments
We thank Professor Byong Lee at Jiangnan University for his discussion and revision. This work was financially supported by the Key Program of National Natural Science Foundation of China (31130043), the National Natural Science Foundation of China (31200020, 31000054, 31171638), the Priority Academic Program Development of Jiangsu Higher Education Institutions, the Jiangsu Planned Projects for Postdoctoral Research Funds (1101053C) and the Independent Innovation Program of Jiangnan University (JUSRP111A23).
References
Henderson JW, Ricker RD, Bidlingmeyer BA et al (2000) Rapid, accurate, sensitive, and reproducible HPLC analysis of amino acids. USA (Agilent App Note 5980–1193E). Agilent Technologies, Santa Clara