Abstract
The dysfunction of placenta development is correlated to the defects of pregnancy and fetal growth. The detailed molecular mechanism of placenta development is not identified in humans due to the lack of material in vivo. Trophoblast (TB) lineage derived from human embryonic stem cells (hESCs) induced by bone morphogenetic protein 4 (BMP4) has been applied as a model for studying TB lineage specification in vitro. With the development of single-cell sequencing technology, it became possible to detect the transcriptome of the post-implantation embryo at unprecedented precision. In this study, we reanalyzed single-cell RNA-seq of post-implantation embryos derived from two separate groups and identified different subtypes of trophoblast cells and their marker, respectively. At the same time, we focused on the gene expression patterns of trophoblast-specific transcription factors in different models. Our analysis sheds new light on the transcription regulation mechanism of trophoblast differentiation at the early stage of pregnancy establishment in human.




Similar content being viewed by others
References
Artus J, Chazaud C (2014) A close look at the mammalian blastocyst: epiblast and primitive endoderm formation. Cell Mol Life Sci 71:3327–3338. https://doi.org/10.1007/s00018-014-1630-3
Blakeley P, Fogarty NME, del Valle I, Wamaitha SE, Hu TX, Elder K, Snell P, Christie L, Robson P, Niakan KK (2015) Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142:3151–3165. https://doi.org/10.1242/dev.123547
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nature Biotechnology 34:525–527. https://doi.org/10.1038/nbt.3519
Chazaud C, Yamanaka Y (2016) Lineage specification in the mouse pre-implantation embryo. Development 143:1063–1074. https://doi.org/10.1242/dev.128314
Chen Y, Chen A (2019) Unveiling the gene regulatory landscape in diseases through the identification of DNase I-hypersensitive sites (Review). Biomedical Reports 11:87–97. https://doi.org/10.3892/br.2019.1233
Chen T, Yang S, Xu J, Lu W, Xie X (2020) Transcriptome sequencing profiles of cervical cancer tissues and SiHa cells. Funct Integr Genomics 20:211–221. https://doi.org/10.1007/s10142-019-00706-y
Chen D, Liu W, Zimmerman J, Pastor WA, Kim R, Hosohama L, Ho J, Aslanyan M, Gell JJ, Jacobsen SE, Clark AT (2018) The TFAP2C-Regulated OCT4 Naive Enhancer Is Involved in Human Germline Formation. Cell Reports 25:3591-3602.e5
Fernandez NF, Gundersen GW, Rahman A et al (2017) Clustergrammer, a web-based heatmap visualization and analysis tool for high-dimensional biological data. sdata 4:170151. https://doi.org/10.1038/sdata.2017.151
He J, Huisken J (2020) Image quality guided smart rotation improves coverage in microscopy. Nat Commun 11:1–9. https://doi.org/10.1038/s41467-019-13821-y
Jain A, Ezashi T, Roberts RM, Tuteja G (2017) Deciphering transcriptional regulation in human embryonic stem cells specified towards a trophoblast differentiation in both normal development and disease. PNAS 113:E3882–E3891. https://doi.org/10.1073/pnas.1604747113
Jékely G, Arnone MI (2019) Editorial overview: Tissue-level dynamics in development and evolution. Current Opinion in Genetics & Development 57:iii–v.fate. Sci Rep 7:1–12. https://doi.org/10.1038/s41598-017-17614-5
Knöfler M, Haider S, Saleh L, Pollheimer J, Gamage TKJB, James J (2019) Human placenta and trophoblast development: key molecular mechanisms and model systems. Cell Mol Life Sci 76:3479–3496. https://doi.org/10.1007/s00018-019-03104-6
Krendl C, Shaposhnikov D, Rishko V, Ori C, Ziegenhain C, Sass S, Simon L, Müller NS, Straub T, Brooks KE, Chavez SL, Enard W, Theis FJ, Drukker M (2017) GATA2/3-TFAP2A/C transcription factor network couples human pluripotent stem cell differentiation to trophectoderm with repression of pluripotency. PNAS 114:E9579–E9588. https://doi.org/10.1073/pnas.1708341114
Kribelbauer JF, Rastogi C, Bussemaker HJ, Mann RS (2019) Low-affinity binding sites and the transcription factor specificity paradox in eukaryotes. Annual Review of Cell and Developmental Biology 35:357–379. https://doi.org/10.1146/annurev-cellbio-100617-062719
Li QV, Rosen BP, Huangfu D (2020) Decoding pluripotency: genetic screens to interrogate the acquisition, maintenance, and exit of pluripotency. WIREs Systems Biology and Medicine 12:e1464. https://doi.org/10.1002/wsbm.1464
Liu Y, Ding D, Liu H, Sun X (2017) The accessible chromatin landscape during conversion of human embryonic stem cells to trophoblast by bone morphogenetic protein 4. Biol Reprod 96:1267–1278. https://doi.org/10.1093/biolre/iox028
Lv B, An Q, Zeng Q, Zhang X, Lu P, Wang Y, Zhu X, Ji Y, Fan G, Xue Z (2019) Single-cell RNA sequencing reveals regulatory mechanism for trophoblast cell-fate divergence in human peri-implantation conceptuses. PLOS Biology 17:e3000187. https://doi.org/10.1371/journal.pbio.3000187
Macabenta F, Stathopoulos A (2019) Sticking to a plan: adhesion and signaling control spatial organization of cells within migrating collectives. Current Opinion in Genetics & Development 57:39–46. https://doi.org/10.1016/j.gde.2019.07.003
Majewska M, Lipka A, Paukszto L, Jastrzebski JP, Myszczynski K, Gowkielewicz M, Jozwik M, Majewski MK (2017) Transcriptome profile of the human placenta. Funct Integr Genomics 17:551–563. https://doi.org/10.1007/s10142-017-0555-y
Marchand M, Horcajadas JA, Esteban FJ, McElroy SL, Fisher SJ, Giudice LC (2011) Transcriptomic signature of trophoblast differentiation in a human embryonic stem cell model. Biol Reprod 84:1258–1271. https://doi.org/10.1095/biolreprod.110.086413
McInnes L, Healy J, Melville J (2018) UMAP: uniform manifold approximation and projection for dimension reduction. arXiv:180203426 [cs, stat]
Pastor WA, Liu W, Chen D, Ho J, Kim R, Hunt TJ, Lukianchikov A, Liu X, Polo JM, Jacobsen SE, Clark AT (2018) TFAP2C regulates transcription in human naive pluripotency by opening enhancers. Nat Cell Biol 20:553–564. https://doi.org/10.1038/s41556-018-0089-0
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Research 43:e47–e47
Rizzo R, Vercammen M, van de Velde H, Horn PA, Rebmann V (2011) The importance of HLA-G expression in embryos, trophoblast cells, and embryonic stem cells. Cell Mol Life Sci 68:341–352. https://doi.org/10.1007/s00018-010-0578-1
Roberts RM, Ezashi T, Sheridan MA, Yang Y (2018) Specification of trophoblast from embryonic stem cells exposed to BMP4. Biol Reprod 99:212–224. https://doi.org/10.1093/biolre/ioy070
Schubert C (2015) Single cell analysis probes variability in gene expression in early human embryo. Biol Reprod 93:108. https://doi.org/10.1095/biolreprod.115.134999
Seita Y, Su J, Wang B, Treff N, Lu CW (2012) CDX2 expression marks trophoblast progenitors from differentiating human pluripotent stem cells. Biol Reprod 87:53–53. https://doi.org/10.1093/biolreprod/87.s1.53
Soncin F, Natale D, Parast MM (2015) Signaling pathways in mouse and human trophoblast differentiation: a comparative review. Cell Mol Life Sci 72:1291–1302. https://doi.org/10.1007/s00018-014-1794-x
The Tabula Muris Consortium et al (2018) Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562:367–372. https://doi.org/10.1038/s41586-018-0590-4
Torres-Padilla M-E, Chambers I (2014) Transcription factor heterogeneity in pluripotent stem cells: a stochastic advantage. Development 141:2173–2181. https://doi.org/10.1242/dev.102624
Traag V (2015) louvain-igraph: v0.5.3. Zenodo
Maaten and Hinton (2008) Visualizing data using t-SNE. Journal of Machine Learning Research 9:2579–2605
Velicky P, Knöfler M, Pollheimer J (2016) Function and control of human invasive trophoblast subtypes: intrinsic vs. maternal control. Cell Adhesion & Migration 10:154–162. https://doi.org/10.1080/19336918.2015.1089376
Whited JL, Levin M (2019) Bioelectrical controls of morphogenesis: from ancient mechanisms of cell coordination to biomedical opportunities. Current Opinion in Genetics & Development 57:61–69. https://doi.org/10.1016/j.gde.2019.06.014
Wolf FA, Angerer P, Theis FJ (2018) SCANPY: large-scale single-cell gene expression data analysis. Genome Biology 19:15. https://doi.org/10.1186/s13059-017-1382-0
Xia B, Yanai I (2019) A periodic table of cell types. Development 146:dev169854. https://doi.org/10.1242/dev.169854
Xie C, Cai X, Wang X, Lin L, Long Y, Cui X (2018) Gene expression profiles of HTR8-S/Vneo cells after changes in ABCA1 expression. Funct Integr Genomics 18:725–735. https://doi.org/10.1007/s10142-018-0621-0
Yabe S, Alexenko AP, Amita M, Yang Y, Schust DJ, Sadovsky Y, Ezashi T, Roberts RM (2016) Comparison of syncytiotrophoblast generated from human embryonic stem cells and from term placentas. Proc Natl Acad Sci USA 113:E2598–E2607. https://doi.org/10.1073/pnas.1601630113
Zhou F, Wang R, Yuan P, Ren Y, Mao Y, Li R, Lian Y, Li J, Wen L, Yan L, Qiao J, Tang F (2019) Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 572:660–664. https://doi.org/10.1038/s41586-019-1500-0
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Funding
Yajun Liu was supported by the “Young scientist startup grand of The Second Affiliated Hospital of Zhengzhou University” and Foundation of Henan Educational Committee (CN) (19A320044).
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
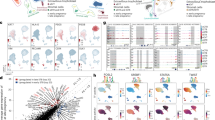
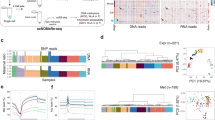
Figure S1 (A) Similarity matrix of the different samples in the RNA-seq dataset (GEO number: GSE73017). The cells in the matrix represent the similarity between rows, where red/blue represent positive/negative similarity (measured as 1-cosine distance). (B) Similarity matrix of the different samples in the RNA-seq dataset (array express number: E-MTAB-8186, E-MTAB-8181). Figure S2 (A) Violin plots showing the expression of EPI-specific TFs (POU5F1, SOX2, NANOG) in represented cell cluster and t-SNE visualization of the expression of these genes in 5911cells. The points show cells positioned in t-SNE space, lower right, with the more intense red hue indicating higher relative expression and gray signifying no expression. (B) Violin plots and t-SNE visualization showing the expression of TE-specific TFs (JUN, FOS, TFAP2C, TBX3, and OVOL1) in represented cell cluster and t-SNE visualization of the expression of these genes in 5911cells. (C) Violin plots showing the expression of PE-specific TFs (GATA4, GATA6, SOX17) are represented cell clusters and t-SNE visualization of the expression of these genes in 5911cells. (PDF 1282 kb)
Supplementary file 1
(SVG 825 kb)
Supplementary file 2
(PDF 234 kb)
Table S1
Transcriptional factors that are enriched in human trophoblast and ESCs.docx. (DOCX 19 kb)
Table S2
differential gene expression analysis between TE and EPI.txt. (TXT 1343 kb)
Table S3
Marker genes in each cluster after cluster by SCANPY. (TSV 27 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Zhang, Y., Li, S. et al. Gene expression pattern of trophoblast-specific transcription factors in trophectoderm by analysis of single-cell RNA-seq data of human blastocyst. Funct Integr Genomics 21, 205–214 (2021). https://doi.org/10.1007/s10142-021-00770-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-021-00770-3