Abstract
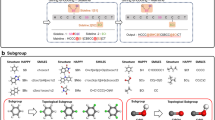
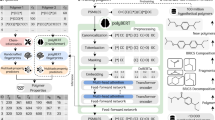
A deep neural network model generally consists of different modules that play essential roles in performing a task. The optimal design of a module for use in modeling a physical problem is directly related to the success of the model. In this work, the effectiveness of a number of special modules, the self-attention mechanism for recognizing the importance of molecular sequence information in a polymer, as well as the big-stride representation and conditional random field for enhancing the network ability to produce desired local configurations, is numerically studied. Network models containing these modules are trained by using the well documented data of the native structures of the HP model and assessed according to their capability in making structural predictions of unseen data. The specific network design of self-attention mechanism adopted here is modified from a similar idea in natural language recognition. The big-stride representation module introduced in this work is shown to drastically improve network’s capability to model polymer segments of strong lattice position correlations.
Similar content being viewed by others
References
Yang, L.; Tan, X.; Wang, Z.; Zhang, X. Supramolecular polymers: historical development, preparation, characterization, and functions. Chem. Rev. 2015, 115, 7196–7239.
Zhang, X.; Wang, C. Supramolecular amphiphiles. Chem. Soc. Rev. 2011, 40, 94–101.
Elshire, R. J.; Glaubitz, J. C.; Sun, Q.; Poland, J. A.; Kawamoto, K. Buckler, E. S.; Mitchel, S. E. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PloS One 2011, 6, e19379.
Saunders, M. G.; Voth, G. A. Coarse-graining methods for computational biology. Annual Rev. Biophys. 2013, 42, 73–93.
Perilla, J. R.; Goh, B. C.; Cassidy, C. K.; Liu, B.; Bernardi, R. C.; Rudack, T.; Yu, H.; Wu, Z.; Schulten, K. Molecular dynamics simulations of large macromolecular complexes. Curr. Opin. Struct. Biol. 2015, 31, 64–74.
Shakhnovich, E.; Farztdinov, G.; Gutin, A.; Karplus, M. Protein folding bottlenecks: A lattice Monte Carlo simulation. Phys. Rev. Lett. 1991, 67, 1665.
Scheraga, H. A.; Khalili, M.; Liwo, A. Protein-folding dynamics: overview of molecular simulation techniques. Annu. Rev. Phys. Chem. 2007, 58, 57–83.
Carrasquilla, J.; Melko, R. G. Machine learning phases of matter. Nat. Phys. 2017, 13, 431–434.
Smith, J. S.; Isayev, O.; Roitberg, A. E. ANI-1: an extensible neural network potential with DFT accuracy at force field computational cost. Chem. Sci. 2017, 8, 3192–3203.
Schütt, K. T.; Arbabzadah, F.; Chmiela, S.; Muller; K. R.; Tkatchenko, A. Quantum-chemical insights from deep tensor neural networks. Nat. Commun. 2017, 8, 1–8.
Wei, Q.; Melko, R. G.; Chen, J. Z. Y. Identifying polymer states by machine learning. Phys. Rev. E 2017, 95, 032504.
Lau, K. F.; Dill, K. A. A lattice statistical mechanics model of the conformational and sequence spaces of proteins. Macromolecules 1989, 22, 3986–3997.
Dill, K. A.; MacCallum, J. L. The protein-folding problem, 50 years on. Science 2012, 338, 1042–1046.
Hossain, M. S.; Salam, A. Text-to-3D Scene Generation using Semantic Parsing and Spatial Knowledge with Rule Based System. Int. J. Comp. Sci. Issues (IJCSI) 2017, 14, 37–41.
LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521(7553), 436–444.
Goodfellow, I.; Bengio, Y.; Courville, A. Deep learning. MIT Press, 2016.
Nielsen, M. A. Neural networks and deep learning. Determination Press San Francisco, CA, USA, 2015; Vol. 25.
Shalev-Shwartz, S.; Ben-David, S. Understanding machine learning: from theory to algorithms. Cambridge University Press, 2014.
Li, J.; Zhang, H.; Chen, J. Z. Y. Structural prediction and inverse design by a strongly correlated neural network. Phys. Rev. Lett. 2019, 123, 108002.
Cheng, J.; Dong, L.; Lapata, M. Long short-term memorynetworks for machine reading. Proceedings of the 2016 Conference on Empirical Methods in Natural Language Processing, Austin, Texas, 2016, pp. 551–561.
Parikh, A. P.; Täckström, O.; Das, D.; Uszkoreit, J. A decomposable attention model for natural language inference. Proceedings of the 2016 Conference on Empirical Methods in Natural Language Processing, Austin, Texas, 2016, pp. 2249–2255.
Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A. N.; Kaiser, L.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst., 2017.
Flory, P. J. Principles of polymer chemistry. Cornell University Press, 1953.
Landau, D.; Binder, K. A guide to Monte Carlo simulations in statistical physics. Cambridge University Press, 2021.
Allen, M. P.; Tildesley, D. J. Computer simulation of liquids. Oxford University Press, 2017.
Wang, S.; Peng, J.; Ma, J.; Xu; J. B. Protein secondary structure prediction using deep convolutional neural fields. Scient. Rep. 2016, 6, 1–11.
Chan, H. S.; Dill, K. A. Transition states and folding dynamics of proteins and heteropolymers. J. Chem. Phys. 1994, 100, 9238–9257.
Please seehttps://github.com/vvoelz/HPSandbox.
Wüst, T.; Landau, D. P. Versatile approach to access the low temperature thermodynamics of lattice polymers and proteins. Phys. Rev. Lett. 2009, 102, 178101.
Boškovič, B.; Brest, J. Genetic algorithm with advanced mechanisms applied to the protein structure prediction in a hydrophobic-polar model and cubic lattice. Appl. Soft Comput. 2016, 45, 61–70.
Yang, C. H.; Wu, K. C.; Lin, Y. S.; Chuang; L. Y.; Chang H. W. Protein folding prediction in the HP model using ions motion optimization with a greedy algorithm. BioData Mining 2018, 11, 1–14.
Li, Y. W.; Wuest, T.; Landau, D. P. Generic folding and transition hierarchies for surface adsorption of hydrophobic-polar lattice model proteins. Phys. Rev. E 2013, 87, 012706.
Wu, H.; Yang, R.; Fu, Q.; Chen, J. P.; Lu, W. Z.; Li, H. O. Research on predicting 2D-HP protein folding using reinforcement learning with full state space. Bmc Bioinformatics 2019, 20.
Please see https://github.com/Titanium-ALarx7/HP-ProteinPrediction-SCN.
He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition, Proceedings of the IEEE conference on computer vision and pattern recognition, 2016; pp. 770–778.
Hochreiter, S.; Schmidhuber, J. Long short-term memory. Neural Comput. 1997, 9, 1735–1780.
Devlin, J.; Chang, M. W.; Lee, K.; Toutanova, K. Bert: Pre-training of deep bidirectional transformers for language understanding. “NAACL-HLT2019”, Minneapolis, Minnesota, 2018, pp. 4171–4186.
Lafferty, J.; McCallum, A.; Pereira, F. C. Conditional random fields: Probabilistic models for segmenting and labeling sequence data. 2001.
Frauenkron, H.; Bastolla, U.; Gerstner, E.; Grassberger, P.; Nadler, W. New Monte Carlo algorithm for protein folding. Phys. Rev. Lett. 1998, 80, 3149.
Thachuk, C.; Shmygelska, A.; Hoos, H. H. A replica exchange Monte Carlo algorithm for protein folding in the HP model. BMC bioinformatics 2007, 8, 1–20.
Wüst, T.; Landau, D. The HP model of protein folding: A challenging testing ground for Wang-Landau sampling. Comp. Phys. Commun. 2008, 179, 124–127.
Acknowledgments
This work was financially supported by the National Natural Science Foundation of China (Nos. 21973018 and 21534002) and the Natural Sciences and Engineering Research Council (NSERC) of Canada.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare no interest conflict.
Electronic Supplementary Information
Rights and permissions
About this article
Cite this article
Wang, TY., Li, JF., Zhang, HD. et al. Designs to Improve Capability of Neural Networks to Make Structural Predictions. Chin J Polym Sci 41, 1477–1485 (2023). https://doi.org/10.1007/s10118-023-2910-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10118-023-2910-x