Abstract
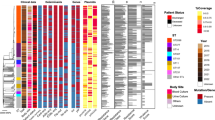
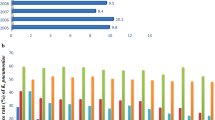
Vancomycin-resistant Enterococcus faecium (VREfm) emerged as an important cause of nosocomial infections worldwide. Previous studies based on molecular typing revealed that VREfm outbreaks are mainly associated with a particular genetic lineage, namely clonal complex 17 (CC17), which harbours either vanA or vanB gene cluster. The University Hospital of Lausanne faced several VREfm episodes of transmissions between 2014 and 2017. In this study, we used whole-genome sequencing (WGS) to investigate the relatedness of 183 VREfm isolates collected from 156 patients. Sequence types (ST) 17, ST80 and ST117 were the most predominant clones. Based on epidemiological data, 10 outbreaks were identified, which were caused by at least 13 distinct genotypes. The majority of isolates involved in outbreaks (91%) differed by only 0 to 3 SNPs. Four outbreaks involved more than one genotype and half of the cases considered as sporadic were possibly linked to an outbreak. By sequencing all isolates, we were able to better understand our local epidemiology of VREfm. The polyclonal structure observed between the different outbreaks strains, the high level of recombination detected in isolates, the time elapsed between admission and the first VREfm detection and the negative screening at admission support the hypothesis of the emergence of new VREfm clones within the hospitalised population.



Similar content being viewed by others
References
Woodford N, Stigter JM (1998) Molecular investigation of glycopeptide resistance in gram-positive bacteria. Methods Mol Med 15:579–615
Woodford N (2001) Epidemiology of the genetic elements responsible for acquired glycopeptide resistance in enterococci. Microb Drug Resist 7(3):229–236
DiazGranados CA, Jernigan JA (2005) Impact of vancomycin resistance on mortality among patients with neutropenia and enterococcal bloodstream infection. J Infect Dis 191(4):588–595
DiazGranados CA, Zimmer SM, Klein M, Jernigan JA (2005) Comparison of mortality associated with vancomycin-resistant and vancomycin-susceptible enterococcal bloodstream infections: a meta-analysis. Clin Infect Dis 41(3):327–333
Talaga-Cwiertnia K, Bulanda M (2018) Analysis of the world epidemiological situation among vancomycin-resistant Enterococcus faecium infections and the current situation in Poland. Przegl Epidemiol 72(1):3–15
Pinholt M, Gumpert H, Bayliss S, Nielsen JB, Vorobieva V, Pedersen M et al (2017) Genomic analysis of 495 vancomycin-resistant Enterococcus faecium reveals broad dissemination of a vanA plasmid in more than 19 clones from Copenhagen, Denmark. J Antimicrob Chemother 72(1):40–47
Raven KE, Gouliouris T, Brodrick H, Coll F, Brown NM, Reynolds R et al (2017) Complex routes of nosocomial vancomycin-resistant Enterococcus faecium transmission revealed by genome sequencing. Clin Infect Dis 64(7):886–893
de Been M, Pinholt M, Top J, Bletz S, Mellmann A, van Schaik W et al (2015) Core genome multilocus sequence typing scheme for high- resolution typing of Enterococcus faecium. J Clin Microbiol 53(12):3788–3797
Inouye M, Dashnow H, Raven LA, Schultz MB, Pope BJ, Tomita T et al (2014) SRST2: rapid genomic surveillance for public health and hospital microbiology labs. Genome Med 6(11):90
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25(14):1754–1760
Garrison EM, Gabor. (2012) Haplotype-based variant detection from short-read sequencing. ARXIV [Internet]
Kurtz S, Phillippy A, Delcher AL, Smoot M, Shumway M, Antonescu C et al (2004) Versatile and open software for comparing large genomes. Genome Biol 5(2):R12
Croucher NJ, Page AJ, Connor TR, Delaney AJ, Keane JA, Bentley SD et al (2015) Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res 43(3):e15
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27(2):221–224
Jia B, Raphenya AR, Alcock B, Waglechner N, Guo P, Tsang KK et al (2017) CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res 45(D1):D566–Dd73
Chen L, Yang J, Yu J, Yao Z, Sun L, Shen Y et al (2005) VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res 33(Database issue):D325–D328
Chen L, Zheng D, Liu B, Yang J, Jin Q (2016) VFDB 2016: hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res 44(D1):D694–D697
Pinholt M, Larner-Svensson H, Littauer P, Moser CE, Pedersen M, Lemming LE et al (2015) Multiple hospital outbreaks of vanA Enterococcus faecium in Denmark, 2012-13, investigated by WGS, MLST and PFGE. J Antimicrob Chemother 70(9):2474–2482
Howden BP, Holt KE, Lam MM, Seemann T, Ballard S, Coombs GW et al (2013) Genomic insights to control the emergence of vancomycin-resistant enterococci. mBio 4(4)
Moradigaravand D, Gouliouris T, Blane B, Naydenova P, Ludden C, Crawley C et al (2017) Within-host evolution of enterococcus faecium during longitudinal carriage and transition to bloodstream infection in immunocompromised patients. Genome Med 9(1):119
van Hal SJ, Ip CL, Ansari MA, Wilson DJ, Espedido BA, Jensen SO et al (2016) Evolutionary dynamics of Enterococcus faecium reveals complex genomic relationships between isolates with independent emergence of vancomycin resistance. Microbial Genomics 2(1)
Funding
The Service of Hospital Preventive Medicine of the Lausanne University Hospital funded this project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in this study involving human participants were in accordance with the ethical standards of the regional and national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
For this type of study, informed consent is not required.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Abdelbary, M.H.H., Senn, L., Greub, G. et al. Whole-genome sequencing revealed independent emergence of vancomycin-resistant Enterococcus faecium causing sequential outbreaks over 3 years in a tertiary care hospital. Eur J Clin Microbiol Infect Dis 38, 1163–1170 (2019). https://doi.org/10.1007/s10096-019-03524-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-019-03524-z