Abstract
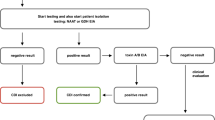
We aimed to assess differences in bacterial intensities of Bacteroidetes phylum and different clostridial species in the human intestines with respect to C. difficile infection. Patients with a stool assay for C. difficile toxin were identified via the microbiology laboratory in our institute. Bacterial populations were quantified from stool samples of four groups of patients: Group I—patients with C. difficile associated diarrhea (CDAD); Group II—asymptomatic C. difficile carriers; Group III—patients with non-C. difficile diarrhea; Group IV—patients with no diarrhea and negative stool samples for the C. difficile toxin (control group). Stool was examined for three genes—C. difficile toxin A gene, 16S rRNA gene from Clostridium thermocellum representing other clostridial species, and 16S rRNA gene from Bacteroides fragilis representing the Bacteroidetes phylum. Fifty-nine patients underwent analysis of the stool (CDAD group 14, carriers group 14, non-C. difficile diarrhea group 16, control group 15). C. difficile concentration was highest in the CDAD group, followed by the carriers group. Higher concentrations of both clostridial species and Bacteriodetes were observed in the control and non-C. difficile diarrhea groups compared to the CDAD and carriers groups. We demonstrated an inverse association between infection with C. difficile and the abundance of Bacteroidetes phylum and other clostridial species in human intestines. Studies with larger samples and broader diagnostic procedures are needed in order to better explore and understand this association.

Similar content being viewed by others
References
Kelly CP, Pothoulakis C, LaMont JT (1994) Clostridium difficile colitis. N Engl J Med 330:257–262
McFarland LV, Mulligan ME, Kwok RYY, Stamm WE (1989) Nosocomial acquisition of Clostridium difficile infection. N Engl J Med 320:204–210
Warny M, Vaerman JP, Avesani V, Delmée M (1994) Human antibody response to Clostridium difficile toxin A in relation to clinical course of infection. Infect Immun 62:384–389
Kyne L, Warny M, Qamar A, Kelly CP (2000) Asymptomatic carriage of Clostridium difficile and serum levels of IgG antibody against toxin A. N Engl J Med 342:390–397
Hopkins MJ, Macfarlane GT (2002) Changes in predominant bacterial populations in human faeces with age and with Clostridium difficile infection. J Med Microbiol 51:448–454
Borody TJ, Warren EF, Leis SM, Surace R, Ashman O, Siarakas SJ (2004) Bacteriotherapy using fecal flora: toying with human motions. Clin Gastroenterol 38:475–483
Sullivan A, Edlund C, Nord CE (2001) Effect of antimicrobial agents on the ecological balance of human microflora. Lancet Infect Dis 1:101–114
Rousseau C, Levenez F, Fouqueray C, Doré J, Collignon A, Lepage P (2011) Clostridium difficile colonization in early infancy is accompanied by changes in intestinal microbiota composition. J Clin Microbiol 49:858–865
Vickerman MM, Brossard KA, Funk DB, Jesionowski AM, Gill SR (2007) Phylogenetic analysis of bacterial and archaeal species in symptomatic and asymptomatic endodontic infections. J Med Microbiol 56:110–118
Bélanger SD, Boissinot M, Clairoux N, Picard FJ, Bergeron MG (2003) Rapid detection of Clostridium difficile in feces by real-time PCR. J Clin Microbiol 41:730–734
Rinttilä T, Kassinen A, Malinen E, Krogius L, Palva A (2004) Development of an extensive set of 16S rDNA-targeted primers for quantification of pathogenic and indigenous bacteria in faecal samples by real-time PCR. J Appl Microbiol 97:1166–1177
Kovacs A (2009) Gut microbial ecology and its use in studying inflammatory bowel diseases. MSc Thesis, Tel-Aviv University
Merrigan MM, Sambol SP, Johnson S, Gerding DN (2003) Prevention of fatal Clostridium difficile-associated disease during continuous administration of clindamycin in hamsters. J Infect Dis 188:1922–1927
Sambol SP, Merrigan MM, Tang JK, Johnson S, Gerding DN (2002) Colonization for the prevention of Clostridium difficile disease in hamsters. J Infect Dis 186:1781–1789
Mazmanian SK, Liu CH, Tzianabos AO, Kasper DL (2005) An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122:107–118
Mazmanian SK, Round JL, Kasper DL (2008) A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453:620–625
van der Waaij D (1988) Evidence of immunoregulation of the composition of intestinal microflora and its practical consequences. Eur J Clin Microbiol Infect Dis 7:103–106
Wilson KH (1993) The microecology of Clostridium difficile. Clin Infect Dis 16(Suppl 4):S214–S218
Manges AR, Labbe A, Loo VG, Atherton JK, Behr MA, Masson L, Tellis PA, Brousseau R (2010) Comparative metagenomic study of alterations to the intestinal microbiota and risk of nosocomial Clostridum difficile-associated disease. J Infect Dis 202:1877–1884
Boyanton BL Jr, Sural P, Loomis CR, Pesta C, Gonzalez-Krellwitz L, Robinson-Dunn B, Riska P (2012) Loop-mediated isothermal amplification compared to real-time PCR and enzyme immunoassay for toxigenic Clostridium difficile detection. J Clin Microbiol 50:640–645
Acknowledgments
The authors thank Prof. Yitzhak Haberfeld from the Department of Labor Studies, The Faculty of Social Sciences, Tel-Aviv University, Israel, for his assistance in statistical analysis.
Disclosure
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
E. Goldberg and I. Amir contributed equally to this work.
Rights and permissions
About this article
Cite this article
Goldberg, E., Amir, I., Zafran, M. et al. The correlation between Clostridium-difficile infection and human gut concentrations of Bacteroidetes phylum and clostridial species. Eur J Clin Microbiol Infect Dis 33, 377–383 (2014). https://doi.org/10.1007/s10096-013-1966-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-013-1966-x