Abstract
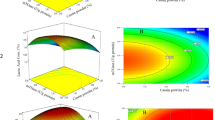
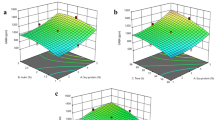
The requirements for the production of optimized Zea mays transglutaminase (TGZo) using Pichia pastoris GS115 (pPIC9K-tgzo) were optimized in this study. Plackett–Burman design was used to screen variables that significantly influence TGZo production. Oleic acid, methanol, and loading volume were identified as the most significant parameters. Central composite design was employed to determine the optimal level of these three parameters for TGZo production. Results showed that 1078 mU/mL of TGZo activity and 7.6 mg/L of TGZo production were obtained under conditions of 0.07% oleic acid, 1.31% methanol, and 7.36% loading volume. To explore the functional characteristics of TGZo, it was used in yogurt. It was found that the addition of TGZo could produce yogurt with stronger acid gel and higher consistency, cohesiveness, index of viscosity, and apparent viscosity than the untreated product. Therefore, TGZo can be used as a substitute for microbial transglutaminase in the yogurt, even in the food industry.


Similar content being viewed by others
References
Lorand L, Graham RM. Transglutaminases: crosslinking enzymes with pleiotropic functions. Nat. Rev. Mol. Cell Biol. 4: 140–156 (2003)
Kieliszek M, Misiewicz A. Microbial transglutaminase and its application in the food industry. A review. Folia Microbiol. 59: 241–250 (2014)
Jaros D, Partschefeld C, Henle T, Rohm H. Transglutaminase in dairy products: chemistry, physics, applications. J. Texture Stud. 37: 113–155 (2006)
Lin YS, Chao ML, Liu CH, Chu WS. Cloning and expression of the transglutaminase gene from Streptoverticillium ladakanum in Streptomyces lividans. Process Biochem. 39: 591–598 (2004)
Icekson I, Apelbaum A. Evidence for transglutaminase activity in plant tissue. Plant Physiol. 84: 972–974 (1987)
Kang H, Cho YD. Purification and properties of transglutaminase from soybean (Glycine max) leaves. Biochem. Biophys. Res. Commun. 223: 288–292 (1996)
Margosiak SA, Dharma A, Bruce-Carver MR, Gonzales AP, Louie D, Kuehn GD. Identification of the large subunit of ribulose 1, 5-bisphosphate carboxylase/oxygenase as a substrate for transglutaminase in Medicago sativa L.(Alfalfa). Plant Physiol. 92: 88–96 (1990)
Serafini-Fracassini D, Del Duca S, Beninati S. Plant transglutaminases. Phytochemistry. 40: 355–365 (1995)
Villalobos E, Santos M, Talavera D, Rodrıguez-Falcón M, Torné J. Molecular cloning and characterization of a maize transglutaminase complementary DNA. Gene. 336: 93–104 (2004)
Signorini M, Beninati S, Bergamini CM. Identification of transglutaminase activity in the leaves of silver beet (Beta vulgaris L.). J. Plant Physiol. 137: 547–552 (1991)
Della Mea M, Caparrós-Ruiz D, Claparols I, Serafini-Fracassini D, Rigau J. AtPng1p. The first plant transglutaminase. Plant Physiol. 135: 2046–2054 (2004)
Carvajal P, Gibert J, Campos N, Lopera O, Barberà E, Torné JM, Santos M. Activity of maize transglutaminase overexpressed in Escherichia coli inclusion bodies: An alternative to protein refolding. Biotechnol. Prog. 27: 232–240 (2011)
Li H, Cui Y, Zhang L, Luo X, Fan R, Xue C, Wang S, Liu W, Zhang S, Jiao Y, Du M, Yi H, Han X. Production of a transglutaminase from Zea mays in Escherichia coli and its impact on yoghurt properties. Int. J. Dairy Technol. 68: 54–61 (2015)
Li H, Cui Y, Zhang L, Yi H, Han X, Jiao Y, Du M, Fan R, Zhang S. Heterologous expression and purification of Zea mays transglutaminase in Pichia pastoris. Food Sci. and Biotech. 23: 1507–1513 (2014)
Li H, Zhang L, Cui Y, Luo X, Xue C, Wang S. Expression of soluble recombinant transglutaminase from Zea mays in Pichia pastoris. World J. Microb. Biot. 29: 939–947 (2013)
Li H, Zhang L, Cui Y, Luo X, Xue C, Wang S, Jiao Y, Zhang S, Liu W, Fan R. Characterization of recombinant Zea mays transglutaminase expressed in Pichia pastoris and its impact on full and non-fat yoghurts. J. Sci. Food Agric. 94: 1225–1230 (2014)
Huang H, Yang P, Luo H, Tang H, Shao N, Yuan T, Wang Y, Bai Y, Yao B. High-level expression of a truncated 1, 3-1, 4-β-D-glucanase from Fibrobacter succinogenes in Pichia pastoris by optimization of codons and fermentation. Appl. Microbiol. Biotechnol. 78: 95–103 (2008)
Liu S, Fang Y, Lv M, Wang S, Chen L. Optimization of the production of organic solvent-stable protease by Bacillus sphaericus DS11 with response surface methodology. Bioresource Technol. 101: 7924–7929 (2010)
Kim HS, Lee AY, Jo JE, Moon BC, Chun JM, Choi G, Kim HK. Optimization of ultrasound-assisted extraction of quercitrin from Houttuynia cordata Thunb. using respon sesurface methodology and UPLC analysis. Food Sci. Biotechnol. 23: 1–7 (2014)
Patil SA, Surwase SN, Jadhav SB, Jadhav JP. Optimization of medium using response surface methodology for L-DOPA production by Pseudomonas sp. SSA. Biochem. Eng. J. 74: 36–45 (2013)
Roukas T, Niavi P, Kotzekidou P. A new medium for spore production of Blakeslea trispora using response surface methodology. World J. Microb. Biot. 27: 307–317 (2011)
22. Şanlı T, Sezgin E, Deveci O, Şenel E, Benli M. Effect of using transglutaminase on physical, chemical and sensory properties of set-type yoghurt. Food Hydrocolloid. 25: 1477–1481 (2011)
Yüksel Z, Erdem Y. The influence of transglutaminase treatment on functional properties of set yoghurt. Int. J. Dairy Technol. 63: 86–97 (2010)
Zhang L, Zhang L, Yi H, Du M, Ma C, Han X, Feng Z, Jiao Y, Zhang Y. Enzymatic characterization of transglutaminase from Streptomyces mobaraensis DSM 40587 in high salt and effect of enzymatic cross-linking of yak milk proteins on functional properties of stirred yogurt. J. Dairy Sci. 95: 3559–3568 (2012)
Grossowicz N, Wainfan E, Borek E, Waelsch H. The enzymatic formation of hydroxamic acids from glutamine and asparagine. J. Biol. Chem. 187: 111–125 (1950)
Farnsworth JP, Li J, Hendricks GM, Guo MR. Effects of transglutaminase treatment on functional properties and probiotic culture survivability of goat milk yogurt. Small Ruminant Res. 65: 113–121 (2006)
Mohana S, Shrivastava S, Divecha J, Madamwar D. Response surface methodology for optimization of medium for decolorization of textile dye Direct Black 22 by a novel bacterial consortium. Bioresource Technol. 99: 562–569 (2008)
Kim S, Warburton S, Boldogh I, Svensson C, Pon L, d’Anjou M, Stadheim TA, Choi BK. Regulation of alcohol oxidase 1 (AOX1) promoter and peroxisome biogenesis in different fermentation processes in Pichia pastoris. J. Biotechnol. 166: 174–181 (2013)
Antonenkov VD, Grunau S, Ohlmeier S, Hiltunen JK. Peroxisomes are oxidative organelles. Antioxid. Redox Signal 13: 525–537 (2010)
Sakai Y, Oku M, van der Klei IJ, Kiel JA. Pexophagy: autophagic degradation of peroxisomes. BBA-Mol. Cell. Res. 1763: 1767–1775 (2006)
Erdmann R, Blobel G. Giant peroxisomes in oleic acid-induced Saccharomyces cerevisiae lacking the peroxisomal membrane protein Pmp27p. J. Cell Biol. 128: 509–523 (1995)
Khatri NK, Hoffmann F. Impact of methanol concentration on secreted protein production in oxygen-limited cultures of recombinant Pichia pastoris. Biotechnol. Bioeng. 93: 871–879 (2006)
Ozer B, Avni Kirmaci H, Oztekin S, Hayaloglu A, Atamer M. Incorporation of microbial transglutaminase into non-fat yogurt production. Int. Dairy J. 17: 199–207 (2007)
Gauche C, Tomazi T, Barreto PLM, Ogliari PJ, Bordignon-Luiz MT. Physical properties of yoghurt manufactured with milk whey and transglutaminase. LWT- Food Sci. Technol. 42: 239–243 (2009)
Acknowledgements
This work was supported by the National Nature Science Foundation of China (Grant No. 31601496 and 31301545), China Postdoctoral Science Foundation (Grant No. 2014M560244), International Postdoctoral Exchange Fellowship Program of China (Grant No. 20150082) and Heilongjiang Postdoctoral Foundation (Grant No. LBH-Z13042).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Li, H., Cui, Y., Zhang, L. et al. Optimization of recombinant Zea mays transglutaminase production and its influence on the functional properties of yogurt. Food Sci Biotechnol 26, 723–730 (2017). https://doi.org/10.1007/s10068-017-0083-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10068-017-0083-5