Abstract
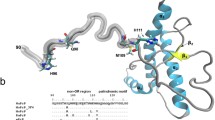
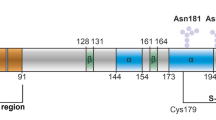
Prion-induced diseases are a global health concern. The lack of effective therapy and 100 % mortality rates for such diseases have made the prion protein an important target for drug discovery. Previous NMR experimental work revealed that thiamine and its derivatives bind the prion protein in a pocket near the N-terminal loop of helix 1, and conserved intermolecular interactions were noted between thiamine and other thiamine-binding proteins. Furthermore, water-mediated interactions were observed in all of the X-ray crystallographic structures of thiamine-binding proteins, but were not observed in the thiamine–prion NMR study. To better understand the potential role of water in thiamine–prion binding, a docking study was employed using structural X-ray solvent. Before energy minimization, docked thiamine assumed a “V” shape similar to some of the known thiamine-dependent proteins. Following minimization with NMR-derived restraints, the “F” conformation was observed. Our findings confirmed that water is involved in ligand stabilization and phosphate group interaction. The resulting refined structure of thiamine bound to the prion protein allowed the 4-aminopyrimidine ring of thiamine to π-stack with Tyr150, and facilitated hydrogen bonding between Asp147 and the amino group of 4-aminopyrimidine. Investigation of the π-stacking interaction through mutation of the tyrosine residue further revealed its importance in ligand placement. The resulting refined structure is in good agreement with previous experimental restraints, and is consistent with the pharmacophore model of thiamine-binding proteins.






Similar content being viewed by others
References
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216:136–144
Prusiner SB (1998) Prions. Proc Natl Acad Sci USA 95:13363–13383
Perez-Pineiro R, Bjorndahl TC, Berjanskii MV, Hau D, Li L, Huang A, Lee R, Gibbs E, Ladner C, Dong YW, Abera A, Cashman NR, Wishart DS (2011) The prion protein binds thiamine. FEBS J 278:4002–4014
Bjorndahl TC, Zhou GP, Liu XH, Perez-Pineiro R, Semenchenko V, Saleem F, Acharya S, Bujold A, Sobsey CA, Wishart DS (2011) Detailed biophysical characterization of the acid-Induced PrPc to PrP beta conversion process. Biochemistry 50:1162–1173
Kuwata K, Nishida N, Matsumoto T, Kamatari YO, Hosokawa-Muto J, Kodama K, Nakamura HK, Kimura K, Kawasaki M, Takakura Y, Shirabe S, Takata J, Kataoka Y, Katamine S (2007) Hot spots in prion protein for pathogenic conversion. Proc Natl Acad Sci USA 104:11921–11926
Timm DE, Liu JY, Baker LJ, Harris RA (2001) Crystal structure of thiamin pyrophosphokinase. J Mol Biol 310:195–204
Sippel KH, Robbins AH, Reutzel R, Domsic J, Boehlein SK, Govindasamy L, Agbandje-McKenna M, Rosser CJ, McKenna R (2008) Structure determination of the cancer-associated Mycoplasma hyorhinis protein Mh-p37. Acta Crystallogr D 64:1172–1178
Nemeria NS, Arjunan P, Chandrasekhar K, Mossad M, Tittmann K, Furey W, Jordan F (2010) Communication between thiamin cofactors in the Escherichia coli pyruvate dehydrogenase complex E1 component active centers: evidence for a “direct pathway” between the 4′-aminopyrimidine N1′ atoms. J Biol Chem 285:11197–11209
Onufriev A, Bashford D, Case DA (2000) Modification of the generalized Born model suitable for macromolecules. J Phys Chem B 104:3712–3720
Mackerell AD, Feig M, Brooks CL (2004) Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J Comput Chem 25:1400–1415
Clark M, Cramer RD, Vanopdenbosch N (1989) Validation of the general-purpose Tripos 5.2 force-field. J Comput Chem 10:982–1012
Pletcher J, Sax M (1972) Crystal and molecular structure of thiamine pyrophosphate hydrochloride. J Am Chem Soc 94:3998
Lindqvist Y, Schneider G, Ermler U, Sundstrom M (1992) Three-dimensional structure of transketolase, a thiamine diphosphate dependent enzyme, at 2.5 Å resolution. EMBO J 11:2373–2379
Muller YA, Schulz GE (1993) Structure of the thiamine and flavin-dependent enzyme pyruvate oxidase. Science 259:965–967
Iwashima A, Nishimura H (1979) Isolation of a thiamine-binding protein from Saccharomyces cerevisiae. Biochim Biophys Acta 577:217–220
Peapus DH, Chiu HJ, Campobasso N, Reddick JJ, Begley TP, Ealick SE (2001) Structural characterization of the enzyme-substrate, enzyme-intermediate, and enzyme-product complexes of thiamine phosphate synthase. Biochemistry 40:10103–10114
Kern D, Kern G, Neef H, Tittmann K, KillenbergJabs M, Wikner C, Schneider G, Hubner G (1997) How thiamine diphosphate is activated in enzymes. Science 275:67–70
Acknowledgments
The authors wish to acknowledge the financial support of the Alberta Prion Research Institute (APRI), Alberta Innovates Bio-Solutions, the National Research Council of Canada (NRC-NINT), and the Canadian Institutes of Health Research (CIHR).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 4.16 MB)
Rights and permissions
About this article
Cite this article
Pagadala, N.S., Bjorndahl, T.C., Blinov, N. et al. Molecular docking of thiamine reveals similarity in binding properties between the prion protein and other thiamine-binding proteins. J Mol Model 19, 5225–5235 (2013). https://doi.org/10.1007/s00894-013-1979-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-013-1979-5