Abstract
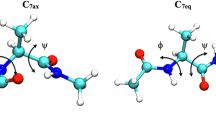
Main chain torsions of alanine dipeptide are parameterized into coupled 2-dimensional Fourier expansions based on quantum mechanical (QM) calculations at M06 2X/aug-cc-pvtz//HF/6-31G** level. Solvation effect is considered by employing polarizable continuum model. Utilization of the M06 2X functional leads to precise potential energy surface that is comparable to or even better than MP2 level, but with much less computational demand. Parameterization of the 2D expansions is against the full main chain torsion space instead of just a few low energy conformations. This procedure is similar to that for the development of AMBER03 force field, except unique weighting factor was assigned to all the grid points. To avoid inconsistency between quantum mechanical calculations and molecular modeling, the model peptide is further optimized at molecular mechanics level with main chain dihedral angles fixed before the calculation of the conformational energy on molecular mechanical level at each grid point, during which generalized Born model is employed. Difference in solvation models at quantum mechanics and molecular mechanics levels makes this parameterization procedure less straightforward. All force field parameters other than main chain torsions are taken from existing AMBER force field. With this new main chain torsion terms, we have studied the main chain dihedral distributions of ALA dipeptide and pentapeptide in aqueous solution. The results demonstrate that 2D main chain torsion is effective in delineating the energy variation associated with rotations along main chain dihedrals. This work is an implication for the necessity of more accurate description of main chain torsions in the future development of ab initio force field and it also raises a challenge to the development of quantum mechanical methods, especially the quantum mechanical solvation models.

A coupled two-dimensional main chain torsional potential for protein dynamics






Similar content being viewed by others
References
Duan Y, Wu C, Chowdhury S, Lee MC, Xiong G, Zhang W, Yang R, Cieplak P, Luo R, Lee T, Caldwell J, Wang J, Kollman P (2003) J Comput Chem 24:1999–2012
MacKerell AD Jr et al. (1998) J Phys Chem B 102:3586–3616
Kaminski GA, Friesner RA, Tirado-Rives J, Jorgensen WL (2001) J Phys Chem B 105:6474–6487
Oostenbrink C, Villa A, Mark AE, Van Gunsteren WF (2004) J Comput Chem 25:1656–1676
Bixon M, Lifson S (1967) Tetrahedron 23:769–784
Li D-W, Brüschweiler R (2010) Angew Chem Int Edit 49:6778–6780
Best RB, Hummer G (2009) J Phys Chem B 113:9004–9015
Nerenberg PS, Head-Gordon T (2011) J Chem Theor Comput 7:1220–1230
Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, Shaw DE (2010) Proteins 78:1950–1958
Jorgensen WL, Tirado-Rives J (2005) Proc Natl Acad Sci USA 102:6665–6670
Mackerell AD (2004) J Comput Chem 25:1584–1604
Ponder JW, Case DA (2003) Protein simulations. Academic, New York 66:27–85
Freddolino PL, Park S, Roux B, Schulten K (2009) Biophys J 96:3772–3780
Okur A, Strockbine B, Hornak V, Simmerling CJ (2003) J Comput Chem 24:21–31
Garcia AE, Sanbonmatsu KY (2002) Proc Natl Acad Sci USA 99:2782–2787
Kamiya N, Higo J, Nakamura H (2002) Protein Sci 11:2297–2307
Higo J, Ito N, Kuroda M, Ono S, Nakajima N, Nakamura H (2001) Protein Sci 10:1160–1171
Ono S, Nakajima N, Higo J, Nakamura HJ (2000) J Comput Chem 21:748–762
Wang L, Duan Y, Shortle R, Imperiali B, Kollman PA (1999) Protein Sci 8:1292–1304
Best RB, Buchete N-V, Hummer G (2008) Biophys J 95:L07–L09
Simmerling C, Strockbine B, Roitberg AE (2002) J Am Chem Soc 124:11258–11259
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Proteins 65:712–725
MacKerell AD, Feig M, Brooks CL (2004) J Am Chem Soc 126:698–699
Mackerell AD, Feig M, Brooks CL (2004) J Comput Chem 25:1400–1415
Wang Z-X, Zhang W, Wu C, Lei H, Cieplak P, Duan Y (2006) J Comput Chem 27:781–790
Sakae Y, Okamoto Y (2010) Mol Simulat 36:138–158
Sorin EJ, Pande VS (2005) Biophys J 88:2472–2493
Piana S, Lindorff-Larsen K, Shaw DE (2011) Biophys J 100:L47–L49
Kamiya N, Watanabe YS, Ono S, Higo J (2005) Chem Phys Lett 401:312–317
Iwaoka M, Tomoda S (2003) J Comput Chem 24:1192–1200
Zaman MH, Shen M-Y, Berry RS, Freed KF, Sosnick TR (2003) J Mol Biol 331:693–711
Liu ZW, Ensing B, Moore PB (2011) J Chem Theor Comput 7:402–419
Buck M, Bouguet-Bonnet S, Pastor RW, MacKerell AD (2006) Biophys J 90:L36–L38
Sakae Y, Okamoto Y (2006) J Phys Soc Jpn 75:054802
Case DA et al. (2010) AMBER 11. University of California, San Francisco
Head-Gordon M, Pople JA, Frisch MJ (1988) Chem Phys Lett 153:503–506
Scuseria GE, Janssen CL, Schaefer HF III (1988) J Chem Phys 89:7382–7387
Zhao Y, Truhlar DG (2007) Theor Chem Accounts 120:215–241
Zhao Y, Truhlar DG (2008) Accounts Chem Res 41:157–167
Tomasi J, Mennucci B, Cancès E (1999) J Mol Struct (Theochem) 464:211–226
Tomasi J, Mennucci B, Cammi R (2005) Chem Rev 105:2999–3094
Frisch MJ et al. (2010) Gaussian 09, revision B.01. Gaussian, Inc, Wallingford
Tsui V, Case DA (2000) Biopolymers 56:275–291
Cao Z, Lin Z, Wang J, Liu H (2009) J Comput Chem 30:645–660
Press WH, Teukolsky SA, Vetterling WT, Flannery BP (1986) Numerical recipes. Cambridge University Press, Cambridge, UK
Blondel A, Karplus M (1996) J Comput Chem 17:1132–1141
Sugita Y, Okamoto Y (1999) Chem Phys Lett 314:141–151
Onufriev A, Bashford D, Case DA (2004) Proteins 55:383–394
Mongan J, Simmerling C, McCammon JA, Case DA, Onufriev A (2007) J Chem Theor Comput 3:156–169
Jorgensen WL, Candrasekhar J, Madura JD, Impey RW, Klein ML (1983) J Chem Phys 79:926–935
Kumar S, Rosenberg JM, Bouzida D, Swendsen RH, Kollman PA (1992) J Comput Chem 13:1011–1021
Chodera JD, Swope WC, Pitera JW, Seok C, Dill KA (2007) J Chem Theor Comput 3:26–41
Kim YS, Wang JP, Hochstrasser RM (2005) J Phys Chem B 109:7511–7521
Grdadolnik J, Golic Grdadolnik S, Avbelj F (2008) J Phys Chem B 112:2712–2718
Mukhopadhyay P, Zuber G, Beratan DN (2008) Biophys J 95:5574–5586
Schweitzer-Stenner R, Measey T, Kakalis L, Jordan F, Pizzanelli S, Forte C, Griebenow K (2007) Biochemistry 46:1587–1596
Mehta MA, Fry EA, Eddy MT, Dedeo MT, Anagnost AE, Long JR (2004) J Phys Chem B 108:2777–2780
Beachy MD, Chasman D, Murphy RB, Halgren TA, Friesner RA (1997) J Am Chem Soc 119:5908–5920
Smith PE (1999) J Chem Phys 111:5568–5579
Vargas R, Garza J, Hay BP, Dixon DA (2002) J Phys Chem A 106:3213–3218
Hu H, Elstner M, Hermans J (2003) Proteins 50:451–463
Wang Z-X, Duan Y (2004) J Comput Chem 25:1699–1716
Kwac K, Lee K-K, Han JB, Oh K-I, Cho M (2008) J Chem Phys 128:105106
de Seabra GM, Walker RC, Roitberg AE (2009) J Phys Chem A 113:11938–11948
Gaigeot M-P (2009) J Phys Chem B 113:10059–10062
Mu YG, Kosov DS, Stock G (2003) J Phys Chem B 107:5064–5073
Graf J, Nguyen PH, Stock G, Schwalbe H (2007) J Am Chem Soc 129:1179–1189
Woutersen S, Pfister R, Hamm P, Mu YG, Kosov DS, Stock G (2002) J Chem Phys 117:6833–6840
Avbelj F, Grdadolnik SG, Grdadolnik J, Baldwin RL (2006) Proc Natl Acad Sci USA 103:1272–1277
Hu J-S, Bax A (1997) J Am Chem Soc 119:6360–6368
Case DA, Scheurer C, Brüschweiler R (2000) J Am Chem Soc 122:10390–10397
Acknowledgments
This work is supported by the National Natural Science Foundation of China (Grant No. 10974054, 20933002, 21173082 and 31200545), the Shanghai PuJiang Program (09PJ1404000), and the Shanghai Rising-Star Program (Grant No. 11QA1402000). We thank Supercomputer Center of East China Normal University for CPU time support.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yongxiu Li and Ya Gao contributed equally to this work
Rights and permissions
About this article
Cite this article
Li, Y., Gao, Y., Zhang, X. et al. A coupled two-dimensional main chain torsional potential for protein dynamics: generation and implementation. J Mol Model 19, 3647–3657 (2013). https://doi.org/10.1007/s00894-013-1879-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-013-1879-8