Abstract
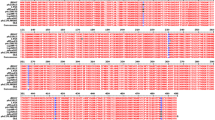
To identify a new morphological phenotype of erythromycin (EM)-resistant Staphylococcus aureus (S. aureus) were isolated in vitro from EM-sensitive parent strain, and the distribution of staphylococcus specific protein A (SpA) on the surface of these strains was examined morphologically by using applied immunoelectron microscopy. The isolated EM-resistant strains had thickened cell walls, and the distribution of SpA on the surfaces of these strains was demonstrated to be lower than that of the parent strain. The SpA suppression was confirmed by enzyme-linked immunosorbent assay (ELISA) using fixed EM-resistant cells. Moreover, the spa gene of EM-resistant cells was detected by polymerase chain reaction (PCR) and confirmed by quantitative real-time PCR assay, showing that the expression of SpA was repressed at the transcriptional level in these strains. Furthermore, ELISA assay showed that whole EM-resistant cell SpA content was significantly decreased. Therefore, it was considered that the suppression of surface SpA on the EM-resistant strain was due to regulated SpA production, and not dependent on the conformational change in SpA molecule expression through cell wall thickening. These results strongly suggest that suppressed SpA distribution on the EM-resistant S. aureus is a phenotypical characteristic in these strains.







Similar content being viewed by others
References
Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG Jr (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28:603–661
Halem M, Trent J, Green J, Kerdel F (2006) Community-acquired methicillin resistant Staphylococcus aureus skin infection. Semin Cutan Med Surg 25:68–71
Merino LA, Ronconi MC, Hreñuk GE, de Pepe MG (2003) Bacteriologic findings in patients with chronic sinusitis. Ear Nose Throat J 82:798–800, 803-804–806
Felmingham D (2004) Comparative antimicrobial susceptibility of respiratory tract pathogens. Chemotherapy 1:3–10
Sarrou S, Malli E, Tsilipounidaki K, Florou Z, Medvecky M, Skoulakis A, Hrabak J, Papagiannitsis CC, Petinaki E (2019) MLSB-Resistant Staphylococcus aureus in Central Greece: rate of resistance and molecular characterization. Microb Drug Resist 25:543–550
Sutcliffe J, Grebe T, Tait-Kamradt A, Wondrack L (1996) Detection of erythromycin-resistant determinants by PCR. Antimicrob Agents Chemother 40:2562–2566
Janosi L, Nakajima Y, Hashimoto H (1990) Characterization of plasmids that confer inducible resistance to 14-membered macrolides and streptogramin type B antibiotics in Staphylococcus aureus. Microbiol Immunol 34:723–735
Cui L, Ma X, Sato K, Okuma K, Tenover FC, Mamizuka EM, Gemmell CG, Kim MN, Ploy MC, El-Solh N, Ferraz V, Hiramatsu K (2003) Cell wall thickening is a common feature of vancomycin resistance in Staphylococcus aureus. J Clin Microbiol 41:5–14
Cui J, Zhang H, Mo Z, Yu M, Liang Z (2021) Cell wall thickness and the molecular mechanism of heterogeneous vancomycin-intermediate Staphylococcus aureus. Lett Appl Microbiol 72:604–609
Nishi H, Komatsuzawa H, Yamada S, Fujiwara T, Ohara M, Ohta K, Sugiyama M, Ishikawa T, Sugai M (2003) Moenomycin-resistance is associated with vancomycin-intermediate susceptibility in Staphylococcus aureus. Microbiol Immunol 47:927–935
Kawai M, Yamada S, Ishidoshiro A, Oyamada Y, Ito H, Yamagishi JI (2009) Cell-wall thickness: possible mechanism of acriflavine resistance in methicillin-resistant Staphylococcus aureus. J Med Microbiol 58:331–336
Hyo Y, Yamada S, Fukutsuji K, Harada T (2013) Thickening of the cell wall in macrolide-resistant Staphylococcus aureus. Med Mol Morphol 46:217–224
Fukutsuji K, Yamada S, Harada T (2013) Ultrastructural cell wall characteristics of clinical gentamycin-resistant Staphylococcus aureus isolates. Med Mol Morphol 46:70–76
Kim HK, Emolo C, DeDent AC, Falugi F, Missiakas DM, Schneewind O (2012) Protein A-specific monoclonal antibodies and prevention of Staphylococcus aureus disease in mice. Infect Immun 80:3460–3470
Pauli NT, Kim HK, Falugi F, Huang M, Dulac J, Henry Dunand C, Zheng NY, Kaur K, Andrews SF, Huang Y, DeDent A, Frank KM, Charnot-Katsikas A, Schneewind O, Wilson PC (2014) Staphylococcus aureus infection induces protein A-mediated immune evasion in humans. J Exp Med 211:2331–2339
Peterson PK, Verhoef J, Sabath LD, Quie PG (1977) Effect of protein A on staphylococcal opsonization. Infect Immun 15:760–764
Brignoli T, Manetti AGO, Rosini R, Haag AF, Scarlato V, Bagnoli F, Delany I (2019) Absence of protein A expression is associated with higher capsule production in staphylococcal isolates. Front Microbiol 10:863
Umeda A, Ikebuchi T, Amako K (1980) Localization of bacteriophage receptor, clumping factor, and protein A on the cell surface of Staphylococcus aureus. J Bacteriol 141:838–844
Yamada S, Matsumoto A (1988) Hemagglutination activity and localization of Fc receptor of group A and G streptococci. Microbiol Immunol 32:15–23
Yamada S, Yamagishi J, Matsumoto A (1996) Detection of Fc receptor genes from Staphylococcus aureus and streptococci by polymerase chain reaction. J Med Microbiol 45:507–511
Yamada S, Matsumoto A (1984) Localization of protein A on the cell surface of Staphylococcus aureus Cowan I and protein A-deficient strains. J Electron Microsc 33:172–174
Yamada S, Matsumoto A (1986) Effect of mannitol and glucose on the distribution and trypsin susceptibility of protein A of Staphylococcus aureus. Microbiol Immunol 30:993–1002
Hyo Y, Yamada S, Harada T (2008) Characteristic cell wall ultrastructure of a macrolide-resistant Staphylococcus capitis strain isolated from a patient with chronic sinusitis. Med Mol Morphol 41:160–164
Nagayama A, Yamaguchi K, Watanabe K, Tanaka M, Kobayashi I, Nagasawa Z (2007) Final report from the committee on antimicrobial susceptibility testing, Japanese society of chemotherapy, on the agar dilution method. J Infect Chemother 14:383–392
Uhlén M, Guss B, Nilsson B, Gatenbeck S, Philipson L, Lindberg M (1984) Complete sequence of the staphylococcal gene encoding protein A. A gene evolved through multiple duplications. J Biol Chem 259:1695–1702
Yamada S, Matsumoto A (1990) Surface properties of Staphylococcus aureus affecting chemiluminescence response of human phagocytes. Microbiol Immunol 34:809–817
Sjöquist J, Meloun B, Hjelm H (1972) Protein A isolated from Staphylococcus aureus after digestion with lysostaphin. Eur J Biochem 29:572–578
Jayakumar J, Kumar VA, Biswas L, Biswas R (2021) Therapeutic applications of lysostaphin against Staphylococcus aureus. J Appl Microbiol 131:1072–1082
Turner NA, Sharma-Kuinkel BK, Maskarinec SA, Eichenberger EM, Shah PP, Carugati M, Holland TL, Fowler VG Jr (2019) Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nat Rev Microbiol 17:203–218
Chaieb K, Zmantar T, Chehab O, Bouchami O, Ben Hasen A, Mahdouani K, Bakhrouf A (2007) Antibiotic resistance genes detected by multiplex PCR assays in Staphylococcus epidermidis strains isolated from dialysis fluid and needles in a dialysis service. Jpn J Infect Dis 60:183–187
Leclercq R (2002) Mechanisms of resistance to macrolides and lincosamides: nature of the resistance elements and their clinical implications. Clin Infect Dis 34:482–492
Martineau F, Picard FJ, Lansac N, Ménard C, Roy PH, Ouellette M, Bergeron MG (2000) Correlation between the resistance genotype determined by multiplex PCR assays and the antibiotic susceptibility patterns of Staphylococcus aureus and Staphylococcus epidermidis. Antimicrob Agents Chemother 44:231–238
Vandenesch F, Naimi T, Enright MC, Lina G, Nimmo GR, Heffernan H, Liassine N, Bes M, Greenland T, Reverdy ME, Etienne J (2003) Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: worldwide emergence. Emerg Infect Dis 9:978–984
Movitz J (1974) A study on the biosynthesis of protein A in Staphylococcus aureus. Eur J Biochem 48:131–136
Forsgren A (1969) Protein A from Staphylococcus aureus. Production of protein A by bacterial and L forms of S. aureus. Acta Pathol Microbiol Scand 75:481–490
Winblad S, Ericson C (1973) Sensitized sheep red cells as a reactant for Staphylococcus aureus protein A. Methodology and epidemiology with special reference to weakly reacting methicillin-resistant strains. Acta Pathol Microbiol Scand B Microbiol Immunol 81:150–156
Santos-Júnior CD, Veríssimo A, Costa J (2016) The recombination dynamics of Staphylococcus aureus inferred from spa gene. BMC Microbiol 16:143
Balachandran M, Bemis DA, Kania SA (2018) Expression and function of protein A in Staphylococcus pseudintermedius. Virulence 9:390–401
Goodyear CS, Silverman GJ (2003) Death by a B cell superantigen: In vivo VH-targeted apoptotic supraclonal B cell deletion by a Staphylococcal Toxin. J Exp Med 197:1125–1139
Kobayashi SD, DeLeo FR (2013) Staphylococcus aureus protein A promotes immune suppression. MBio 4:e00764-e813
Merino N, Toledo-Arana A, Vergara-Irigaray M, Valle J, Solano C, Calvo E, Lopez JA, Foster TJ, Penadés JR, Lasa I (2009) Protein A-mediated multicellular behavior in Staphylococcus aureus. J Bacteriol 191:832–843
Hoppenbrouwers T, Sultan AR, Abraham TE, Lemmens-den Toom NA, Hansenová Maňásková S, van Cappellen WA, Houtsmuller AB, van Wamel WJB, de Maat MPM, van Neck JW (2018) Staphylococcal protein A is a key factor in neutrophil extracellular traps formation. Front Immunol 9:165
Shi M, Willing SE, Kim HK, Schneewind O, Missiakas D (2021) Peptidoglycan contribution to the B cell superantigen activity of Staphylococcal protein A. MBio 12:e00039-e121
Cheung AL, Eberhardt K, Heinrichs JH (1997) Regulation of protein A synthesis by the sar and agr loci of Staphylococcus aureus. Infect Immun 65:2243–2249
Jenul C, Horswill AR (2019) Regulation of Staphylococcus aureus virulence. Microbiol Spectr. https://doi.org/10.1128/microbiolspec
Bronner S, Monteil H, Prévost G (2004) Regulation of virulence determinants in Staphylococcus aureus: complexity and applications. FEMS Microbiol Rev 8:183–200
Tan L, Li SR, Jiang B, Hu XM, Li S (2018) Therapeutic targeting of the Staphylococcus aureus accessory gene regulator (agr) system. Front Microbiol 9:55
Huntzinger E, Boisset S, Saveanu C, Benito Y, Geissmann T, Namane A, Lina G, Etienne J, Ehresmann B, Ehresmann C, Jacquier A, Vandenesch F, Romby P (2005) Staphylococcus aureus RNAIII and the endoribonuclease III coordinately regulate spa gene expression. EMBO J 24:824–835
Gao J, Stewart GC (2004) Regulatory elements of the Staphylococcus aureus protein A (SpA) promoter. J Bacteriol 186:3738–3748
Schmidt KA, Manna AC, Cheung AL (2003) SarT influences sarS expression in Staphylococcus aureus. Infect Immun 71:5139–5148
Acknowledgements
We thank K. Isoda from the Electron Microscope Center, Kawasaki Medical School, for the excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Okabe, K., Chikasue, K., Murakami, K. et al. Suppressed distribution of protein A on the surface of Staphylococcus aureus as a morphological characteristic of erythromycin-resistant strain. Med Mol Morphol (2024). https://doi.org/10.1007/s00795-023-00379-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00795-023-00379-4