Abstract
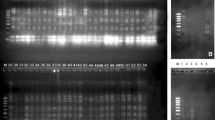
Proteins are important biochemical parameters in genetic diversity and controlling morphological characteristics in plants. In this study, the proteomic and morphometric data of an important medicinal herb “Aci Paşa” (Andrographis paniculata) were combined together to illustrate their impacts on genetic variation of the plant’s population and to realize the connection between protein patterns and phenotypic behavior of the species. We used three protein extraction buffers including Tris, potassium phosphate, and sodium citrate. The Tris buffer was significantly different (p ≤ 0.01) than other two in terms of the quality and quantity of protein bands by producing 15 types of proteins ranged from 13 to 105 kDa of which two of them were polymorphic. Consequently, a total of 12 accessions of A. paniculata were subjected to morpho-proteomic analyses. The unweighted pair group method with arithmetic average cluster analysis of the accessions based on the protein data and morphological characteristics generated three and four clusters, respectively, at a Euclidean distance of 2.53 for the morphological traits. Moreover, seed proteins analysis revealed that the two polymorphic protein bands sized 20.5 (protein “b”) and 30 kDa (protein “a”) effectively diversified the morphological characteristics and phylogenetic relationships among the 12 accessions of A. paniculata. Interestingly, the protein “b” acted as an activator agent for the number of branches, leaves and total dry weight, while the protein “a” performed a suppressive role for the same traits. Additionally, the two high-weighted faint bands “c” (75 kDa) and “d” (100 kDa) with a very low expression in accession 11228 proved their suppressive role along with the “a” band, while these bands were strongly expressed in the rest of the accessions. These findings suggest that these four proteins should be sequenced and perfectly established for further proteomic analyses. Ultimately, the mentioned proteins can be developed for any prospective breeding program or gene identification.





Similar content being viewed by others
References
Akhila H, Beevy SS (2011) Morphological and seed protein characterization of the cultivated and the wild taxa of Sesamum L. (Pedaliaceae). Plant Syst Evol 293(1):65–70
Andrews M, Raven JA, Lea PJ, Sprent JI (2006) A role for shoot protein in shoot–root dry matter allocation in higher plants. Ann Bot 97:3–10
Asante I, Offei S, Addy R, Carson A (2009) Phenotypic and seed protein analysis in 31 Lima bean (Phaseolus lunatus) accessions in Ghana. West African J Appl Ecol 12(1):1–10
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254
Campa A, Pañeda A, Pérez-Vega E, Giraldez R, Ferreira JJ (2011) Mapping and use of seed protein loci for marker-assisted selection of growth habit and photoperiod response in Nuña bean (Phaseolus vulgaris L.). Euphytica 179:383–391
Chang YP, Ding SF, Chou CC, Du BS, Chen WS (1998) Daylength affects protein pattern and flowering in tuberose (Polianthes tuberosa L.). Bot Bull Acad Sin 39:199–203
Cho K, Agrawal GK, Shibato J, Jung YH, Kim YK, Nahm BH, Jwa NS, Tamogami S, Han O, Kohda K, Iwahashi H, Rakwal R (2007) Survey of differentially expressed proteins and genes in jasmonic acid treated rice seedling shoot and root at the proteomics and transcriptomics levels. J Proteome Res 6:3581–3603
Durst RA, Staples BR (1972) Tris/Tris HCl: a standard buffer for use in the physiologic pH range. Clin Chem 18(3):206–208
Gepts P, Brown A, Clegg M, Kahler A, Weir B (1990) Genetic diversity of seed storage proteins in plants. In: Brown AHD, Clegg MT, Khaler AL, Weir BS (eds) Plant population genetics, breeding, and genetic resources. Sinauer Associates Inc, Sunderland, pp 64–82
Hanada K, Kuromori T, Myouga F, Toyoda T, Shinozaki K (2009) Increased expression and protein divergence in duplicate genes is associated with morphological diversification. PLoS Genet 5(12):e1000781. doi:10.1371/journal.pgen.1000781
He C, Tian Y, Saedler R, Efremova N, Riss S, Khan MR, Yephremov A, Saedler H (2010) The MADS-domain protein MPF1 of Physalis floridana controls plant architecture, seed development and flowering time. Planta 231:767–777
Huang JF, Zhang ML, Cohen JI (2013) Phylogenetic analysis of Lappula Moench (Boraginaceae) based on molecular and morphological data. Plant Syst Evol 299(5):913–926. doi:10.1007/s00606-013-0772-3
Jehan T, Lakhanpaul S (2006) Single nucleotide polymorphism (SNP)-methods and applications in plant genetics: a review. Indian J Biotechnol 5:435–459
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Ries-Kautt M, Ducruix A (1997) Inferences drawn from physicochemical studies of crystallogenesis and precrystalline state. Meth Enzymol 276:23–59
Sabu KK, Padmesh P, Seeni S (2001) Intraspecific variation in active principle content and isozymes of Andrographis paniculata Nees (Kalmegh): a traditional hepatoprotective medicinal herb of Indian J Medic Aroma. Plant Sci 23:637–647
SAS Institute Inc (2009a) SAS, SAS/Sat user’s guide, Release 9.1. SAS Institute Inc, Cary (Licenced to Institute UPM)
SAS Institute Inc (2009b) JMP® 8 User Guide, 2nd edn. SAS Institute Inc, Cary
Schmittschmitt JP, Scholtz JM (2009) The role of protein stability, solubility, and net charge in amyloid fibril formation. Protein Sci 12(10):2374–2378
Sheidai M, Hamta A, Jaffari A, Noori-Daloii M (2000) Morphometric and seed protein studies of Trifolium species and cultivars in Iran. Plant Genet Res Newsletter 120:52–54
Talei D, Fotokian MH (2008) Assessment of relationship between bacterial stripe resistance and leaf protein bands in rice (Oryza sativa L.) varieties. AIP Conf Proc 971:211–215
Talei D, Valdiani A, Abdullah MP, Hassan SA (2012) A rapid and effective method for dormancy breakage and germination of King of Bitters (Andrographis paniculata Nees) seeds. Maydica 57:98–105
Valdiani A, Kadir MA, Tan SG, Talei D, Puad MA, Nikzad S (2012a) Nain-e Havandi (Andrographis paniculata) present yesterday, absent today: a plenary review on underutilized herb of Iran’s pharmaceutical plants. Mol Biol Rep 39:5409–5424
Valdiani A, Kadir MA, Saad MS, Talei D, Tan SG (2012b) Intra-specific hybridization: generator of genetic diversification and heterosis in Andrographis paniculata Nees. A bridge from extinction to survival. Gene 505(1):23–36
Valdiani A, Javanmard A, Talei D, Tan SG, Nikzad S, Kadir MA, Abdullah SNA (2013) Microsatellite-based evidences of genetic bottlenecks in the cryptic species “Andrographis paniculata Nees”: a potential anticancer agent. Mol Biol Rep 40:1775–1784
Victor KJ, Fennell AY, Grimplet J (2010) Proteomic analysis of shoot tissue during photoperiod induced growth cessation in V. riparia Michx. grapevines. Proteome Sci 8(44):2–17
Acknowledgments
The authors express their gratitude to Universiti Putra Malaysia for supporting this research.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Talei, D., Valdiani, A. & Abdullah, M.P. Impact of protein diversification on morphometric behavior of Andrographis paniculata Nees. Plant Syst Evol 300, 1003–1010 (2014). https://doi.org/10.1007/s00606-013-0938-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-013-0938-z