Abstract
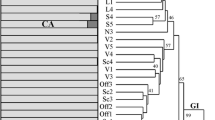
DNA fingerprinting (AFLP) and chemical analyses of essential oils were utilized to define the extent of variation existing in the genus Ocimum. Research was carried out on 22 Ocimum accessions representing seven species. Concerning the essential oil composition of all investigated accessions, 115 compounds were identified. UPGMA cluster analysis, based on Euclidian distances of essential oil constituents between all pairs of accessions, showed four well-supported clusters (O. tenuiflorum, O. basilicum/O. africanum, O. basilicum, and O. americanum/O. africanum). Relating to the essential oil composition of all of the investigated accessions, 17 compounds were identified as the main ones, and according to them 13 chemotypes were determined. AFLP relationships were determined by neighbor-joining (NJ) cluster analysis based on Dice’s distance matrix and by maximum parsimony (MP) analysis. O. basilicum, O. americanum/O. africanum, O. tenuiflorum, and O. gratissimum represented four clusters supported with high bootstrap values. A neighbor-net diagram allowed the visualization of apparently conflicting data by revealing relationships between genotypes and chemotypes. Concerning the O. africanum species, two distinct chemotypes, geranial/neral (accession 11) and estragol (accession 10), have been established, while all the studied O. americanum accessions belong to the geranial/neral chemotype. This could be additional evidence that O. americanum is one of the parents of O. africanum. Furthermore, the fact that the O. africanum accession (10) as well as O. basilicum ‘Purpurascens’ and O. basilicum ‘Erevanskii’ accessions belong to the estragol chemotype supports the theory that O. africanum is one of the parents of these two O. basilicum accessions.



Similar content being viewed by others
References
Adams RP (1995) Identification of essential oil components by gas chromatography and mass spectroscopy. Allured Publishing Co, Carol Stream
Bryant D, Moulton V (2004) Neighbor-net: an agglomerative method for the construction of phylogenetic networks. Mol Biol Evol 21(2):255–265
Carović-Stanko K, Liber Z, Besendorfer V, Javornik B, Bohanec B, Kolak I, Satovic Z (2010a) Genetic relations among basil taxa (Ocimum L.) based on molecular markers, nuclear DNA content, and chromosome number. Plant Syst Evol 285(1-2):13–22
Carović-Stanko K, Orlić S, Politeo O, Strikić F, Kolak I, Milos M, Satovic Z (2010b) Composition and antibacterial activities of essential oils of seven Ocimum taxa. Food Chem 119(1):196–201
Carović-Stanko K, Šalinović A, Grdiša M, Liber Z, Kolak I, Satovic Z (2011) Efficiency of morphological trait descriptors in discrimination of Ocimum basilicum L. accessions. Plant Biosyst 145 (in press)
De Masi L, Siviero P, Esposito C, Castaldo D, Siano F, Laratta B (2006) Assessment of agronomic, chemical and genetic variability in common Basil (Ocimum basilicum L.). Eur Food Res Technol 223:273–281
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Farris JS (1969) A successive approximations approach to character weighting. Syst Zool 18:374–385
Felsenstein J (1985) Confidence limits on phylogenesis: an approach using the bootstrap. Evolution 39:783–791
Grayer RJ, Kite GC, Goldstone FJ, Bryan SE, Paton A, Putievsky E (1996) Intraspecific taxonomy and essential oil chemotypes in sweet basil, Ocimum basilicum. Phytochemistry 43:1033–1039
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontolo Electron 4(1):9
Hillis DM, Huelsenbeck JP (1992) Signal, noise and reliability in molecular phylogenetic analyses. J Hered 83:189–195
Hiltunen R, Holm Y (1999) Essential oil of Ocimum. In: Hiltunen H, Holm Y (eds) Basil: the genus Ocimum. Harwood Academic Publishers, Amsterdam, pp 77–111
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Khosla MK (1995) Study of inter-relationship, phylogeny and evolutionary tendencies in genus Ocimum. Ind J Genet 55:71–83
Kothari SK, Bhattacharya AK, Ramesh S, garg SN, Khanuja SPS (2005) Volatile constituents in oil from different plant parts of methyl eugenol-rich Ocimum tenuiflorum L. f. (syn. O. sanctum) growing in South India. J Essent Oil Res 17:656–658
Labra M, Miele M, Ledda B, Grassi F, Mazzei M, Sala F (2004) Morphological characterization, essential oil composition and DNA genotyping of Ocimum basilicum L. cultivars. Plant Sci 167:725–731
Lawrence BM (1993) Labiatae oils: mother nature’s chemical factory. In: Essential oils. Allured Publishing, Carol Stream, pp 188–206
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Paton A (1992) A synopsis of Ocimum L. (Labiatae) in Africa. Kew Bull 47:405–437
Paton A, Harley MR, Harley MM (1999) Ocimum: an overview of classification and relationships. In: Hiltunen R, Holm Y (eds) Basil: the genus Ocimum. Harwood Academic Publishers, Amsterdam, pp 1–38
Pino J, Rosado A, Fuentes V (1996) Composition of the essential oil from the leaves and flowers of Ocimum gratissimum L. grown in Cuba. J Ess Oil Res 8:139–141
Pushpangadan P, Bradu BL (1995) Medicinal and aromatic plants. In: Chadha KL, Gupta R (eds) Advances in horticulture, vol 11. Malhotra Publishing House, New Delhi, pp 627–657
Rohlf FJ (2000) NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 2.1. Exeter Publications, New York
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 6:514–525
Satovic Z, Liber Z, Karlovic K, Kolak I (2002) Genetic relatednessamong basil (Ocimum spp.) accessions using RAPD markers. Acta Biol Cracov Bot 44:155–160
Simon JE, Quinn J, Murray RG (1990) Basil: a source of essential oils. In: Janick J, Simon JE (eds) Advances in new crops. Timber Press, Portland, pp 484–989
Singh AP, Dwivedi S, Bharti S, Srivastava A, Singh V, Khanuja SPS (2004) Phylogenetic relationships as in Ocimum revealed by RAPD markers. Euphytica 136:11–20
Sobti SN, Pushpangadan P (1982) Studies in the genus Ocimum: cytogenetics, breeding and utilization of aromatic plants. In: Atal CK, Kapur BM (eds) Regional research laboratory. Jammu-Tawi, India, pp 457–472
Swofford DL (2003) PAUP*. Phylogenetic analysis using parsimony (*and Other Methods), Version 4. Sinauer, Sunderland
Van de Peer Y, De Wachter R (1994) TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Comput Appl Biosci 10:569–570
Van Den Dool H, Kratz PD (1963) A generalization of the retention index system including linear temperature programmed gas–liquid partition chromatography. J Chromatogr 11:463–471
Vieira RF, Simon JE (2000) Chemical characterization of basil (Ocimum spp.) founding the markets and used in traditional medicine in Brazil. Econ Bot 54(2):207–216
Vieira RF, Grayer RJ, Paton A, Simon JE (2001) Genetic diversity of Ocimum gratissimum L. based on volatile oil constituents, flavonoids and RAPD markers. Biochem Syst Ecol 29:287–304
Vieira RF, Goldsbrough P, Simon JE (2003) Genetic diversity of basil (Ocimum spp.) based on RAPD markers. J Am Soc Hort Sci 128:94–99
Vos P, Hogers R, Bleeker M, Reijans M, Van de Lee T, Hornes M, Frijters A, Plot J, Peleman J, Kuipe M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucl Acids Res 23:4407–4414
Zheljazkov VD, Canterll CL, Tekwani B, Khan SI (2008a) Content, Composition, and Bioactivity of Three Basil Genotypes as a Function of Harvesting. J Agric Food Chem 56:385–385
Zheljazkov VD, Callahan A, Canterll CL (2008b) Yield and Oil Composition of 38 Basil (Ocimum basilicum L.) accessions grown in Mississippi. J Agr Food Chem 56:241–245
Acknowledgments
This study was supported by the Ministry of Science, Education, and Sports of the Republic of Croatia, within the framework of Project No. 178-1191193-0212 and Project No. 119-1191193-1232.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Carović-Stanko, K., Liber, Z., Politeo, O. et al. Molecular and chemical characterization of the most widespread Ocimum species. Plant Syst Evol 294, 253–262 (2011). https://doi.org/10.1007/s00606-011-0471-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-011-0471-x